| Record Information |
|---|
| Version | 2.0 |
|---|
| Creation Date | 2009-03-06 18:58:04 UTC |
|---|
| Update Date | 2014-12-24 20:21:06 UTC |
|---|
| Accession Number | T3D0099 |
|---|
| Identification |
|---|
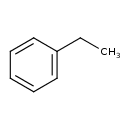
| Common Name | Ethylbenzene |
|---|
| Class | Small Molecule |
|---|
| Description | Ethylbenzene is an organic compound with the formula C6H5CH2CH3. This aromatic hydrocarbon is important in the petrochemical industry as an intermediate in the production of styrene, which in turn is used for making polystyrene, a commonly used plastic material. Although often present in small amounts in crude oil, ethylbenzene is produced in bulk quantities by combining benzene and ethylene in an acid-catalyzed chemical reaction. It is one ingredient of cigarette. The acute toxicity of ethylbenzene is low, with an LD50 of about 4 grams per kilogram of body weight. The longer term toxicity and carcinogenicity is ambiguous. Eye and throat sensitivity can occur when high level exposure to ethylbenzene in the air occurs. At higher level exposure, ethylbenzene can cause dizziness. |
|---|
| Compound Type | - Aromatic Hydrocarbon
- Cigarette Toxin
- Food Toxin
- Household Toxin
- Industrial Precursor/Intermediate
- Industrial/Workplace Toxin
- Metabolite
- Organic Compound
- Pollutant
- Synthetic Compound
|
|---|
| Chemical Structure | |
|---|
| Synonyms | | Synonym | | Aethylbenzol | | alpha-methyltoluene | | Ethylbenzol | | Phenylethane |
|
|---|
| Chemical Formula | C8H10 |
|---|
| Average Molecular Mass | 106.165 g/mol |
|---|
| Monoisotopic Mass | 106.078 g/mol |
|---|
| CAS Registry Number | 100-41-4 |
|---|
| IUPAC Name | ethylbenzene |
|---|
| Traditional Name | ethylbenzene |
|---|
| SMILES | CCC1=CC=CC=C1 |
|---|
| InChI Identifier | InChI=1S/C8H10/c1-2-8-6-4-3-5-7-8/h3-7H,2H2,1H3 |
|---|
| InChI Key | InChIKey=YNQLUTRBYVCPMQ-UHFFFAOYSA-N |
|---|
| Chemical Taxonomy |
|---|
| Description | belongs to the class of organic compounds known as benzene and substituted derivatives. These are aromatic compounds containing one monocyclic ring system consisting of benzene. |
|---|
| Kingdom | Organic compounds |
|---|
| Super Class | Benzenoids |
|---|
| Class | Benzene and substituted derivatives |
|---|
| Sub Class | Not Available |
|---|
| Direct Parent | Benzene and substituted derivatives |
|---|
| Alternative Parents | |
|---|
| Substituents | - Monocyclic benzene moiety
- Aromatic hydrocarbon
- Unsaturated hydrocarbon
- Hydrocarbon
- Aromatic homomonocyclic compound
|
|---|
| Molecular Framework | Aromatic homomonocyclic compounds |
|---|
| External Descriptors | |
|---|
| Biological Properties |
|---|
| Status | Detected and Not Quantified |
|---|
| Origin | Exogenous |
|---|
| Cellular Locations | |
|---|
| Biofluid Locations | Not Available |
|---|
| Tissue Locations | Not Available |
|---|
| Pathways | Not Available |
|---|
| Applications | Not Available |
|---|
| Biological Roles | Not Available |
|---|
| Chemical Roles | Not Available |
|---|
| Physical Properties |
|---|
| State | Liquid |
|---|
| Appearance | Not Available |
|---|
| Experimental Properties | | Property | Value |
|---|
| Melting Point | -94.9°C | | Boiling Point | Not Available | | Solubility | 0.169 mg/mL at 25 °C [SANEMASA,I et al. (1982)] | | LogP | Not Available |
|
|---|
| Predicted Properties | |
|---|
| Spectra |
|---|
| Spectra | | Spectrum Type | Description | Splash Key | Deposition Date | View |
|---|
| GC-MS | GC-MS Spectrum - EI-B (Non-derivatized) | splash10-0006-9000000000-e311097b1353d1f46e6e | 2017-09-12 | View Spectrum | | GC-MS | GC-MS Spectrum - EI-B (Non-derivatized) | splash10-0006-9200000000-c2b5306fbaeb48134d6b | 2017-09-12 | View Spectrum | | GC-MS | GC-MS Spectrum - CI-B (Non-derivatized) | splash10-0a4i-1900000000-ff5c54f00a3ae0d72793 | 2017-09-12 | View Spectrum | | GC-MS | GC-MS Spectrum - EI-B (Non-derivatized) | splash10-0006-9000000000-e311097b1353d1f46e6e | 2018-05-18 | View Spectrum | | GC-MS | GC-MS Spectrum - EI-B (Non-derivatized) | splash10-0006-9200000000-c2b5306fbaeb48134d6b | 2018-05-18 | View Spectrum | | GC-MS | GC-MS Spectrum - CI-B (Non-derivatized) | splash10-0a4i-1900000000-ff5c54f00a3ae0d72793 | 2018-05-18 | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | splash10-052f-9300000000-c7e86064f086caf02674 | 2016-09-22 | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | Not Available | 2021-10-12 