| Record Information |
|---|
| Version | 2.0 |
|---|
| Creation Date | 2009-03-06 18:58:21 UTC |
|---|
| Update Date | 2014-12-24 20:21:24 UTC |
|---|
| Accession Number | T3D0246 |
|---|
| Identification |
|---|
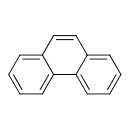
| Common Name | Phenanthrene |
|---|
| Class | Small Molecule |
|---|
| Description | Phenanthrene is one of over 100 different polycyclic aromatic hydrocarbons (PAHs). PAHs are chemicals that are formed during the incomplete burning of organic substances, such as fossil fuels. They are usually found as a mixture containing two or more of these compounds. (4) |
|---|
| Compound Type | - Aromatic Hydrocarbon
- Food Toxin
- Industrial By-product/Pollutant
- Industrial/Workplace Toxin
- Natural Compound
- Organic Compound
- Pollutant
- Polycyclic Aromatic Hydrocarbon
|
|---|
| Chemical Structure | |
|---|
| Synonyms | | Synonym | | 9,10-Dehydrophenanthrene | | Phenanthracene | | Phenanthrin | | Phenantrin | | Ravatite |
|
|---|
| Chemical Formula | C14H10 |
|---|
| Average Molecular Mass | 178.229 g/mol |
|---|
| Monoisotopic Mass | 178.078 g/mol |
|---|
| CAS Registry Number | 85-01-8 |
|---|
| IUPAC Name | phenanthrene |
|---|
| Traditional Name | phenanthrene |
|---|
| SMILES | C1=CC=C2C(C=CC3=CC=CC=C23)=C1 |
|---|
| InChI Identifier | InChI=1S/C14H10/c1-3-7-13-11(5-1)9-10-12-6-2-4-8-14(12)13/h1-10H |
|---|
| InChI Key | InChIKey=YNPNZTXNASCQKK-UHFFFAOYSA-N |
|---|
| Chemical Taxonomy |
|---|
| Description | belongs to the class of organic compounds known as phenanthrenes and derivatives. These are polycyclic compounds containing a phenanthrene moiety, which is a tricyclic aromatic compound with three non-linearly fused benzene. |
|---|
| Kingdom | Organic compounds |
|---|
| Super Class | Benzenoids |
|---|
| Class | Phenanthrenes and derivatives |
|---|
| Sub Class | Not Available |
|---|
| Direct Parent | Phenanthrenes and derivatives |
|---|
| Alternative Parents | |
|---|
| Substituents | - Phenanthrene
- Naphthalene
- Aromatic hydrocarbon
- Polycyclic hydrocarbon
- Unsaturated hydrocarbon
- Hydrocarbon
- Aromatic homopolycyclic compound
|
|---|
| Molecular Framework | Aromatic homopolycyclic compounds |
|---|
| External Descriptors | |
|---|
| Biological Properties |
|---|
| Status | Detected and Not Quantified |
|---|
| Origin | Exogenous |
|---|
| Cellular Locations | |
|---|
| Biofluid Locations | Not Available |
|---|
| Tissue Locations | Not Available |
|---|
| Pathways | Not Available |
|---|
| Applications | Not Available |
|---|
| Biological Roles | Not Available |
|---|
| Chemical Roles | |
|---|
| Physical Properties |
|---|
| State | Solid |
|---|
| Appearance | Colorless solid. |
|---|
| Experimental Properties | | Property | Value |
|---|
| Melting Point | 99.2°C | | Boiling Point | Not Available | | Solubility | 0.00115 mg/mL at 25 °C [SCHWARZ,FP (1977)] | | LogP | Not Available |
|
|---|
| Predicted Properties | |
|---|
| Spectra |
|---|
| Spectra | | Spectrum Type | Description | Splash Key | Deposition Date | View |
|---|
| Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | splash10-004i-0900000000-73152ac8902a46a51b0e | 2021-09-24 | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | Not Available | 2021-10-12 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-004i-0900000000-9b7fcb41bcf755cbca82 | 2016-06-03 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-004i-0900000000-e07754c44a96c14bf7b0 | 2016-06-03 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-0fb9-0900000000-fcb3480d5ef5e6c5c5c9 | 2016-06-03 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-004i-0900000000-309955037ba0840b44e2 | 2016-08-03 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-004i-0900000000-309955037ba0840b44e2 | 2016-08-03 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-004i-0900000000-de91208ace0944a1f046 | 2016-08-03 | View Spectrum | | MS | Mass Spectrum (Electron Ionization) | splash10-004i-3900000000-2c8c905707eff1036da1 | 2014-09-20 | View Spectrum | | 1D NMR | 1H NMR Spectrum (1D, 400 MHz, CDCl3, experimental) | Not Available | 2014-09-20 | View Spectrum | | 1D NMR | 13C NMR Spectrum (1D, 25.16 MHz, CDCl3, experimental) | Not Available | 2014-09-23 | View Spectrum |
|
|---|
| Toxicity Profile |
|---|
| Route of Exposure | Oral (4) ; inhalation (4) |
|---|
