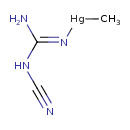
Methylmercuric dicyanamide (T3D1350)
| Record Information | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Version | 2.0 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Creation Date | 2009-06-19 21:58:39 UTC | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Update Date | 2014-12-24 20:23:44 UTC | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Accession Number | T3D1350 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Identification | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Common Name | Methylmercuric dicyanamide | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Class | Small Molecule | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Description | Methylmercuric dicyanamide is an organomercuric compound of mercury and cyanide. Mercury is a heavy, silvery d-block metal and one of six elements that are liquid at or near room temperature and pressure. It is a naturally occuring substance, and combines with other elements such as chlorine, sulfur, or oxygen to form inorganic mercury compounds (salts). Mercury also combines with carbon to make organic mercury compounds. (8) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Compound Type |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Chemical Structure | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Synonyms |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Chemical Formula | C3H6HgN4 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Average Molecular Mass | 298.700 g/mol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Monoisotopic Mass | 300.030 g/mol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| CAS Registry Number | 502-39-6 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| IUPAC Name | (E)-1-cyano-2-(methylmercurio)guanidine | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Traditional Name | (E)-1-cyano-2-(methylmercurio)guanidine | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| SMILES | C[Hg]\N=C(/N)NC#N | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| InChI Identifier | InChI=1S/C2H3N4.CH3.Hg/c3-1-6-2(4)5;;/h(H3-,4,5,6);1H3;/q-1;;+1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| InChI Key | InChIKey=JVJUWCMBRUMDDQ-UHFFFAOYSA-N | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Chemical Taxonomy | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Description | belongs to the class of organic compounds known as carbene-type 1,3-dipolar compounds. These are 1,3-dipolar compounds with the general structure X:-C=Z<-> X+=C-Z- (X = C or N; Z = C, N, or O). | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Kingdom | Organic compounds | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Super Class | Organic 1,3-dipolar compounds | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Class | Carbene-type 1,3-dipolar compounds | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Sub Class | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Direct Parent | Carbene-type 1,3-dipolar compounds | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Alternative Parents | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Substituents |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Molecular Framework | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| External Descriptors | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Biological Properties | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Status | Detected and Not Quantified | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Origin | Exogenous | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Cellular Locations |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Biofluid Locations | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Tissue Locations | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Pathways | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Applications | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Biological Roles | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Chemical Roles | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Physical Properties | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| State | Solid | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Appearance | Colorless crystals. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Experimental Properties |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Predicted Properties |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Spectra | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Spectra |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Toxicity Profile | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Route of Exposure | Oral (9) ; inhalation (9) ; dermal (9) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Mechanism of Toxicity | High-affinity binding of the divalent mercuric ion to thiol or sulfhydryl groups of proteins is believed to be the major mechanism for the activity of mercury. Through alterations in intracellular thiol status, mercury can promote oxidative stress, lipid peroxidation, mitochondrial dysfunction, and changes in heme metabolism. Mercury is known to bind to microsomal and mitochondrial enzymes, resulting in cell injury and death. For example, mercury is known to inhibit aquaporins, halting water flow across the cell membrane. It also inhibits the protein LCK, which causes decreased T-cell signalling and immune system depression. Mercury is also believed to inhibit neuronal excitability by acting on the postsynaptic neuronal membrane. It also affects the nervous system by inhibiting protein kinase C and alkaline phosphatase, which impairs brain microvascular formation and function, as well as alters the blood-brain barrier. Mercury also produces an autoimmune response, likely by modification of major histocompatibility complex (MHC) class II molecules, self peptides, T-cell receptors, or cell-surface adhesion molecules. Organic nitriles decompose into cyanide ions both in vivo and in vitro. Consequently the primary mechanism of toxicity for organic nitriles is their production of toxic cyanide ions or hydrogen cyanide. Cyanide is an inhibitor of cytochrome c oxidase in the fourth complex of the electron transport chain (found in the membrane of the mitochondria of eukaryotic cells). It complexes with the ferric iron atom in this enzyme. The binding of cyanide to this cytochrome prevents transport of electrons from cytochrome c oxidase to oxygen. As a result, the electron transport chain is disrupted and the cell can no longer aerobically produce ATP for energy. Tissues that mainly depend on aerobic respiration, such as the central nervous system and the heart, are particularly affected. Cyanide is also known produce some of its toxic effects by binding to catalase, glutathione peroxidase, methemoglobin, hydroxocobalamin, phosphatase, tyrosinase, ascorbic acid oxidase, xanthine oxidase, succinic dehydrogenase, and Cu/Zn superoxide dismutase. Cyanide binds to the ferric ion of methemoglobin to form inactive cyanmethemoglobin. (9, 4, 5, 6, 12) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Metabolism | Mercury is absorbed mainly via ingestion and inhalation, then distributed throughout the body via the bloodstream, where a portion binds to sulfhydryl groups on haemoglobin. Mercury can undergo oxidation to mercuric mercury, which takes place via the catalase-hydrogen peroxide pathway. The mercury atom is able to diffuse down the cleft in the catalase enzyme to reach the active site where the heme ring is located. Oxidation most likely occurs in all tissue, as the catalase hydrogen peroxide pathway is ubiquitous. Following oxidation, mercury tends to accumulate in the kidneys. Mercury is excreted mainly by exhalation and in the faeces. Organic nitriles are converted into cyanide ions through the action of cytochrome P450 enzymes in the liver. Cyanide is rapidly absorbed and distributed throughout the body. Cyanide is mainly metabolized into thiocyanate by either rhodanese or 3-mercaptopyruvate sulfur transferase. Cyanide metabolites are excreted in the urine. (2, 9, 11) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Toxicity Values | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Lethal Dose | 200 to 300 milligrams for an adult human (cyanide salts). (7) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Carcinogenicity (IARC Classification) | 2B, possibly carcinogenic to humans. (14) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Uses/Sources | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Minimum Risk Level | Chronic Inhalation: 0.0002 mg/m3 (Mercury) (13) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Health Effects | Mercury mainly affects the nervous system. Exposure to high levels of metallic, inorganic, or organic mercury can permanently damage the brain, kidneys, and developing fetus. Effects on brain functioning may result in irritability, shyness, tremors, changes in vision or hearing, and memory problems. Acrodynia, a type of mercury poisoning in children, is characterized by pain and pink discoloration of the hands and feet. Mercury poisoning can also cause Hunter-Russell syndrome and Minamata disease. Exposure to high levels of cyanide for a short time harms the brain and heart and can even cause coma, seizures, apnea, cardiac arrest and death. Chronic inhalation of cyanide causes breathing difficulties, chest pain, vomiting, blood changes, headaches, and enlargement of the thyroid gland. Skin contact with cyanide salts can irritate and produce sores. (9, 11, 12) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symptoms | Common symptoms include peripheral neuropathy (presenting as paresthesia or itching, burning or pain), skin discoloration (pink cheeks, fingertips and toes), edema (swelling), and desquamation (dead skin peels off in layers). Cyanide poisoning is identified by rapid, deep breathing and shortness of breath, general weakness, giddiness, headaches, vertigo, confusion, convulsions/seizures and eventually loss of consciousness. (1, 11, 12) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Treatment | Mercury poisoning is treated by immediate decontamination and chelation therapy using DMSA, DMPS, DPCN, or dimercaprol. Antidotes to cyanide poisoning include hydroxocobalamin and sodium nitrite, which release the cyanide from the cytochrome system, and rhodanase, which is an enzyme occurring naturally in mammals that combines serum cyanide with thiosulfate, producing comparatively harmless thiocyanate. Oxygen therapy can also be administered. (3, 12) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Normal Concentrations | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Not Available | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Abnormal Concentrations | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Not Available | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| External Links | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| DrugBank ID | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| HMDB ID | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PubChem Compound ID | 16682942 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| ChEMBL ID | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| ChemSpider ID | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| KEGG ID | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| UniProt ID | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| OMIM ID | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| ChEBI ID | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| BioCyc ID | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| CTD ID | C005232 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Stitch ID | Methylmercuric dicyanamide | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PDB ID | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| ACToR ID | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Wikipedia Link | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| References | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Synthesis Reference | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| MSDS | T3D1350.pdf | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| General References |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Gene Regulation | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Up-Regulated Genes | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Down-Regulated Genes | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Targets
- General Function:
- Metal ion binding
- Specific Function:
- Not Available
- Gene Name:
- ALPPL2
- Uniprot ID:
- P10696
- Molecular Weight:
- 57376.515 Da
References
- Gerbitz KD: Human alkaline phosphatases. II. Metalloenzyme properties of the enzyme from human liver. Hoppe Seylers Z Physiol Chem. 1977 Nov;358(11):1491-7. [924371 ]
- General Function:
- Pyrophosphatase activity
- Specific Function:
- This isozyme may play a role in skeletal mineralization.
- Gene Name:
- ALPL
- Uniprot ID:
- P05186
- Molecular Weight:
- 57304.435 Da
References
- ATSDR - Agency for Toxic Substances and Disease Registry (2008). Toxicological profile for mercury. U.S. Public Health Service in collaboration with U.S. Environmental Protection Agency (EPA). [Link]
- General Function:
- Water transmembrane transporter activity
- Specific Function:
- Forms a water-specific channel that provides the plasma membranes of red cells and kidney proximal tubules with high permeability to water, thereby permitting water to move in the direction of an osmotic gradient.
- Gene Name:
- AQP1
- Uniprot ID:
- P29972
- Molecular Weight:
- 28525.68 Da
References
- Yukutake Y, Tsuji S, Hirano Y, Adachi T, Takahashi T, Fujihara K, Agre P, Yasui M, Suematsu M: Mercury chloride decreases the water permeability of aquaporin-4-reconstituted proteoliposomes. Biol Cell. 2008 Jun;100(6):355-63. doi: 10.1042/BC20070132. [18167118 ]
- General Function:
- Water channel activity
- Specific Function:
- Water channel required to promote glycerol permeability and water transport across cell membranes. May contribute to water transport in the upper portion of small intestine. Isoform 2 is not permeable to urea and glycerol.
