Chlorfenapyr (T3D1699)
| Record Information | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Version | 2.0 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Creation Date | 2009-06-22 16:08:28 UTC | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Update Date | 2014-12-24 20:24:29 UTC | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Accession Number | T3D1699 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Identification | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Common Name | Chlorfenapyr | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Class | Small Molecule | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Description | Chlorfenapyr is a chemical compound of cyanide and a pesticide derived from a class of microbially produced compounds known as halogenated pyrroles. Its use is regulated in most areas due to its toxicity. (5) | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Compound Type |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
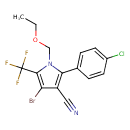
| Chemical Structure | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Synonyms |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Chemical Formula | C15H11BrClF3N2O | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Average Molecular Mass | 407.613 g/mol | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Monoisotopic Mass | 405.970 g/mol | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| CAS Registry Number | 122453-73-0 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| IUPAC Name | 4-bromo-2-(4-chlorophenyl)-1-(ethoxymethyl)-5-(trifluoromethyl)-1H-pyrrole-3-carbonitrile | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Traditional Name | pylon | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| SMILES | CCOCN1C(=C(C#N)C(Br)=C1C(F)(F)F)C1=CC=C(Cl)C=C1 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| InChI Identifier | InChI=1S/C15H11BrClF3N2O/c1-2-23-8-22-13(9-3-5-10(17)6-4-9)11(7-21)12(16)14(22)15(18,19)20/h3-6H,2,8H2,1H3 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| InChI Key | InChIKey=CWFOCCVIPCEQCK-UHFFFAOYSA-N | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Chemical Taxonomy | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Description | belongs to the class of organic compounds known as phenylpyrroles. These are polycyclic aromatic compounds containing a benzene ring linked to a pyrrole ring through a CC or CN bond. | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Kingdom | Organic compounds | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Super Class | Organoheterocyclic compounds | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Class | Pyrroles | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Sub Class | Substituted pyrroles | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Direct Parent | Phenylpyrroles | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Alternative Parents | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Substituents |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Molecular Framework | Aromatic heteromonocyclic compounds | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| External Descriptors |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Biological Properties | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Status | Detected and Not Quantified | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Origin | Exogenous | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Cellular Locations |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Biofluid Locations | Not Available | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Tissue Locations | Not Available | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Pathways | Not Available | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Applications | Not Available | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Biological Roles | Not Available | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Chemical Roles | Not Available | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Physical Properties | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| State | Solid | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Appearance | White powder. | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Experimental Properties |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Predicted Properties |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Spectra | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Spectra |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Toxicity Profile | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Route of Exposure | Oral (3) ; inhalation (3) ; dermal (3) | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Mechanism of Toxicity | Organic nitriles decompose into cyanide ions both in vivo and in vitro. Consequently the primary mechanism of toxicity for organic nitriles is their production of toxic cyanide ions or hydrogen cyanide. Cyanide is an inhibitor of cytochrome c oxidase in the fourth complex of the electron transport chain (found in the membrane of the mitochondria of eukaryotic cells). It complexes with the ferric iron atom in this enzyme. The binding of cyanide to this cytochrome prevents transport of electrons from cytochrome c oxidase to oxygen. As a result, the electron transport chain is disrupted and the cell can no longer aerobically produce ATP for energy. Tissues that mainly depend on aerobic respiration, such as the central nervous system and the heart, are particularly affected. Cyanide is also known produce some of its toxic effects by binding to catalase, glutathione peroxidase, methemoglobin, hydroxocobalamin, phosphatase, tyrosinase, ascorbic acid oxidase, xanthine oxidase, succinic dehydrogenase, and Cu/Zn superoxide dismutase. Cyanide binds to the ferric ion of methemoglobin to form inactive