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-0a4i-0900000000-3d28e81794c61465b233 | 2016-08-03 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-0a4i-1900000000-ea2447a1e61e24730a44 | 2016-08-03 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-052f-9100000000-1824048a45b025a1f4aa | 2016-08-03 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-0a4i-0900000000-8ca4acb96694435a7851 | 2016-08-03 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-0a4i-0900000000-5b9ea9d3d5b6f3bbb587 | 2016-08-03 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-0a70-9600000000-dc68589987b1b4f1e625 | 2016-08-03 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-0a4i-0900000000-f8066873e5f243968d61 | 2021-10-12 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-056r-9300000000-2aecf2aac0a60843c353 | 2021-10-12 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-00ou-9000000000-2708ae8f35c96209bba0 | 2021-10-12 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-0a4i-0900000000-861947f0491f909a2588 | 2021-10-12 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-0a4i-2900000000-1204b096ce8c38491d80 | 2021-10-12 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-004i-9000000000-fc58e0949de9ca4842ff | 2021-10-12 | View Spectrum | | MS | Mass Spectrum (Electron Ionization) | splash10-0006-9100000000-1cec483f116a110c0c16 | 2014-09-20 | View Spectrum | | 1D NMR | 1H NMR Spectrum (1D, 90 MHz, CDCl3, experimental) | Not Available | 2014-09-20 | View Spectrum | | 1D NMR | 13C NMR Spectrum (1D, 25.16 MHz, CDCl3, experimental) | Not Available | 2014-09-23 | View Spectrum |
|
|---|
| Toxicity Profile |
|---|
| Route of Exposure | Oral (7); inhalation (7) ; dermal (7). |
|---|
| Mechanism of Toxicity | Changes in the integrity of the cell membrane after partitioning of ethylbenzene into the lipid bilayer may subsequently affect the function of membrane, particularly as a barrier and in energy transduction, and in the formation of a matrix for proteins and enzymes. Ethybenzene inhibits the activity of the astrocytic membrane ATPases, which helps regulate adequate intercellular levels of ions, nutrients, metabolic intermediates and precursors in the central nervous system. Thus, this may disturb the ability of the cells to maintain homeostasis. (7, 1) |
|---|
| Metabolism | Ethylbenzene is metabolized mainly through hydroxylation and then through conjugation reactions from which numerous metabolites have been isolated. Hydroxylation of ethylbenzene to 1-phenylethanol is catalyzed by cytochrome P-450 isoforms CYP2E1 and CYP2B6. 1-Phenylethanol is conjugated to glucuronide, which then is either excreted or converted to subsequent metabolites. Oxidation of 1-phenylethanol yields acetophenone, which is both excreted in the urine as a minor metabolite and further transformed. Continued oxidation of the side chain leads to the sequential formation of 2-hydroxyacetophenone, 1-phenyl-1,2-ethanediol, mandelic acid, and phenylglyoxylic acid. Minor pathways (e.g., ring hydroxylation) include glucuronide and sulfate conjugation with hydroxylated derivatives to form glucuronides and sulfates that are excreted in the urine. In humans exposed via inhalation, the major metabolites of ethylbenzene in the urine are mandelic acid (70%) and phenylglyoxylic acid (25%). Following dermal exposure of humans, however, excretion of mandelic acid was shown to be only 4.6% of the absorbed dose, which may indicate differences in the metabolic fate between inhalation and dermal exposure routes. (7) |
|---|
| Toxicity Values | LD50: 3.5 g/kg (Oral, Rat) (9)
LD50: 77.4 g/kg (Dermal, Rabbit)(9)
LC50: 17.2 g/m3 (4000 ppm) (Inhalation, Rat) (9) |
|---|
| Lethal Dose | Not Available |
|---|
| Carcinogenicity (IARC Classification) | 2B, possibly carcinogenic to humans. (6) |
|---|
| Uses/Sources | Ethylbenzene is used in the petrochemical industry as an intermediate in the production of styrene, which in turn is used for making polystyrene, a commonly used plastic material. Exposure may occur from breathing contaminated air, drinking or eating food prepared with ethylbenzene-contaminated water, and through skin contact with products containing ethylbenzene, such as gasoline. (7) |
|---|
| Minimum Risk Level | Acute Inhalation: 10 ppm (5)
Intermediate Inhalation: 0.7 ppm (5)
Chronic Inhalation: 0.3 ppm (5)
Intermediate Oral: 0.5 mg/kg/day (5) |
|---|