- Gene Name:
- AQP10
- Uniprot ID:
- Q96PS8
- Molecular Weight:
- 31762.97 Da
References
- Yukutake Y, Tsuji S, Hirano Y, Adachi T, Takahashi T, Fujihara K, Agre P, Yasui M, Suematsu M: Mercury chloride decreases the water permeability of aquaporin-4-reconstituted proteoliposomes. Biol Cell. 2008 Jun;100(6):355-63. doi: 10.1042/BC20070132. [18167118 ]
- General Function:
- Transporter activity
- Specific Function:
- Aquaporins facilitate the transport of water and small neutral solutes across cell membranes.
- Gene Name:
- AQP11
- Uniprot ID:
- Q8NBQ7
- Molecular Weight:
- 30202.59 Da
References
- Yukutake Y, Tsuji S, Hirano Y, Adachi T, Takahashi T, Fujihara K, Agre P, Yasui M, Suematsu M: Mercury chloride decreases the water permeability of aquaporin-4-reconstituted proteoliposomes. Biol Cell. 2008 Jun;100(6):355-63. doi: 10.1042/BC20070132. [18167118 ]
- General Function:
- Transporter activity
- Specific Function:
- Aquaporins facilitate the transport of water and small neutral solutes across cell membranes.
- Gene Name:
- AQP12A
- Uniprot ID:
- Q8IXF9
- Molecular Weight:
- 31474.21 Da
References
- Yukutake Y, Tsuji S, Hirano Y, Adachi T, Takahashi T, Fujihara K, Agre P, Yasui M, Suematsu M: Mercury chloride decreases the water permeability of aquaporin-4-reconstituted proteoliposomes. Biol Cell. 2008 Jun;100(6):355-63. doi: 10.1042/BC20070132. [18167118 ]
- General Function:
- Transporter activity
- Specific Function:
- Aquaporins facilitate the transport of water and small neutral solutes across cell membranes.
- Gene Name:
- AQP12B
- Uniprot ID:
- A6NM10
- Molecular Weight:
- 31475.13 Da
References
- Yukutake Y, Tsuji S, Hirano Y, Adachi T, Takahashi T, Fujihara K, Agre P, Yasui M, Suematsu M: Mercury chloride decreases the water permeability of aquaporin-4-reconstituted proteoliposomes. Biol Cell. 2008 Jun;100(6):355-63. doi: 10.1042/BC20070132. [18167118 ]
- General Function:
- Water transmembrane transporter activity
- Specific Function:
- Forms a water-specific channel that provides the plasma membranes of renal collecting duct with high permeability to water, thereby permitting water to move in the direction of an osmotic gradient.
- Gene Name:
- AQP2
- Uniprot ID:
- P41181
- Molecular Weight:
- 28837.17 Da
References
- Yukutake Y, Tsuji S, Hirano Y, Adachi T, Takahashi T, Fujihara K, Agre P, Yasui M, Suematsu M: Mercury chloride decreases the water permeability of aquaporin-4-reconstituted proteoliposomes. Biol Cell. 2008 Jun;100(6):355-63. doi: 10.1042/BC20070132. [18167118 ]
- General Function:
- Water channel activity
- Specific Function:
- Water channel required to promote glycerol permeability and water transport across cell membranes. Acts as a glycerol transporter in skin and plays an important role in regulating SC (stratum corneum) and epidermal glycerol content. Involved in skin hydration, wound healing, and tumorigenesis. Provides kidney medullary collecting duct with high permeability to water, thereby permitting water to move in the direction of an osmotic gradient. Slightly permeable to urea and may function as a water and urea exit mechanism in antidiuresis in collecting duct cells. It may play an important role in gastrointestinal tract water transport and in glycerol metabolism (By similarity).
- Gene Name:
- AQP3
- Uniprot ID:
- Q92482
- Molecular Weight:
- 31543.605 Da
References
- Yukutake Y, Tsuji S, Hirano Y, Adachi T, Takahashi T, Fujihara K, Agre P, Yasui M, Suematsu M: Mercury chloride decreases the water permeability of aquaporin-4-reconstituted proteoliposomes. Biol Cell. 2008 Jun;100(6):355-63. doi: 10.1042/BC20070132. [18167118 ]
- General Function:
- Water transmembrane transporter activity
- Specific Function:
- Forms a water-specific channel. Osmoreceptor which regulates body water balance and mediates water flow within the central nervous system.
- Gene Name:
- AQP4
- Uniprot ID:
- P55087
- Molecular Weight:
- 34829.43 Da
References
- Yukutake Y, Tsuji S, Hirano Y, Adachi T, Takahashi T, Fujihara K, Agre P, Yasui M, Suematsu M: Mercury chloride decreases the water permeability of aquaporin-4-reconstituted proteoliposomes. Biol Cell. 2008 Jun;100(6):355-63. doi: 10.1042/BC20070132. [18167118 ]
- General Function:
- Water channel activity
- Specific Function:
- Forms a water-specific channel. Implicated in the generation of saliva, tears, and pulmonary secretions. Required for TRPV4 activation by hypotonicity (PubMed:16571723). Together with TRPV4, controls regulatory volume decrease in salivary epithelial cells (PubMed:16571723).
- Gene Name:
- AQP5
- Uniprot ID:
- P55064
- Molecular Weight:
- 28291.89 Da
References
- Yukutake Y, Tsuji S, Hirano Y, Adachi T, Takahashi T, Fujihara K, Agre P, Yasui M, Suematsu M: Mercury chloride decreases the water permeability of aquaporin-4-reconstituted proteoliposomes. Biol Cell. 2008 Jun;100(6):355-63. doi: 10.1042/BC20070132. [18167118 ]
- General Function:
- Water channel activity
- Specific Function:
- Forms a water-specific channel that participates in distinct physiological functions such as glomerular filtration, tubular endocytosis and acid-base metabolism.
- Gene Name:
- AQP6
- Uniprot ID:
- Q13520
- Molecular Weight:
- 29370.215 Da
References
- Yukutake Y, Tsuji S, Hirano Y, Adachi T, Takahashi T, Fujihara K, Agre P, Yasui M, Suematsu M: Mercury chloride decreases the water permeability of aquaporin-4-reconstituted proteoliposomes. Biol Cell. 2008 Jun;100(6):355-63. doi: 10.1042/BC20070132. [18167118 ]
- General Function:
- Water channel activity
- Specific Function:
- Forms a channel for water and glycerol.
- Gene Name:
- AQP7
- Uniprot ID:
- O14520
- Molecular Weight:
- 37231.325 Da
References
- Yukutake Y, Tsuji S, Hirano Y, Adachi T, Takahashi T, Fujihara K, Agre P, Yasui M, Suematsu M: Mercury chloride decreases the water permeability of aquaporin-4-reconstituted proteoliposomes. Biol Cell. 2008 Jun;100(6):355-63. doi: 10.1042/BC20070132. [18167118 ]
- General Function:
- Water channel activity
- Specific Function:
- Forms a water-specific channel; mercury-sensitive. Not permeable to glycerol or urea.
- Gene Name:
- AQP8
- Uniprot ID:
- O94778
- Molecular Weight:
- 27381.01 Da
References
- Yukutake Y, Tsuji S, Hirano Y, Adachi T, Takahashi T, Fujihara K, Agre P, Yasui M, Suematsu M: Mercury chloride decreases the water permeability of aquaporin-4-reconstituted proteoliposomes. Biol Cell. 2008 Jun;100(6):355-63. doi: 10.1042/BC20070132. [18167118 ]
- General Function:
- Water transmembrane transporter activity
- Specific Function:
- Forms a channel with a broad specificity. Mediates passage of a wide variety of non-charged solutes including carbamides, polyols, purines, and pyrimidines in a phloretin- and mercury-sensitive manner, whereas amino acids, cyclic sugars, Na(+), K(+), Cl(-), and deprotonated monocarboxylates are excluded. Also permeable to urea and glycerol.
- Gene Name:
- AQP9
- Uniprot ID:
- O43315
- Molecular Weight:
- 31430.77 Da
References
- Yukutake Y, Tsuji S, Hirano Y, Adachi T, Takahashi T, Fujihara K, Agre P, Yasui M, Suematsu M: Mercury chloride decreases the water permeability of aquaporin-4-reconstituted proteoliposomes. Biol Cell. 2008 Jun;100(6):355-63. doi: 10.1042/BC20070132. [18167118 ]
- General Function:
- Receptor binding
- Specific Function:
- Occurs in almost all aerobically respiring organisms and serves to protect cells from the toxic effects of hydrogen peroxide. Promotes growth of cells including T-cells, B-cells, myeloid leukemia cells, melanoma cells, mastocytoma cells and normal and transformed fibroblast cells.
- Gene Name:
- CAT
- Uniprot ID:
- P04040
- Molecular Weight:
- 59755.82 Da
References
- Kang YS, Lee DH, Yoon BJ, Oh DC: Purification and characterization of a catalase from photosynthetic bacterium Rhodospirillum rubrum S1 grown under anaerobic conditions. J Microbiol. 2006 Apr;44(2):185-91. [16728955 ]
- General Function:
- Iron ion binding
- Specific Function:
- Cytochrome c oxidase is the component of the respiratory chain that catalyzes the reduction of oxygen to water. Subunits 1-3 form the functional core of the enzyme complex. CO I is the catalytic subunit of the enzyme. Electrons originating in cytochrome c are transferred via the copper A center of subunit 2 and heme A of subunit 1 to the bimetallic center formed by heme A3 and copper B.