cyanmethemoglobin. (4) | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Metabolism | Organic nitriles are converted into cyanide ions through the action of cytochrome P450 enzymes in the liver. Cyanide is rapidly absorbed and distributed throughout the body. Cyanide is mainly metabolized into thiocyanate by either rhodanese or 3-mercaptopyruvate sulfur transferase. Cyanide metabolites are excreted in the urine. (3) | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Toxicity Values | LD50: 441 mg/kg (Oral, Rat) (2) | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Lethal Dose | 215 to 300 milligrams for an adult human (cyanide salts). (1) | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Carcinogenicity (IARC Classification) | No indication of carcinogenicity to humans (not listed by IARC). | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Uses/Sources | Chlorfenapyr is a pesticide. (5) | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Minimum Risk Level | Not Available | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Health Effects | Exposure to high levels of cyanide for a short time harms the brain and heart and can even cause coma, seizures, apnea, cardiac arrest and death. Chronic inhalation of cyanide causes breathing difficulties, chest pain, vomiting, blood changes, headaches, and enlargement of the thyroid gland. Skin contact with cyanide salts can irritate and produce sores. (3, 4) | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symptoms | Cyanide poisoning is identified by rapid, deep breathing and shortness of breath, general weakness, giddiness, headaches, vertigo, confusion, convulsions/seizures and eventually loss of consciousness. (3, 4) | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Treatment | Antidotes to cyanide poisoning include hydroxocobalamin and sodium nitrite, which release the cyanide from the cytochrome system, and rhodanase, which is an enzyme occurring naturally in mammals that combines serum cyanide with thiosulfate, producing comparatively harmless thiocyanate. Oxygen therapy can also be administered. (4) | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Normal Concentrations | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Abnormal Concentrations | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| External Links | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| DrugBank ID | Not Available | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| HMDB ID | Not Available | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PubChem Compound ID | 91778 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| ChEMBL ID | CHEMBL1869551 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| ChemSpider ID | 82875 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| KEGG ID | C18455 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| UniProt ID | Not Available | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| OMIM ID | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| ChEBI ID | 39347 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| BioCyc ID | Not Available | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| CTD ID | Not Available | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Stitch ID | Chlorfenapyr | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PDB ID | Not Available | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| ACToR ID | 7879 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Wikipedia Link | Not Available | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| References | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Synthesis Reference | Not Available | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| MSDS | T3D1699.pdf | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| General References |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Gene Regulation | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Up-Regulated Genes | Not Available | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Down-Regulated Genes | Not Available | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Targets
- General Function:
- Metal ion binding
- Specific Function:
- Not Available
- Gene Name:
- ALPPL2
- Uniprot ID:
- P10696
- Molecular Weight:
- 57376.515 Da
References
- Gerbitz KD: Human alkaline phosphatases. II. Metalloenzyme properties of the enzyme from human liver. Hoppe Seylers Z Physiol Chem. 1977 Nov;358(11):1491-7. [924371 ]
- General Function:
- Pyrophosphatase activity
- Specific Function:
- This isozyme may play a role in skeletal mineralization.
- Gene Name:
- ALPL
- Uniprot ID:
- P05186
- Molecular Weight:
- 57304.435 Da
References
- Gerbitz KD: Human alkaline phosphatases. II. Metalloenzyme properties of the enzyme from human liver. Hoppe Seylers Z Physiol Chem. 1977 Nov;358(11):1491-7. [924371 ]
- General Function:
- Receptor binding
- Specific Function:
- Occurs in almost all aerobically respiring organisms and serves to protect cells from the toxic effects of hydrogen peroxide. Promotes growth of cells including T-cells, B-cells, myeloid leukemia cells, melanoma cells, mastocytoma cells and normal and transformed fibroblast cells.
- Gene Name:
- CAT
- Uniprot ID:
- P04040
- Molecular Weight:
- 59755.82 Da
References
- Kang YS, Lee DH, Yoon BJ, Oh DC: Purification and characterization of a catalase from photosynthetic bacterium Rhodospirillum rubrum S1 grown under anaerobic conditions. J Microbiol. 2006 Apr;44(2):185-91. [16728955 ]
- General Function:
- Iron ion binding
- Specific Function:
- Cytochrome c oxidase is the component of the respiratory chain that catalyzes the reduction of oxygen to water. Subunits 1-3 form the functional core of the enzyme complex. CO I is the catalytic subunit of the enzyme. Electrons originating in cytochrome c are transferred via the copper A center of subunit 2 and heme A of subunit 1 to the bimetallic center formed by heme A3 and copper B.