| Health Effects | Chronic exposure to etylbenzene can lead to an increase in the mean number of lymphocytes and a decrease in hemoglobin levels. Acute duration and intermediate duration studies suggest that the auditory system is a sensitive target of ethylbenzene toxicity. Exposure ethylbenzene can lead to functional and organic disturbances (nervous system disturbances, toxic hepatitis and upper respiratory tract complaints). Metabolites of ethylbenzene have been shown to produce oxidative damage to DNA. (7, 4) |
|---|
| Symptoms | Cough, sore throat, dizziness, drowsiness, and headache follow inhalation or ingestion exposure to ethylbenzene. Ingestion exposure can also lead to burning sensation in the throat and chest. Skin or eyes contact to ethylbenzene can lead to redness and pain of the exposed surface. (8) |
|---|
| Treatment | Following oral exposure, a gastric lavage is recommended. Protect airway by placement in Trendelenburg and left lateral decubitus position or by endotracheal intubation. Control any seizures first. Following inhalation, move patient to fresh air. Monitor for respiratory distress. If cough or difficulty breathing develops, evaluate for respiratory tract irritation, bronchitis, or pneumonitis. Administer oxygen and assist ventilation as required. Following eye exposure, irrigate exposed eyes with copious amounts of room temperature water for at least 15 minutes. In case of dermal exposure, remove contaminated clothing and wash exposed area thoroughly with soap and water. Treat dermal irritation or burns with standard topical therapy. Patients developing dermal hypersensitivity reactions may require treatment with systemic or topical corticosteroids or antihistamines. Some chemicals can produce systemic poisoning by absorption through intact skin. Carefully observe patients with dermal exposure for the development of any systemic signs or symptoms and administer symptomatic treatment as necessary. (3) |
|---|
| Normal Concentrations |
|---|
| Not Available |
|---|
| Abnormal Concentrations |
|---|
| Not Available |
|---|
| External Links |
|---|
| DrugBank ID | DB01722 |
|---|
| HMDB ID | HMDB59905 |
|---|
| PubChem Compound ID | 7500 |
|---|
| ChEMBL ID | CHEMBL371561 |
|---|
| ChemSpider ID | 7219 |
|---|
| KEGG ID | C07111 |
|---|
| UniProt ID | Not Available |
|---|
| OMIM ID | |
|---|
| ChEBI ID | 16101 |
|---|
| BioCyc ID | CPD-9502 |
|---|
| CTD ID | C004912 |
|---|
| Stitch ID | Ethylbenzene |
|---|
| PDB ID | Not Available |
|---|
| ACToR ID | 606 |
|---|
| Wikipedia Link | Ethylbenzene |
|---|
| References |
|---|
| Synthesis Reference | Guenther Heimlich, Gregor Tremmel, Manfred Lieb, “Continuous preparation of ethylbenzene in a heterogeneous-phase reaction.” U.S. Patent US4431854, issued July, 1950. |
|---|
| MSDS | T3D0099.pdf |
|---|
| General References | - Vaalavirta L, Tahti H: Astrocyte membrane Na+, K(+)-ATPase and Mg(2+)-ATPase as targets of organic solvent impact. Life Sci. 1995;57(24):2223-30. [7475975 ]
- Sams C, Loizou GD, Cocker J, Lennard MS: Metabolism of ethylbenzene by human liver microsomes and recombinant human cytochrome P450s (CYP). Toxicol Lett. 2004 Mar 7;147(3):253-60. [15104117 ]
- Rumack BH (2009). POISINDEX(R) Information System. Englewood, CO: Micromedex, Inc. CCIS Volume 141, edition expires Aug, 2009.
- International Labour Office (1983). Encyclopedia of Occupational Health and Safety. Volumes. I and II. Geneva, Switzerland: International Labour Office.
- ATSDR - Agency for Toxic Substances and Disease Registry (2001). Minimal Risk Levels (MRLs) for Hazardous Substances. U.S. Public Health Service in collaboration with U.S. Environmental Protection Agency (EPA). [Link]
- International Agency for Research on Cancer (2014). IARC Monographs on the Evaluation of Carcinogenic Risks to Humans. [Link]
- ATSDR - Agency for Toxic Substances and Disease Registry (2007). Toxicological profile for ethylbenzene. U.S. Public Health Service in collaboration with U.S. Environmental Protection Agency (EPA). [Link]
- International Programme on Chemical Safety (IPCS) INCHEM (2007). Poison Information Monograph for Ethylbenzne. [Link]
- International Programme on Chemical Safety (IPCS) INCHEM (1996). Environmental Health Criteria for Ethylbenzene. [Link]
|
|---|
| Gene Regulation |
|---|
| Up-Regulated Genes | | Gene | Gene Symbol | Gene ID | Interaction | Chromosome | Details |
|---|
|
|---|
| Down-Regulated Genes | | Gene | Gene Symbol | Gene ID | Interaction | Chromosome | Details |
|---|
|
|---|