- Gene Name:
- MT-CO1
- Uniprot ID:
- P00395
- Molecular Weight:
- 57040.91 Da
References
- Wikipedia. Cyanide poisoning. Last Updated 30 March 2009. [Link]
- General Function:
- Cytochrome-c oxidase activity
- Specific Function:
- Cytochrome c oxidase is the component of the respiratory chain that catalyzes the reduction of oxygen to water. Subunits 1-3 form the functional core of the enzyme complex. Subunit 2 transfers the electrons from cytochrome c via its binuclear copper A center to the bimetallic center of the catalytic subunit 1.
- Gene Name:
- MT-CO2
- Uniprot ID:
- P00403
- Molecular Weight:
- 25564.73 Da
References
- Wikipedia. Cyanide poisoning. Last Updated 30 March 2009. [Link]
- General Function:
- Cytochrome-c oxidase activity
- Specific Function:
- This protein is one of the nuclear-coded polypeptide chains of cytochrome c oxidase, the terminal oxidase in mitochondrial electron transport.
- Gene Name:
- COX4I1
- Uniprot ID:
- P13073
- Molecular Weight:
- 19576.6 Da
References
- Wikipedia. Cyanide poisoning. Last Updated 30 March 2009. [Link]
- General Function:
- Cytochrome-c oxidase activity
- Specific Function:
- This protein is one of the nuclear-coded polypeptide chains of cytochrome c oxidase, the terminal oxidase in mitochondrial electron transport.
- Gene Name:
- COX4I2
- Uniprot ID:
- Q96KJ9
- Molecular Weight:
- 20010.02 Da
References
- Wikipedia. Cyanide poisoning. Last Updated 30 March 2009. [Link]
- General Function:
- Metal ion binding
- Specific Function:
- This is the heme A-containing chain of cytochrome c oxidase, the terminal oxidase in mitochondrial electron transport.
- Gene Name:
- COX5A
- Uniprot ID:
- P20674
- Molecular Weight:
- 16761.985 Da
References
- Wikipedia. Cyanide poisoning. Last Updated 30 March 2009. [Link]
- General Function:
- Metal ion binding
- Specific Function:
- This protein is one of the nuclear-coded polypeptide chains of cytochrome c oxidase, the terminal oxidase in mitochondrial electron transport.
- Gene Name:
- COX5B
- Uniprot ID:
- P10606
- Molecular Weight:
- 13695.57 Da
References
- Wikipedia. Cyanide poisoning. Last Updated 30 March 2009. [Link]
- General Function:
- Cytochrome-c oxidase activity
- Specific Function:
- This protein is one of the nuclear-coded polypeptide chains of cytochrome c oxidase, the terminal oxidase in mitochondrial electron transport.
- Gene Name:
- COX6A1
- Uniprot ID:
- P12074
- Molecular Weight:
- 12154.8 Da
References
- Wikipedia. Cyanide poisoning. Last Updated 30 March 2009. [Link]
- General Function:
- Cytochrome-c oxidase activity
- Specific Function:
- This protein is one of the nuclear-coded polypeptide chains of cytochrome c oxidase, the terminal oxidase in mitochondrial electron transport.
- Gene Name:
- COX6A2
- Uniprot ID:
- Q02221
- Molecular Weight:
- 10815.32 Da
References
- Wikipedia. Cyanide poisoning. Last Updated 30 March 2009. [Link]
- General Function:
- Cytochrome-c oxidase activity
- Specific Function:
- This protein is one of the nuclear-coded polypeptide chains of cytochrome c oxidase, the terminal oxidase in mitochondrial electron transport.
- Gene Name:
- COX6C
- Uniprot ID:
- P09669
- Molecular Weight:
- 8781.36 Da
References
- Wikipedia. Cyanide poisoning. Last Updated 30 March 2009. [Link]
- General Function:
- Cytochrome-c oxidase activity
- Specific Function:
- This protein is one of the nuclear-coded polypeptide chains of cytochrome c oxidase, the terminal oxidase in mitochondrial electron transport.
- Gene Name:
- COX7A1
- Uniprot ID:
- P24310
- Molecular Weight:
- 9117.44 Da
References
- Wikipedia. Cyanide poisoning. Last Updated 30 March 2009. [Link]
- General Function:
- Cytochrome-c oxidase activity
- Specific Function:
- This protein is one of the nuclear-coded polypeptide chains of cytochrome c oxidase, the terminal oxidase in mitochondrial electron transport.
- Gene Name:
- COX7A2
- Uniprot ID:
- P14406
- Molecular Weight:
- 9395.89 Da
References
- Wikipedia. Cyanide poisoning. Last Updated 30 March 2009. [Link]
- General Function:
- Cytochrome-c oxidase activity
- Specific Function:
- This protein is one of the nuclear-coded polypeptide chains of cytochrome c oxidase, the terminal oxidase in mitochondrial electron transport. Plays a role in proper central nervous system (CNS) development in vertebrates.
- Gene Name:
- COX7B
- Uniprot ID:
- P24311
- Molecular Weight:
- 9160.485 Da
References
- Wikipedia. Cyanide poisoning. Last Updated 30 March 2009. [Link]
- General Function:
- Cytochrome-c oxidase activity
- Specific Function:
- This protein is one of the nuclear-coded polypeptide chains of cytochrome c oxidase, the terminal oxidase in mitochondrial electron transport.
- Gene Name:
- COX7B2
- Uniprot ID:
- Q8TF08
- Molecular Weight:
- 9077.43 Da
References
- Wikipedia. Cyanide poisoning. Last Updated 30 March 2009. [Link]
- General Function:
- Cytochrome-c oxidase activity
- Specific Function:
- This protein is one of the nuclear-coded polypeptide chains of cytochrome c oxidase, the terminal oxidase in mitochondrial electron transport.
- Gene Name:
- COX7C
- Uniprot ID:
- P15954
- Molecular Weight:
- 7245.45 Da
References
- Wikipedia. Cyanide poisoning. Last Updated 30 March 2009. [Link]
- General Function:
- Cytochrome-c oxidase activity
- Specific Function:
- This protein is one of the nuclear-coded polypeptide chains of cytochrome c oxidase, the terminal oxidase in mitochondrial electron transport.
- Gene Name:
- COX8A
- Uniprot ID:
- P10176
- Molecular Weight:
- 7579.0 Da
References
- Wikipedia. Cyanide poisoning. Last Updated 30 March 2009. [Link]
- General Function:
- Cytochrome-c oxidase activity
- Specific Function:
- This protein is one of the nuclear-coded polypeptide chains of cytochrome c oxidase, the terminal oxidase in mitochondrial electron transport.
- Gene Name:
- COX8C
- Uniprot ID:
- Q7Z4L0
- Molecular Weight:
- 8128.575 Da
References
- Wikipedia. Cyanide poisoning. Last Updated 30 March 2009. [Link]
- General Function:
- Glutathione peroxidase activity
- Specific Function:
- Protects cells and enzymes from oxidative damage, by catalyzing the reduction of hydrogen peroxide, lipid peroxides and organic hydroperoxide, by glutathione. May constitute a glutathione peroxidase-like protective system against peroxide damage in sperm membrane lipids.
- Gene Name:
- GPX5
- Uniprot ID:
- O75715
- Molecular Weight:
- 25202.14 Da
References
- Kraus RJ, Ganther HE: Reaction of cyanide with glutathione peroxidase. Biochem Biophys Res Commun. 1980 Oct 16;96(3):1116-22. [7437059 ]
- General Function:
- Zinc ion binding
- Specific Function:
- Protect the extracellular space from toxic effect of reactive oxygen intermediates by converting superoxide radicals into hydrogen peroxide and oxygen.
- Gene Name:
- SOD3
- Uniprot ID:
- P08294
- Molecular Weight:
- 25850.675 Da
References
- Lee WG, Hwang JH, Na BK, Cho JH, Lee HW, Cho SH, Kong Y, Song CY, Kim TS: Functional expression of a recombinant copper/zinc superoxide dismutase of filarial nematode, Brugia malayi. J Parasitol. 2005 Feb;91(1):205-8. [15856906 ]
- General Function:
- Sh3 domain binding
- Specific Function:
- Protects the hemoglobin in erythrocytes from oxidative breakdown.
- Gene Name:
- GPX1
- Uniprot ID:
- P07203
- Molecular Weight:
- 22087.94 Da
References
- Kraus RJ, Ganther HE: Reaction of cyanide with glutathione peroxidase. Biochem Biophys Res Commun. 1980 Oct 16;96(3):1116-22. [7437059 ]
- General Function:
- Glutathione peroxidase activity
- Specific Function:
- Could play a major role in protecting mammals from the toxicity of ingested organic hydroperoxides. Tert-butyl hydroperoxide, cumene hydroperoxide and linoleic acid hydroperoxide but not phosphatidycholine hydroperoxide, can act as acceptors.
- Gene Name:
- GPX2
- Uniprot ID:
- P18283
- Molecular Weight:
- 21953.835 Da
References
- Kraus RJ, Ganther HE: Reaction of cyanide with glutathione peroxidase. Biochem Biophys Res Commun. 1980 Oct 16;96(3):1116-22. [7437059 ]
- General Function:
- Transcription factor binding
- Specific Function:
- Protects cells and enzymes from oxidative damage, by catalyzing the reduction of hydrogen peroxide, lipid peroxides and organic hydroperoxide, by glutathione.
- Gene Name:
- GPX3
- Uniprot ID:
- P22352
- Molecular Weight:
- 25552.185 Da
References
- Kraus RJ, Ganther HE: Reaction of cyanide with glutathione peroxidase. Biochem Biophys Res Commun. 1980 Oct 16;96(3):1116-22. [7437059 ]
- General Function:
- Peroxidase activity
- Specific Function:
- It protects esophageal epithelia from hydrogen peroxide-induced oxidative stress. It suppresses acidic bile acid-induced reactive oxigen species (ROS) and protects against oxidative DNA damage and double-strand breaks.