- Gene Name:
- MT-CO1
- Uniprot ID:
- P00395
- Molecular Weight:
- 57040.91 Da
References
- Wikipedia. Cyanide poisoning. Last Updated 30 March 2009. [Link]
- General Function:
- Cytochrome-c oxidase activity
- Specific Function:
- Cytochrome c oxidase is the component of the respiratory chain that catalyzes the reduction of oxygen to water. Subunits 1-3 form the functional core of the enzyme complex. Subunit 2 transfers the electrons from cytochrome c via its binuclear copper A center to the bimetallic center of the catalytic subunit 1.
- Gene Name:
- MT-CO2
- Uniprot ID:
- P00403
- Molecular Weight:
- 25564.73 Da
References
- Wikipedia. Cyanide poisoning. Last Updated 30 March 2009. [Link]
- General Function:
- Cytochrome-c oxidase activity
- Specific Function:
- This protein is one of the nuclear-coded polypeptide chains of cytochrome c oxidase, the terminal oxidase in mitochondrial electron transport.
- Gene Name:
- COX4I1
- Uniprot ID:
- P13073
- Molecular Weight:
- 19576.6 Da
References
- Wikipedia. Cyanide poisoning. Last Updated 30 March 2009. [Link]
- General Function:
- Cytochrome-c oxidase activity
- Specific Function:
- This protein is one of the nuclear-coded polypeptide chains of cytochrome c oxidase, the terminal oxidase in mitochondrial electron transport.
- Gene Name:
- COX4I2
- Uniprot ID:
- Q96KJ9
- Molecular Weight:
- 20010.02 Da
References
- Wikipedia. Cyanide poisoning. Last Updated 30 March 2009. [Link]
- General Function:
- Metal ion binding
- Specific Function:
- This is the heme A-containing chain of cytochrome c oxidase, the terminal oxidase in mitochondrial electron transport.
- Gene Name:
- COX5A
- Uniprot ID:
- P20674
- Molecular Weight:
- 16761.985 Da
References
- Wikipedia. Cyanide poisoning. Last Updated 30 March 2009. [Link]
- General Function:
- Metal ion binding
- Specific Function:
- This protein is one of the nuclear-coded polypeptide chains of cytochrome c oxidase, the terminal oxidase in mitochondrial electron transport.
- Gene Name:
- COX5B
- Uniprot ID:
- P10606
- Molecular Weight:
- 13695.57 Da
References
- Wikipedia. Cyanide poisoning. Last Updated 30 March 2009. [Link]
- General Function:
- Cytochrome-c oxidase activity
- Specific Function:
- This protein is one of the nuclear-coded polypeptide chains of cytochrome c oxidase, the terminal oxidase in mitochondrial electron transport.
- Gene Name:
- COX6A1
- Uniprot ID:
- P12074
- Molecular Weight:
- 12154.8 Da
References
- Wikipedia. Cyanide poisoning. Last Updated 30 March 2009. [Link]
- General Function:
- Cytochrome-c oxidase activity
- Specific Function:
- This protein is one of the nuclear-coded polypeptide chains of cytochrome c oxidase, the terminal oxidase in mitochondrial electron transport.
- Gene Name:
- COX6A2
- Uniprot ID:
- Q02221
- Molecular Weight:
- 10815.32 Da
References
- Wikipedia. Cyanide poisoning. Last Updated 30 March 2009. [Link]
- General Function:
- Cytochrome-c oxidase activity
- Specific Function:
- This protein is one of the nuclear-coded polypeptide chains of cytochrome c oxidase, the terminal oxidase in mitochondrial electron transport.
- Gene Name:
- COX6C
- Uniprot ID:
- P09669
- Molecular Weight:
- 8781.36 Da
References
- Wikipedia. Cyanide poisoning. Last Updated 30 March 2009. [Link]
- General Function:
- Cytochrome-c oxidase activity
- Specific Function:
- This protein is one of the nuclear-coded polypeptide chains of cytochrome c oxidase, the terminal oxidase in mitochondrial electron transport.
- Gene Name:
- COX7A1
- Uniprot ID:
- P24310
- Molecular Weight:
- 9117.44 Da
References
- Wikipedia. Cyanide poisoning. Last Updated 30 March 2009. [Link]
- General Function:
- Cytochrome-c oxidase activity
- Specific Function:
- This protein is one of the nuclear-coded polypeptide chains of cytochrome c oxidase, the terminal oxidase in mitochondrial electron transport.