| Mechanism of Toxicity | The ability of PAH's to bind to blood proteins such as albumin allows them to be transported throughout the body. Many PAH's induce the expression of cytochrome P450 enzymes, especially CYP1A1, CYP1A2, and CYP1B1, by binding to the aryl hydrocarbon receptor or glycine N-methyltransferase protein. These enzymes metabolize PAH's into their toxic intermediates. The reactive metabolites of PAHs (epoxide intermediates, dihydrodiols, phenols, quinones, and their various combinations) covalently bind to DNA and other cellular macromolecules, initiating mutagenesis and carcinogenesis. (4, 5, 2, 3) |
|---|
| Metabolism | PAH metabolism occurs in all tissues, usually by cytochrome P-450 and its associated enzymes. PAHs are metabolized into reactive intermediates, which include epoxide intermediates, dihydrodiols, phenols, quinones, and their various combinations. The phenols, quinones, and dihydrodiols can all be conjugated to glucuronides and sulfate esters; the quinones also form glutathione conjugates. (4) |
|---|
| Toxicity Values | LD50: 700 mg/kg (Oral, Mouse) (7)
LD50: 700 mg/kg (Intraperitoneal, Mouse) (7)
LD50: 56 mg/kg (Intravenous, Mouse) (7) |
|---|
| Lethal Dose | Not Available |
|---|
| Carcinogenicity (IARC Classification) | 3, not classifiable as to its carcinogenicity to humans. (6) |
|---|
| Uses/Sources | PAHs are released into the environment via the combustion of fossil fuels, coke oven emissions and vehicle exhausts, as well as naturally from forest fires and volcanic eruptions. PAHs from these sources may contaminate nearly water systems. They are also found in coal tar and charbroiled food. (4) |
|---|
| Minimum Risk Level | Not Available |
|---|
| Health Effects | PAHs are carcinogens and have been associated with the increased risk of skin, respiratory tract, bladder, stomach, and kidney cancers. They may also cause reproductive effects and depress the immune system. (4) |
|---|
| Symptoms | Acute exposure to PAHs causes irritation and inflammation of the skin and lung tissue. (1) |
|---|
| Treatment | There is no known antidote for PAHs. Exposure is usually handled with symptomatic treatment. (4) |
|---|
| Normal Concentrations |
|---|
| Not Available |
|---|
| Abnormal Concentrations |
|---|
| Not Available |
|---|
| External Links |
|---|
| DrugBank ID | DB08381 |
|---|
| HMDB ID | Not Available |
|---|
| PubChem Compound ID | 995 |
|---|
| ChEMBL ID | CHEMBL46730 |
|---|
| ChemSpider ID | 970 |
|---|
| KEGG ID | C11422 |
|---|
| UniProt ID | Not Available |
|---|
| OMIM ID | |
|---|
| ChEBI ID | 28851 |
|---|
| BioCyc ID | CPD-1904 |
|---|
| CTD ID | C031181 |
|---|
| Stitch ID | Phenanthrene |
|---|
| PDB ID | Not Available |
|---|
| ACToR ID | 6508 |
|---|
| Wikipedia Link | Phenanthrene |
|---|
| References |
|---|
| Synthesis Reference | Kurt Handrick, Georg Kolling, Fritz Mensch, “Anthracene production from phenanthrene.” U.S. Patent US4384152, issued July, 1953. |
|---|
| MSDS | T3D0246.pdf |
|---|
| General References | - Santodonato J, Howard P, Basu D: Health and ecological assessment of polynuclear aromatic hydrocarbons. J Environ Pathol Toxicol. 1981 Sep;5(1):1-364. [7310260 ]
- Uno S, Dragin N, Miller ML, Dalton TP, Gonzalez FJ, Nebert DW: Basal and inducible CYP1 mRNA quantitation and protein localization throughout the mouse gastrointestinal tract. Free Radic Biol Med. 2008 Feb 15;44(4):570-83. Epub 2007 Nov 12. [17997381 ]
- Padros J, Pelletier E: In vivo formation of (+)-anti-benzo[a]pyrene diol-epoxide-plasma albumin adducts in fish. Mar Environ Res. 2000 Jul-Dec;50(1-5):347-51. [11460716 ]
- ATSDR - Agency for Toxic Substances and Disease Registry (1995). Toxicological profile for PAHs. U.S. Public Health Service in collaboration with U.S. Environmental Protection Agency (EPA). [Link]
- Wikipedia. Benzopyrene. Last Updated 22 January 2009. [Link]
- International Agency for Research on Cancer (2014). IARC Monographs on the Evaluation of Carcinogenic Risks to Humans. [Link]
- The Physical and Theoretical Chemistry Laboratory of Oxford University (2004). Material Safety Data Sheet (MSDS) for phenanthrene. [Link]
|
|---|
| Gene Regulation |
|---|
| Up-Regulated Genes | | Gene | Gene Symbol | Gene ID | Interaction | Chromosome | Details |
|---|
|
|---|
| Down-Regulated Genes | | Gene | Gene Symbol | Gene ID | Interaction | Chromosome | Details |
|---|
|
|---|