- Gene Name:
- GPX7
- Uniprot ID:
- Q96SL4
- Molecular Weight:
- 20995.88 Da
References
- Kraus RJ, Ganther HE: Reaction of cyanide with glutathione peroxidase. Biochem Biophys Res Commun. 1980 Oct 16;96(3):1116-22. [7437059 ]
- General Function:
- Nadp binding
- Specific Function:
- Maintains high levels of reduced glutathione in the cytosol.
- Gene Name:
- GSR
- Uniprot ID:
- P00390
- Molecular Weight:
- 56256.565 Da
References
- Ardelt BK, Borowitz JL, Isom GE: Brain lipid peroxidation and antioxidant protectant mechanisms following acute cyanide intoxication. Toxicology. 1989 Jun 1;56(2):147-54. [2734799 ]
- General Function:
- Water channel activity
- Specific Function:
- Water channel. Channel activity is down-regulated by CALM when cytoplasmic Ca(2+) levels are increased. May be responsible for regulating the osmolarity of the lens. Interactions between homotetramers from adjoining membranes may stabilize cell junctions in the eye lens core (By similarity).
- Gene Name:
- MIP
- Uniprot ID:
- P30301
- Molecular Weight:
- 28121.5 Da
References
- Yukutake Y, Tsuji S, Hirano Y, Adachi T, Takahashi T, Fujihara K, Agre P, Yasui M, Suematsu M: Mercury chloride decreases the water permeability of aquaporin-4-reconstituted proteoliposomes. Biol Cell. 2008 Jun;100(6):355-63. doi: 10.1042/BC20070132. [18167118 ]
- General Function:
- Phospholipid-hydroperoxide glutathione peroxidase activity
- Specific Function:
- Protects cells against membrane lipid peroxidation and cell death. Required for normal sperm development and male fertility. Could play a major role in protecting mammals from the toxicity of ingested lipid hydroperoxides. Essential for embryonic development. Protects from radiation and oxidative damage.
- Gene Name:
- GPX4
- Uniprot ID:
- P36969
- Molecular Weight:
- 22174.52 Da
References
- Kraus RJ, Ganther HE: Reaction of cyanide with glutathione peroxidase. Biochem Biophys Res Commun. 1980 Oct 16;96(3):1116-22. [7437059 ]
- General Function:
- Zinc ion binding
- Specific Function:
- Calcium-activated, phospholipid- and diacylglycerol (DAG)-dependent serine/threonine-protein kinase that is involved in positive and negative regulation of cell proliferation, apoptosis, differentiation, migration and adhesion, tumorigenesis, cardiac hypertrophy, angiogenesis, platelet function and inflammation, by directly phosphorylating targets such as RAF1, BCL2, CSPG4, TNNT2/CTNT, or activating signaling cascade involving MAPK1/3 (ERK1/2) and RAP1GAP. Involved in cell proliferation and cell growth arrest by positive and negative regulation of the cell cycle. Can promote cell growth by phosphorylating and activating RAF1, which mediates the activation of the MAPK/ERK signaling cascade, and/or by up-regulating CDKN1A, which facilitates active cyclin-dependent kinase (CDK) complex formation in glioma cells. In intestinal cells stimulated by the phorbol ester PMA, can trigger a cell cycle arrest program which is associated with the accumulation of the hyper-phosphorylated growth-suppressive form of RB1 and induction of the CDK inhibitors CDKN1A and CDKN1B. Exhibits anti-apoptotic function in glioma cells and protects them from apoptosis by suppressing the p53/TP53-mediated activation of IGFBP3, and in leukemia cells mediates anti-apoptotic action by phosphorylating BCL2. During macrophage differentiation induced by macrophage colony-stimulating factor (CSF1), is translocated to the nucleus and is associated with macrophage development. After wounding, translocates from focal contacts to lamellipodia and participates in the modulation of desmosomal adhesion. Plays a role in cell motility by phosphorylating CSPG4, which induces association of CSPG4 with extensive lamellipodia at the cell periphery and polarization of the cell accompanied by increases in cell motility. Is highly expressed in a number of cancer cells where it can act as a tumor promoter and is implicated in malignant phenotypes of several tumors such as gliomas and breast cancers. Negatively regulates myocardial contractility and positively regulates angiogenesis, platelet aggregation and thrombus formation in arteries. Mediates hypertrophic growth of neonatal cardiomyocytes, in part through a MAPK1/3 (ERK1/2)-dependent signaling pathway, and upon PMA treatment, is required to induce cardiomyocyte hypertrophy up to heart failure and death, by increasing protein synthesis, protein-DNA ratio and cell surface area. Regulates cardiomyocyte function by phosphorylating cardiac troponin T (TNNT2/CTNT), which induces significant reduction in actomyosin ATPase activity, myofilament calcium sensitivity and myocardial contractility. In angiogenesis, is required for full endothelial cell migration, adhesion to vitronectin (VTN), and vascular endothelial growth factor A (VEGFA)-dependent regulation of kinase activation and vascular tube formation. Involved in the stabilization of VEGFA mRNA at post-transcriptional level and mediates VEGFA-induced cell proliferation. In the regulation of calcium-induced platelet aggregation, mediates signals from the CD36/GP4 receptor for granule release, and activates the integrin heterodimer ITGA2B-ITGB3 through the RAP1GAP pathway for adhesion. During response to lipopolysaccharides (LPS), may regulate selective LPS-induced macrophage functions involved in host defense and inflammation. But in some inflammatory responses, may negatively regulate NF-kappa-B-induced genes, through IL1A-dependent induction of NF-kappa-B inhibitor alpha (NFKBIA/IKBA). Upon stimulation with 12-O-tetradecanoylphorbol-13-acetate (TPA), phosphorylates EIF4G1, which modulates EIF4G1 binding to MKNK1 and may be involved in the regulation of EIF4E phosphorylation. Phosphorylates KIT, leading to inhibition of KIT activity. Phosphorylates ATF2 which promotes cooperation between ATF2 and JUN, activating transcription.
- Gene Name:
- PRKCA
- Uniprot ID:
- P17252
- Molecular Weight:
- 76749.445 Da
References
- ATSDR - Agency for Toxic Substances and Disease Registry (2008). Toxicological profile for mercury. U.S. Public Health Service in collaboration with U.S. Environmental Protection Agency (EPA). [Link]
- General Function:
- Zinc ion binding
- Specific Function:
- Calcium-activated, phospholipid- and diacylglycerol (DAG)-dependent serine/threonine-protein kinase involved in various cellular processes such as regulation of the B-cell receptor (BCR) signalosome, oxidative stress-induced apoptosis, androgen receptor-dependent transcription regulation, insulin signaling and endothelial cells proliferation. Plays a key role in B-cell activation by regulating BCR-induced NF-kappa-B activation. Mediates the activation of the canonical NF-kappa-B pathway (NFKB1) by direct phosphorylation of CARD11/CARMA1 at 'Ser-559', 'Ser-644' and 'Ser-652'. Phosphorylation induces CARD11/CARMA1 association with lipid rafts and recruitment of the BCL10-MALT1 complex as well as MAP3K7/TAK1, which then activates IKK complex, resulting in nuclear translocation and activation of NFKB1. Plays a direct role in the negative feedback regulation of the BCR signaling, by down-modulating BTK function via direct phosphorylation of BTK at 'Ser-180', which results in the alteration of BTK plasma membrane localization and in turn inhibition of BTK activity. Involved in apoptosis following oxidative damage: in case of oxidative conditions, specifically phosphorylates 'Ser-36' of isoform p66Shc of SHC1, leading to mitochondrial accumulation of p66Shc, where p66Shc acts as a reactive oxygen species producer. Acts as a coactivator of androgen receptor (ANDR)-dependent transcription, by being recruited to ANDR target genes and specifically mediating phosphorylation of 'Thr-6' of histone H3 (H3T6ph), a specific tag for epigenetic transcriptional activation that prevents demethylation of histone H3 'Lys-4' (H3K4me) by LSD1/KDM1A. In insulin signaling, may function downstream of IRS1 in muscle cells and mediate insulin-dependent DNA synthesis through the RAF1-MAPK/ERK signaling cascade. May participate in the regulation of glucose transport in adipocytes by negatively modulating the insulin-stimulated translocation of the glucose transporter SLC2A4/GLUT4. Under high glucose in pancreatic beta-cells, is probably involved in the inhibition of the insulin gene transcription, via regulation of MYC expression. In endothelial cells, activation of PRKCB induces increased phosphorylation of RB1, increased VEGFA-induced cell proliferation, and inhibits PI3K/AKT-dependent nitric oxide synthase (NOS3/eNOS) regulation by insulin, which causes endothelial dysfunction. Also involved in triglyceride homeostasis (By similarity). Phosphorylates ATF2 which promotes cooperation between ATF2 and JUN, activating transcription.
- Gene Name:
- PRKCB
- Uniprot ID:
- P05771
- Molecular Weight:
- 76868.45 Da
References
- ATSDR - Agency for Toxic Substances and Disease Registry (2008). Toxicological profile for mercury. U.S. Public Health Service in collaboration with U.S. Environmental Protection Agency (EPA). [Link]
- General Function:
- Protein serine/threonine kinase activity
- Specific Function:
- Calcium-independent, phospholipid- and diacylglycerol (DAG)-dependent serine/threonine-protein kinase that plays contrasting roles in cell death and cell survival by functioning as a pro-apoptotic protein during DNA damage-induced apoptosis, but acting as an anti-apoptotic protein during cytokine receptor-initiated cell death, is involved in tumor suppression as well as survival of several cancers, is required for oxygen radical production by NADPH oxidase and acts as positive or negative regulator in platelet functional responses. Negatively regulates B cell proliferation and also has an important function in self-antigen induced B cell tolerance induction. Upon DNA damage, activates the promoter of the death-promoting transcription factor BCLAF1/Btf to trigger BCLAF1-mediated p53/TP53 gene transcription and apoptosis. In response to oxidative stress, interact with and activate CHUK/IKKA in the nucleus, causing the phosphorylation of p53/TP53. In the case of ER stress or DNA damage-induced apoptosis, can form a complex with the tyrosine-protein kinase ABL1 which trigger apoptosis independently of p53/TP53. In cytosol can trigger apoptosis by activating MAPK11 or MAPK14, inhibiting AKT1 and decreasing the level of X-linked inhibitor of apoptosis protein (XIAP), whereas in nucleus induces apoptosis via the activation of MAPK8 or MAPK9. Upon ionizing radiation treatment, is required for the activation of the apoptosis regulators BAX and BAK, which trigger the mitochondrial cell death pathway. Can phosphorylate MCL1 and target it for degradation which is sufficient to trigger for BAX activation and apoptosis. Is required for the control of cell cycle progression both at G1/S and G2/M phases. Mediates phorbol 12-myristate 13-acetate (PMA)-induced inhibition of cell cycle progression at G1/S phase by up-regulating the CDK inhibitor CDKN1A/p21 and inhibiting the cyclin CCNA2 promoter activity. In response to UV irradiation can phosphorylate CDK1, which is important for the G2/M DNA damage checkpoint activation. Can protect glioma cells from the apoptosis induced by TNFSF10/TRAIL, probably by inducing increased phosphorylation and subsequent activation of AKT1. Is highly expressed in a number of cancer cells and promotes cell survival and resistance against chemotherapeutic drugs by inducing cyclin D1 (CCND1) and hyperphosphorylation of RB1, and via several pro-survival pathways, including NF-kappa-B, AKT1 and MAPK1/3 (ERK1/2). Can also act as tumor suppressor upon mitogenic stimulation with PMA or TPA. In N-formyl-methionyl-leucyl-phenylalanine (fMLP)-treated cells, is required for NCF1 (p47-phox) phosphorylation and activation of NADPH oxidase activity, and regulates TNF-elicited superoxide anion production in neutrophils, by direct phosphorylation and activation of NCF1 or indirectly through MAPK1/3 (ERK1/2) signaling pathways. May also play a role in the regulation of NADPH oxidase activity in eosinophil after stimulation with IL5, leukotriene B4 or PMA. In collagen-induced platelet aggregation, acts a negative regulator of filopodia formation and actin polymerization by interacting with and negatively regulating VASP phosphorylation. Downstream of PAR1, PAR4 and CD36/GP4 receptors, regulates differentially platelet dense granule secretion; acts as a positive regulator in PAR-mediated granule secretion, whereas it negatively regulates CD36/GP4-mediated granule release. Phosphorylates MUC1 in the C-terminal and regulates the interaction between MUC1 and beta-catenin. The catalytic subunit phosphorylates 14-3-3 proteins (YWHAB, YWHAZ and YWHAH) in a sphingosine-dependent fashion (By similarity).
- Gene Name:
- PRKCD
- Uniprot ID:
- Q05655
- Molecular Weight:
- 77504.445 Da
References
- ATSDR - Agency for Toxic Substances and Disease Registry (2008). Toxicological profile for mercury. U.S. Public Health Service in collaboration with U.S. Environmental Protection Agency (EPA). [Link]
- General Function:
- Signal transducer activity
- Specific Function:
- Calcium-independent, phospholipid- and diacylglycerol (DAG)-dependent serine/threonine-protein kinase that plays essential roles in the regulation of multiple cellular processes linked to cytoskeletal proteins, such as cell adhesion, motility, migration and cell cycle, functions in neuron growth and ion channel regulation, and is involved in immune response, cancer cell invasion and regulation of apoptosis. Mediates cell adhesion to the extracellular matrix via integrin-dependent signaling, by mediating angiotensin-2-induced activation of integrin beta-1 (ITGB1) in cardiac fibroblasts. Phosphorylates MARCKS, which phosphorylates and activates PTK2/FAK, leading to the spread of cardiomyocytes. Involved in the control of the directional transport of ITGB1 in mesenchymal cells by phosphorylating vimentin (VIM), an intermediate filament (IF) protein. In epithelial cells, associates with and phosphorylates keratin-8 (KRT8), which induces targeting of desmoplakin at desmosomes and regulates cell-cell contact. Phosphorylates IQGAP1, which binds to CDC42, mediating epithelial cell-cell detachment prior to migration. In HeLa cells, contributes to hepatocyte growth factor (HGF)-induced cell migration, and in human corneal epithelial cells, plays a critical role in wound healing after activation by HGF. During cytokinesis, forms a complex with YWHAB, which is crucial for daughter cell separation, and facilitates abscission by a mechanism which may implicate the regulation of RHOA. In cardiac myocytes, regulates myofilament function and excitation coupling at the Z-lines, where it is indirectly associated with F-actin via interaction with COPB1. During endothelin-induced cardiomyocyte hypertrophy, mediates activation of PTK2/FAK, which is critical for cardiomyocyte survival and regulation of sarcomere length. Plays a role in the pathogenesis of dilated cardiomyopathy via persistent phosphorylation of troponin I (TNNI3). Involved in nerve growth factor (NFG)-induced neurite outgrowth and neuron morphological change independently of its kinase activity, by inhibition of RHOA pathway, activation of CDC42 and cytoskeletal rearrangement. May be involved in presynaptic facilitation by mediating phorbol ester-induced synaptic potentiation. Phosphorylates gamma-aminobutyric acid receptor subunit gamma-2 (GABRG2), which reduces the response of GABA receptors to ethanol and benzodiazepines and may mediate acute tolerance to the intoxicating effects of ethanol. Upon PMA treatment, phosphorylates the capsaicin- and heat-activated cation channel TRPV1, which is required for bradykinin-induced sensitization of the heat response in nociceptive neurons. Is able to form a complex with PDLIM5 and N-type calcium channel, and may enhance channel activities and potentiates fast synaptic transmission by phosphorylating the pore-forming alpha subunit CACNA1B (CaV2.2). In prostate cancer cells, interacts with and phosphorylates STAT3, which increases DNA-binding and transcriptional activity of STAT3 and seems to be essential for prostate cancer cell invasion. Downstream of TLR4, plays an important role in the lipopolysaccharide (LPS)-induced immune response by phosphorylating and activating TICAM2/TRAM, which in turn activates the transcription factor IRF3 and subsequent cytokines production. In differentiating erythroid progenitors, is regulated by EPO and controls the protection against the TNFSF10/TRAIL-mediated apoptosis, via BCL2. May be involved in the regulation of the insulin-induced phosphorylation and activation of AKT1.
- Gene Name:
- PRKCE
- Uniprot ID:
- Q02156
- Molecular Weight:
- 83673.2 Da
References
- ATSDR - Agency for Toxic Substances and Disease Registry (2008). Toxicological profile for mercury. U.S. Public Health Service in collaboration with U.S. Environmental Protection Agency (EPA). [Link]
- General Function:
- Protein kinase c activity
- Specific Function:
- Calcium-independent, phospholipid- and diacylglycerol (DAG)-dependent serine/threonine-protein kinase that is involved in the regulation of cell differentiation in keratinocytes and pre-B cell receptor, mediates regulation of epithelial tight junction integrity and foam cell formation, and is required for glioblastoma proliferation and apoptosis prevention in MCF-7 cells. In keratinocytes, binds and activates the tyrosine kinase FYN, which in turn blocks epidermal growth factor receptor (EGFR) signaling and leads to keratinocyte growth arrest and differentiation. Associates with the cyclin CCNE1-CDK2-CDKN1B complex and inhibits CDK2 kinase activity, leading to RB1 dephosphorylation and thereby G1 arrest in keratinocytes. In association with RALA activates actin depolymerization, which is necessary for keratinocyte differentiation. In the pre-B cell receptor signaling, functions downstream of BLNK by up-regulating IRF4, which in turn activates L chain gene rearrangement. Regulates epithelial tight junctions (TJs) by phosphorylating occludin (OCLN) on threonine residues, which is necessary for the assembly and maintenance of TJs. In association with PLD2 and via TLR4 signaling, is involved in lipopolysaccharide (LPS)-induced RGS2 down-regulation and foam cell formation. Upon PMA stimulation, mediates glioblastoma cell proliferation by activating the mTOR pathway, the PI3K/AKT pathway and the ERK1-dependent phosphorylation of ELK1. Involved in the protection of glioblastoma cells from irradiation-induced apoptosis by preventing caspase-9 activation. In camptothecin-treated MCF-7 cells, regulates NF-kappa-B upstream signaling by activating IKBKB, and confers protection against DNA damage-induced apoptosis. Promotes oncogenic functions of ATF2 in the nucleus while blocking its apoptotic function at mitochondria. Phosphorylates ATF2 which promotes its nuclear retention and transcriptional activity and negatively regulates its mitochondrial localization.
- Gene Name:
- PRKCH
- Uniprot ID:
- P24723
- Molecular Weight:
- 77827.96 Da
References
- ATSDR - Agency for Toxic Substances and Disease Registry (2008). Toxicological profile for mercury. U.S. Public Health Service in collaboration with U.S. Environmental Protection Agency (EPA). [Link]
- General Function:
- Zinc ion binding
- Specific Function:
- Calcium-activated, phospholipid- and diacylglycerol (DAG)-dependent serine/threonine-protein kinase that plays diverse roles in neuronal cells and eye tissues, such as regulation of the neuronal receptors GRIA4/GLUR4 and GRIN1/NMDAR1, modulation of receptors and neuronal functions related to sensitivity to opiates, pain and alcohol, mediation of synaptic function and cell survival after ischemia, and inhibition of gap junction activity after oxidative stress. Binds and phosphorylates GRIA4/GLUR4 glutamate receptor and regulates its function by increasing plasma membrane-associated GRIA4 expression. In primary cerebellar neurons treated with the agonist 3,5-dihyidroxyphenylglycine, functions downstream of the metabotropic glutamate receptor GRM5/MGLUR5 and phosphorylates GRIN1/NMDAR1 receptor which plays a key role in synaptic plasticity, synaptogenesis, excitotoxicity, memory acquisition and learning. May be involved in the regulation of hippocampal long-term potentiation (LTP), but may be not necessary for the process of synaptic plasticity. May be involved in desensitization of mu-type opioid receptor-mediated G-protein activation in the spinal cord, and may be critical for the development and/or maintenance of morphine-induced reinforcing effects in the limbic forebrain. May modulate the functionality of mu-type-opioid receptors by participating in a signaling pathway which leads to the phosphorylation and degradation of opioid receptors. May also contributes to chronic morphine-induced changes in nociceptive processing. Plays a role in neuropathic pain mechanisms and contributes to the maintenance of the allodynia pain produced by peripheral inflammation. Plays an important role in initial sensitivity and tolerance to ethanol, by mediating the behavioral effects of ethanol as well as the effects of this drug on the GABA(A) receptors. During and after cerebral ischemia modulate neurotransmission and cell survival in synaptic membranes, and is involved in insulin-induced inhibition of necrosis, an important mechanism for minimizing ischemic injury. Required for the elimination of multiple climbing fibers during innervation of Purkinje cells in developing cerebellum. Is activated in lens epithelial cells upon hydrogen peroxide treatment, and phosphorylates connexin-43 (GJA1/CX43), resulting in disassembly of GJA1 gap junction plaques and inhibition of gap junction activity which could provide a protective effect against oxidative stress (By similarity). Phosphorylates p53/TP53 and promotes p53/TP53-dependent apoptosis in response to DNA damage. Involved in the phase resetting of the cerebral cortex circadian clock during temporally restricted feeding. Stabilizes the core clock component ARNTL/BMAL1 by interfering with its ubiquitination, thus suppressing its degradation, resulting in phase resetting of the cerebral cortex clock (By similarity).
- Gene Name:
- PRKCG
- Uniprot ID:
- P05129
- Molecular Weight:
- 78447.23 Da
References
- ATSDR - Agency for Toxic Substances and Disease Registry (2008). Toxicological profile for mercury. U.S. Public Health Service in collaboration with U.S. Environmental Protection Agency (EPA). [Link]
- General Function:
- Protein serine/threonine kinase activity
- Specific Function:
- Calcium- and diacylglycerol-independent serine/ threonine-protein kinase that plays a general protective role against apoptotic stimuli, is involved in NF-kappa-B activation, cell survival, differentiation and polarity, and contributes to the regulation of microtubule dynamics in the early secretory pathway. Is necessary for BCR-ABL oncogene-mediated resistance to apoptotic drug in leukemia cells, protecting leukemia cells against drug-induced apoptosis. In cultured neurons, prevents amyloid beta protein-induced apoptosis by interrupting cell death process at a very early step. In glioblastoma cells, may function downstream of phosphatidylinositol 3-kinase (PI(3)K) and PDPK1 in the promotion of cell survival by phosphorylating and inhibiting the pro-apoptotic factor BAD. Can form a protein complex in non-small cell lung cancer (NSCLC) cells with PARD6A and ECT2 and regulate ECT2 oncogenic activity by phosphorylation, which in turn promotes transformed growth and invasion. In response to nerve growth factor (NGF), acts downstream of SRC to phosphorylate and activate IRAK1, allowing the subsequent activation of NF-kappa-B and neuronal cell survival. Functions in the organization of the apical domain in epithelial cells by phosphorylating EZR. This step is crucial for activation and normal distribution of EZR at the early stages of intestinal epithelial cell differentiation. Forms a protein complex with LLGL1 and PARD6B independently of PARD3 to regulate epithelial cell polarity. Plays a role in microtubule dynamics in the early secretory pathway through interaction with RAB2A and GAPDH and recruitment to vesicular tubular clusters (VTCs). In human coronary artery endothelial cells (HCAEC), is activated by saturated fatty acids and mediates lipid-induced apoptosis.
- Gene Name:
- PRKCI
- Uniprot ID:
- P41743
- Molecular Weight:
- 68261.855 Da
References
- ATSDR - Agency for Toxic Substances and Disease Registry (2008). Toxicological profile for mercury. U.S. Public Health Service in collaboration with U.S. Environmental Protection Agency (EPA). [Link]
- General Function:
- Ubiquitin-protein transferase activity
- Specific Function:
- Calcium-independent, phospholipid- and diacylglycerol (DAG)-dependent serine/threonine-protein kinase that mediates non-redundant functions in T-cell receptor (TCR) signaling, including T-cells activation, proliferation, differentiation and survival, by mediating activation of multiple transcription factors such as NF-kappa-B, JUN, NFATC1 and NFATC2. In TCR-CD3/CD28-co-stimulated T-cells, is required for the activation of NF-kappa-B and JUN, which in turn are essential for IL2 production, and participates to the calcium-dependent NFATC1 and NFATC2 transactivation. Mediates the activation of the canonical NF-kappa-B pathway (NFKB1) by direct phosphorylation of CARD11 on several serine residues, inducing CARD11 association with lipid rafts and recruitment of the BCL10-MALT1 complex, which then activates IKK complex, resulting in nuclear translocation and activation of NFKB1. May also play an indirect role in activation of the non-canonical NF-kappa-B (NFKB2) pathway. In the signaling pathway leading to JUN activation, acts by phosphorylating the mediator STK39/SPAK and may not act through MAP kinases signaling. Plays a critical role in TCR/CD28-induced NFATC1 and NFATC2 transactivation by participating in the regulation of reduced inositol 1,4,5-trisphosphate generation and intracellular calcium mobilization. After costimulation of T-cells through CD28 can phosphorylate CBLB and is required for the ubiquitination and subsequent degradation of CBLB, which is a prerequisite for the activation of TCR. During T-cells differentiation, plays an important role in the development of T-helper 2 (Th2) cells following immune and inflammatory responses, and, in the development of inflammatory autoimmune diseases, is necessary for the activation of IL17-producing Th17 cells. May play a minor role in Th1 response. Upon TCR stimulation, mediates T-cell protective survival signal by phosphorylating BAD, thus protecting T-cells from BAD-induced apoptosis, and by up-regulating BCL-X(L)/BCL2L1 levels through NF-kappa-B and JUN pathways. In platelets, regulates signal transduction downstream of the ITGA2B, CD36/GP4, F2R/PAR1 and F2RL3/PAR4 receptors, playing a positive role in 'outside-in' signaling and granule secretion signal transduction. May relay signals from the activated ITGA2B receptor by regulating the uncoupling of WASP and WIPF1, thereby permitting the regulation of actin filament nucleation and branching activity of the Arp2/3 complex. May mediate inhibitory effects of free fatty acids on insulin signaling by phosphorylating IRS1, which in turn blocks IRS1 tyrosine phosphorylation and downstream activation of the PI3K/AKT pathway. Phosphorylates MSN (moesin) in the presence of phosphatidylglycerol or phosphatidylinositol. Phosphorylates PDPK1 at 'Ser-504' and 'Ser-532' and negatively regulates its ability to phosphorylate PKB/AKT1.
- Gene Name:
- PRKCQ
- Uniprot ID:
- Q04759
- Molecular Weight:
- 81864.145 Da
References
- ATSDR - Agency for Toxic Substances and Disease Registry (2008). Toxicological profile for mercury. U.S. Public Health Service in collaboration with U.S. Environmental Protection Agency (EPA). [Link]
- General Function:
- Protein serine/threonine kinase activity
- Specific Function:
- Calcium- and diacylglycerol-independent serine/threonine-protein kinase that functions in phosphatidylinositol 3-kinase (PI3K) pathway and mitogen-activated protein (MAP) kinase cascade, and is involved in NF-kappa-B activation, mitogenic signaling, cell proliferation, cell polarity, inflammatory response and maintenance of long-term potentiation (LTP). Upon lipopolysaccharide (LPS) treatment in macrophages, or following mitogenic stimuli, functions downstream of PI3K to activate MAP2K1/MEK1-MAPK1/ERK2 signaling cascade independently of RAF1 activation. Required for insulin-dependent activation of AKT3, but may function as an adapter rather than a direct activator. Upon insulin treatment may act as a downstream effector of PI3K and contribute to the activation of translocation of the glucose transporter SLC2A4/GLUT4 and subsequent glucose transport in adipocytes. In EGF-induced cells, binds and activates MAP2K5/MEK5-MAPK7/ERK5 independently of its kinase activity and can activate JUN promoter through MEF2C. Through binding with SQSTM1/p62, functions in interleukin-1 signaling and activation of NF-kappa-B with the specific adapters RIPK1 and TRAF6. Participates in TNF-dependent transactivation of NF-kappa-B by phosphorylating and activating IKBKB kinase, which in turn leads to the degradation of NF-kappa-B inhibitors. In migrating astrocytes, forms a cytoplasmic complex with PARD6A and is recruited by CDC42 to function in the establishment of cell polarity along with the microtubule motor and dynein. In association with FEZ1, stimulates neuronal differentiation in PC12 cells. In the inflammatory response, is required for the T-helper 2 (Th2) differentiation process, including interleukin production, efficient activation of JAK1 and the subsequent phosphorylation and nuclear translocation of STAT6. May be involved in development of allergic airway inflammation (asthma), a process dependent on Th2 immune response. In the NF-kappa-B-mediated inflammatory response, can relieve SETD6-dependent repression of NF-kappa-B target genes by phosphorylating the RELA subunit at 'Ser-311'. Necessary and sufficient for LTP maintenance in hippocampal CA1 pyramidal cells. In vein endothelial cells treated with the oxidant peroxynitrite, phosphorylates STK11 leading to nuclear export of STK11, subsequent inhibition of PI3K/Akt signaling, and increased apoptosis. Phosphorylates VAMP2 in vitro (PubMed:17313651).
- Gene Name:
- PRKCZ
- Uniprot ID:
- Q05513
- Molecular Weight:
- 67659.335 Da
References
- ATSDR - Agency for Toxic Substances and Disease Registry (2008). Toxicological profile for mercury. U.S. Public Health Service in collaboration with U.S. Environmental Protection Agency (EPA). [Link]
- General Function:
- Not Available
- Specific Function:
- Not Available
- Gene Name:
- Not Available
- Uniprot ID:
- A6NKZ8
- Molecular Weight:
- Not Available
References
- ATSDR - Agency for Toxic Substances and Disease Registry (2008). Toxicological profile for mercury. U.S. Public Health Service in collaboration with U.S. Environmental Protection Agency (EPA). [Link]
- General Function:
- Not Available
- Specific Function:
- Not Available
- Gene Name:
- Not Available
- Uniprot ID:
- Q99867
- Molecular Weight:
- Not Available
References
- ATSDR - Agency for Toxic Substances and Disease Registry (2008). Toxicological profile for mercury. U.S. Public Health Service in collaboration with U.S. Environmental Protection Agency (EPA). [Link]
- General Function:
- Structural constituent of cytoskeleton
- Specific Function:
- Not Available
- Gene Name:
- TUBA4B
- Uniprot ID:
- Q9H853
- Molecular Weight:
- 27551.01 Da
References
- ATSDR - Agency for Toxic Substances and Disease Registry (2008). Toxicological profile for mercury. U.S. Public Health Service in collaboration with U.S. Environmental Protection Agency (EPA). [Link]
- General Function:
- Ubiquinone binding
- Specific Function:
- Membrane-anchoring subunit of succinate dehydrogenase (SDH) that is involved in complex II of the mitochondrial electron transport chain and is responsible for transferring electrons from succinate to ubiquinone (coenzyme Q).
- Gene Name:
- SDHD
- Uniprot ID:
- O14521
- Molecular Weight:
- 17042.82 Da
References
- Ardelt BK, Borowitz JL, Isom GE: Brain lipid peroxidation and antioxidant protectant mechanisms following acute cyanide intoxication. Toxicology. 1989 Jun 1;56(2):147-54. [2734799 ]
- General Function:
- Succinate dehydrogenase activity
- Specific Function:
- Flavoprotein (FP) subunit of succinate dehydrogenase (SDH) that is involved in complex II of the mitochondrial electron transport chain and is responsible for transferring electrons from succinate to ubiquinone (coenzyme Q). Can act as a tumor suppressor.
- Gene Name:
- SDHA
- Uniprot ID:
- P31040
- Molecular Weight:
- 72690.975 Da
References
- Ardelt BK, Borowitz JL, Isom GE: Brain lipid peroxidation and antioxidant protectant mechanisms following acute cyanide intoxication. Toxicology. 1989 Jun 1;56(2):147-54. [2734799 ]
- General Function:
- Ubiquinone binding
- Specific Function:
- Iron-sulfur protein (IP) subunit of succinate dehydrogenase (SDH) that is involved in complex II of the mitochondrial electron transport chain and is responsible for transferring electrons from succinate to ubiquinone (coenzyme Q).
- Gene Name:
- SDHB
- Uniprot ID:
- P21912
- Molecular Weight:
- 31629.365 Da
References
- Ardelt BK, Borowitz JL, Isom GE: Brain lipid peroxidation and antioxidant protectant mechanisms following acute cyanide intoxication. Toxicology. 1989 Jun 1;56(2):147-54. [2734799 ]
- General Function:
- Succinate dehydrogenase activity
- Specific Function:
- Membrane-anchoring subunit of succinate dehydrogenase (SDH) that is involved in complex II of the mitochondrial electron transport chain and is responsible for transferring electrons from succinate to ubiquinone (coenzyme Q).
- Gene Name:
- SDHC
- Uniprot ID:
- Q99643
- Molecular Weight:
- 18610.03 Da
References
- Ardelt BK, Borowitz JL, Isom GE: Brain lipid peroxidation and antioxidant protectant mechanisms following acute cyanide intoxication. Toxicology. 1989 Jun 1;56(2):147-54. [2734799 ]
- General Function:
- Zinc ion binding
- Specific Function:
- Destroys radicals which are normally produced within the cells and which are toxic to biological systems.
- Gene Name:
- SOD1
- Uniprot ID:
- P00441
- Molecular Weight:
- 15935.685 Da
References
- Lee WG, Hwang JH, Na BK, Cho JH, Lee HW, Cho SH, Kong Y, Song CY, Kim TS: Functional expression of a recombinant copper/zinc superoxide dismutase of filarial nematode, Brugia malayi. J Parasitol. 2005 Feb;91(1):205-8. [15856906 ]
- General Function:
- Structural constituent of cytoskeleton
- Specific Function:
- Tubulin is the major constituent of microtubules. It binds two moles of GTP, one at an exchangeable site on the beta chain and one at a non-exchangeable site on the alpha chain (By similarity).
- Gene Name:
- TUBAL3
- Uniprot ID:
- A6NHL2
- Molecular Weight:
- 49908.305 Da
References
- ATSDR - Agency for Toxic Substances and Disease Registry (2008). Toxicological profile for mercury. U.S. Public Health Service in collaboration with U.S. Environmental Protection Agency (EPA). [Link]
- General Function:
- Tubulin is the major constituent of microtubules. It binds two moles of GTP, one at an exchangeable site on the beta chain and one at a non-exchangeable site on the alpha chain.
- Specific Function:
- Gtp binding
- Gene Name:
- TUBA1A
- Uniprot ID:
- Q71U36
- Molecular Weight:
- 50135.25 Da
References
- ATSDR - Agency for Toxic Substances and Disease Registry (2008). Toxicological profile for mercury. U.S. Public Health Service in collaboration with U.S. Environmental Protection Agency (EPA). [Link]
- General Function:
- Ubiquitin protein ligase binding
- Specific Function:
- Tubulin is the major constituent of microtubules. It binds two moles of GTP, one at an exchangeable site on the beta chain and one at a non-exchangeable site on the alpha chain.
- Gene Name:
- TUBA1B
- Uniprot ID:
- P68363
- Molecular Weight:
- 50151.24 Da
References
- ATSDR - Agency for Toxic Substances and Disease Registry (2008). Toxicological profile for mercury. U.S. Public Health Service in collaboration with U.S. Environmental Protection Agency (EPA). [Link]
- General Function:
- Structural molecule activity
- Specific Function:
- Tubulin is the major constituent of microtubules. It binds two moles of GTP, one at an exchangeable site on the beta chain and one at a non-exchangeable site on the alpha chain.
- Gene Name:
- TUBA1C
- Uniprot ID:
- Q9BQE3
- Molecular Weight:
- 49894.93 Da
References
- ATSDR - Agency for Toxic Substances and Disease Registry (2008). Toxicological profile for mercury. U.S. Public Health Service in collaboration with U.S. Environmental Protection Agency (EPA). [Link]
- General Function:
- Structural constituent of cytoskeleton
- Specific Function:
- Tubulin is the major constituent of microtubules. It binds two moles of GTP, one at an exchangeable site on the beta chain and one at a non-exchangeable site on the alpha chain.
- Gene Name:
- TUBA3C
- Uniprot ID:
- Q13748
- Molecular Weight:
- 49959.145 Da
References
- ATSDR - Agency for Toxic Substances and Disease Registry (2008). Toxicological profile for mercury. U.S. Public Health Service in collaboration with U.S. Environmental Protection Agency (EPA). [Link]
- General Function:
- Structural constituent of cytoskeleton
- Specific Function:
- Tubulin is the major constituent of microtubules. It binds two moles of GTP, one at an exchangeable site on the beta chain and one at a non-exchangeable site on the alpha chain (By similarity).
- Gene Name:
- TUBA3E
- Uniprot ID:
- Q6PEY2
- Molecular Weight:
- 49858.135 Da
References
- ATSDR - Agency for Toxic Substances and Disease Registry (2008). Toxicological profile for mercury. U.S. Public Health Service in collaboration with U.S. Environmental Protection Agency (EPA). [Link]
- General Function:
- Structural constituent of cytoskeleton
- Specific Function:
- Tubulin is the major constituent of microtubules. It binds two moles of GTP, one at an exchangeable site on the beta chain and one at a non-exchangeable site on the alpha chain.
- Gene Name:
- TUBA4A
- Uniprot ID:
- P68366
- Molecular Weight:
- 49923.995 Da
References
- ATSDR - Agency for Toxic Substances and Disease Registry (2008). Toxicological profile for mercury. U.S. Public Health Service in collaboration with U.S. Environmental Protection Agency (EPA). [Link]
- General Function:
- Tubulin is the major constituent of microtubules. It binds two moles of GTP, one at an exchangeable site on the beta chain and one at a non-exchangeable site on the alpha chain.
- Specific Function:
- Gtp binding
- Gene Name:
- TUBA8
- Uniprot ID:
- Q9NY65
- Molecular Weight:
- 50093.12 Da
References
- ATSDR - Agency for Toxic Substances and Disease Registry (2008). Toxicological profile for mercury. U.S. Public Health Service in collaboration with U.S. Environmental Protection Agency (EPA). [Link]
- General Function:
- Ubiquitin protein ligase binding
- Specific Function:
- Tubulin is the major constituent of microtubules. It binds two moles of GTP, one at an exchangeable site on the beta chain and one at a non-exchangeable site on the alpha chain.
- Gene Name:
- TUBB
- Uniprot ID:
- P07437
- Molecular Weight:
- 49670.515 Da
References
- ATSDR - Agency for Toxic Substances and Disease Registry (2008). Toxicological profile for mercury. U.S. Public Health Service in collaboration with U.S. Environmental Protection Agency (EPA). [Link]
- General Function:
- Structural constituent of cytoskeleton
- Specific Function:
- Tubulin is the major constituent of microtubules. It binds two moles of GTP, one at an exchangeable site on the beta chain and one at a non-exchangeable site on the alpha chain (By similarity).
- Gene Name:
- TUBB1
- Uniprot ID:
- Q9H4B7
- Molecular Weight:
- 50326.56 Da
References
- ATSDR - Agency for Toxic Substances and Disease Registry (2008). Toxicological profile for mercury. U.S. Public Health Service in collaboration with U.S. Environmental Protection Agency (EPA). [Link]
- General Function:
- Structural constituent of cytoskeleton
- Specific Function:
- Tubulin is the major constituent of microtubules. It binds two moles of GTP, one at an exchangeable site on the beta chain and one at a non-exchangeable site on the alpha chain (By similarity).
- Gene Name:
- TUBB2A
- Uniprot ID:
- Q13885
- Molecular Weight:
- 49906.67 Da
References
- ATSDR - Agency for Toxic Substances and Disease Registry (2008). Toxicological profile for mercury. U.S. Public Health Service in collaboration with U.S. Environmental Protection Agency (EPA). [Link]
- General Function:
- Structural constituent of cytoskeleton
- Specific Function:
- Tubulin is the major constituent of microtubules. It binds two moles of GTP, one at an exchangeable site on the beta chain and one at a non-exchangeable site on the alpha chain (By similarity). TUBB2B is implicated in neuronal migration.
- Gene Name:
- TUBB2B
- Uniprot ID:
- Q9BVA1
- Molecular Weight:
- 49952.76 Da
References
- ATSDR - Agency for Toxic Substances and Disease Registry (2008). Toxicological profile for mercury. U.S. Public Health Service in collaboration with U.S. Environmental Protection Agency (EPA). [Link]
- General Function:
- Structural constituent of cytoskeleton
- Specific Function:
- Tubulin is the major constituent of microtubules. It binds two moles of GTP, one at an exchangeable site on the beta chain and one at a non-exchangeable site on the alpha chain. TUBB3 plays a critical role in proper axon guidance and mantainance.
- Gene Name:
- TUBB3
- Uniprot ID:
- Q13509
- Molecular Weight:
- 50432.355 Da
References
- ATSDR - Agency for Toxic Substances and Disease Registry (2008). Toxicological profile for mercury. U.S. Public Health Service in collaboration with U.S. Environmental Protection Agency (EPA). [Link]
- General Function:
- Structural constituent of cytoskeleton
- Specific Function:
- Tubulin is the major constituent of microtubules. It binds two moles of GTP, one at an exchangeable site on the beta chain and one at a non-exchangeable site on the alpha chain.
- Gene Name:
- TUBB4A
- Uniprot ID:
- P04350
- Molecular Weight:
- 49585.475 Da
References
- ATSDR - Agency for Toxic Substances and Disease Registry (2008). Toxicological profile for mercury. U.S. Public Health Service in collaboration with U.S. Environmental Protection Agency (EPA). [Link]
- General Function:
- Unfolded protein binding
- Specific Function:
- Tubulin is the major constituent of microtubules. It binds two moles of GTP, one at an exchangeable site on the beta chain and one at a non-exchangeable site on the alpha chain.
- Gene Name:
- TUBB4B
- Uniprot ID:
- P68371
- Molecular Weight:
- 49830.72 Da
References
- ATSDR - Agency for Toxic Substances and Disease Registry (2008). Toxicological profile for mercury. U.S. Public Health Service in collaboration with U.S. Environmental Protection Agency (EPA). [Link]
- General Function:
- Structural constituent of cytoskeleton
- Specific Function:
- Tubulin is the major constituent of microtubules. It binds two moles of GTP, one at an exchangeable site on the beta chain and one at a non-exchangeable site on the alpha chain (By similarity).
- Gene Name:
- TUBB6
- Uniprot ID:
- Q9BUF5
- Molecular Weight:
- 49856.785 Da
References
- ATSDR - Agency for Toxic Substances and Disease Registry (2008). Toxicological profile for mercury. U.S. Public Health Service in collaboration with U.S. Environmental Protection Agency (EPA). [Link]
- General Function:
- Structural constituent of cytoskeleton
- Specific Function:
- Tubulin is the major constituent of microtubules. It binds two moles of GTP, one at an exchangeable site on the beta chain and one at a non-exchangeable site on the alpha chain (By similarity).
- Gene Name:
- TUBB8
- Uniprot ID:
- Q3ZCM7
- Molecular Weight:
- 49775.655 Da
References
- ATSDR - Agency for Toxic Substances and Disease Registry (2008). Toxicological profile for mercury. U.S. Public Health Service in collaboration with U.S. Environmental Protection Agency (EPA). [Link]
- General Function:
- Structural constituent of cytoskeleton
- Specific Function:
- Tubulin is the major constituent of microtubules. It binds two moles of GTP, one at an exchangeable site on the beta chain and one at a non-exchangeable site on the alpha chain (By similarity).
- Gene Name:
- Not Available
- Uniprot ID:
- A6NNZ2
- Molecular Weight:
- 49572.265 Da
References
- ATSDR - Agency for Toxic Substances and Disease Registry (2008). Toxicological profile for mercury. U.S. Public Health Service in collaboration with U.S. Environmental Protection Agency (EPA). [Link]
- General Function:
- Structural constituent of cytoskeleton
- Specific Function:
- In the elongating spermatid it is associated with the manchette, a specialized microtubule system present during reshaping of the sperm head.
- Gene Name:
- TUBD1
- Uniprot ID:
- Q9UJT1
- Molecular Weight:
- 51033.86 Da
References
- ATSDR - Agency for Toxic Substances and Disease Registry (2008). Toxicological profile for mercury. U.S. Public Health Service in collaboration with U.S. Environmental Protection Agency (EPA). [Link]
- General Function:
- Structural constituent of cytoskeleton
- Specific Function:
- Not Available
- Gene Name:
- TUBE1
- Uniprot ID:
- Q9UJT0
- Molecular Weight:
- 52931.4 Da
References
- ATSDR - Agency for Toxic Substances and Disease Registry (2008). Toxicological profile for mercury. U.S. Public Health Service in collaboration with U.S. Environmental Protection Agency (EPA). [Link]
- General Function:
- Structural constituent of cytoskeleton
- Specific Function:
- Tubulin is the major constituent of microtubules. The gamma chain is found at microtubule organizing centers (MTOC) such as the spindle poles or the centrosome. Pericentriolar matrix component that regulates alpha/beta chain minus-end nucleation, centrosome duplication and spindle formation.
- Gene Name:
- TUBG1
- Uniprot ID:
- P23258
- Molecular Weight:
- 51169.48 Da
References
- ATSDR - Agency for Toxic Substances and Disease Registry (2008). Toxicological profile for mercury. U.S. Public Health Service in collaboration with U.S. Environmental Protection Agency (EPA). [Link]
- General Function:
- Structural molecule activity
- Specific Function:
- Tubulin is the major constituent of microtubules. The gamma chain is found at microtubule organizing centers (MTOC) such as the spindle poles or the centrosome. Pericentriolar matrix component that regulates alpha/beta chain minus-end nucleation, centrosome duplication and spindle formation (By similarity).
- Gene Name:
- TUBG2
- Uniprot ID:
- Q9NRH3
- Molecular Weight:
- 51091.32 Da
References
- ATSDR - Agency for Toxic Substances and Disease Registry (2008). Toxicological profile for mercury. U.S. Public Health Service in collaboration with U.S. Environmental Protection Agency (EPA). [Link]
- General Function:
- Protein homodimerization activity
- Specific Function:
- This is a copper-containing oxidase that functions in the formation of pigments such as melanins and other polyphenolic compounds. Catalyzes the rate-limiting conversions of tyrosine to DOPA, DOPA to DOPA-quinone and possibly 5,6-dihydroxyindole to indole-5,6 quinone.
- Gene Name:
- TYR
- Uniprot ID:
- P14679
- Molecular Weight:
- 60392.69 Da
References
- Laufer Z, Beckett RP, Minibayeva FV: Co-occurrence of the multicopper oxidases tyrosinase and laccase in lichens in sub-order peltigerineae. Ann Bot. 2006 Nov;98(5):1035-42. Epub 2006 Sep 1. [16950829 ]
- General Function:
- Sh2 domain binding
- Specific Function:
- Non-receptor tyrosine-protein kinase that plays an essential role in the selection and maturation of developing T-cells in the thymus and in the function of mature T-cells. Plays a key role in T-cell antigen receptor (TCR)-linked signal transduction pathways. Constitutively associated with the cytoplasmic portions of the CD4 and CD8 surface receptors. Association of the TCR with a peptide antigen-bound MHC complex facilitates the interaction of CD4 and CD8 with MHC class II and class I molecules, respectively, thereby recruiting the associated LCK protein to the vicinity of the TCR/CD3 complex. LCK then phosphorylates tyrosines residues within the immunoreceptor tyrosine-based activation motifs (ITAM) of the cytoplasmic tails of the TCR-gamma chains and CD3 subunits, initiating the TCR/CD3 signaling pathway. Once stimulated, the TCR recruits the tyrosine kinase ZAP70, that becomes phosphorylated and activated by LCK. Following this, a large number of signaling molecules are recruited, ultimately leading to lymphokine production. LCK also contributes to signaling by other receptor molecules. Associates directly with the cytoplasmic tail of CD2, which leads to hyperphosphorylation and activation of LCK. Also plays a role in the IL2 receptor-linked signaling pathway that controls the T-cell proliferative response. Binding of IL2 to its receptor results in increased activity of LCK. Is expressed at all stages of thymocyte development and is required for the regulation of maturation events that are governed by both pre-TCR and mature alpha beta TCR. Phosphorylates other substrates including RUNX3, PTK2B/PYK2, the microtubule-associated protein MAPT, RHOH or TYROBP.
- Gene Name:
- LCK
- Uniprot ID:
- P06239
- Molecular Weight:
- 58000.15 Da
References
- Ziemba SE, Menard SL, McCabe MJ Jr, Rosenspire AJ: T-cell receptor signaling is mediated by transient Lck activity, which is inhibited by inorganic mercury. FASEB J. 2009 Jun;23(6):1663-71. doi: 10.1096/fj.08-117283. Epub 2009 Jan 23. [19168706 ]