- Gene Name:
- COX7A2
- Uniprot ID:
- P14406
- Molecular Weight:
- 9395.89 Da
References
- Wikipedia. Cyanide poisoning. Last Updated 30 March 2009. [Link]
- General Function:
- Cytochrome-c oxidase activity
- Specific Function:
- This protein is one of the nuclear-coded polypeptide chains of cytochrome c oxidase, the terminal oxidase in mitochondrial electron transport. Plays a role in proper central nervous system (CNS) development in vertebrates.
- Gene Name:
- COX7B
- Uniprot ID:
- P24311
- Molecular Weight:
- 9160.485 Da
References
- Wikipedia. Cyanide poisoning. Last Updated 30 March 2009. [Link]
- General Function:
- Cytochrome-c oxidase activity
- Specific Function:
- This protein is one of the nuclear-coded polypeptide chains of cytochrome c oxidase, the terminal oxidase in mitochondrial electron transport.
- Gene Name:
- COX7B2
- Uniprot ID:
- Q8TF08
- Molecular Weight:
- 9077.43 Da
References
- Wikipedia. Cyanide poisoning. Last Updated 30 March 2009. [Link]
- General Function:
- Cytochrome-c oxidase activity
- Specific Function:
- This protein is one of the nuclear-coded polypeptide chains of cytochrome c oxidase, the terminal oxidase in mitochondrial electron transport.
- Gene Name:
- COX7C
- Uniprot ID:
- P15954
- Molecular Weight:
- 7245.45 Da
References
- Wikipedia. Cyanide poisoning. Last Updated 30 March 2009. [Link]
- General Function:
- Cytochrome-c oxidase activity
- Specific Function:
- This protein is one of the nuclear-coded polypeptide chains of cytochrome c oxidase, the terminal oxidase in mitochondrial electron transport.
- Gene Name:
- COX8A
- Uniprot ID:
- P10176
- Molecular Weight:
- 7579.0 Da
References
- Wikipedia. Cyanide poisoning. Last Updated 30 March 2009. [Link]
- General Function:
- Cytochrome-c oxidase activity
- Specific Function:
- This protein is one of the nuclear-coded polypeptide chains of cytochrome c oxidase, the terminal oxidase in mitochondrial electron transport.
- Gene Name:
- COX8C
- Uniprot ID:
- Q7Z4L0
- Molecular Weight:
- 8128.575 Da
References
- Wikipedia. Cyanide poisoning. Last Updated 30 March 2009. [Link]
- General Function:
- Glutathione peroxidase activity
- Specific Function:
- Protects cells and enzymes from oxidative damage, by catalyzing the reduction of hydrogen peroxide, lipid peroxides and organic hydroperoxide, by glutathione. May constitute a glutathione peroxidase-like protective system against peroxide damage in sperm membrane lipids.
- Gene Name:
- GPX5
- Uniprot ID:
- O75715
- Molecular Weight:
- 25202.14 Da
References
- Kraus RJ, Ganther HE: Reaction of cyanide with glutathione peroxidase. Biochem Biophys Res Commun. 1980 Oct 16;96(3):1116-22. [7437059 ]
- General Function:
- Zinc ion binding
- Specific Function:
- Protect the extracellular space from toxic effect of reactive oxygen intermediates by converting superoxide radicals into hydrogen peroxide and oxygen.
- Gene Name:
- SOD3
- Uniprot ID:
- P08294
- Molecular Weight:
- 25850.675 Da
References
- Lee WG, Hwang JH, Na BK, Cho JH, Lee HW, Cho SH, Kong Y, Song CY, Kim TS: Functional expression of a recombinant copper/zinc superoxide dismutase of filarial nematode, Brugia malayi. J Parasitol. 2005 Feb;91(1):205-8. [15856906 ]
- General Function:
- Sh3 domain binding
- Specific Function:
- Protects the hemoglobin in erythrocytes from oxidative breakdown.
- Gene Name:
- GPX1
- Uniprot ID:
- P07203
- Molecular Weight:
- 22087.94 Da
References
- Kraus RJ, Ganther HE: Reaction of cyanide with glutathione peroxidase. Biochem Biophys Res Commun. 1980 Oct 16;96(3):1116-22. [7437059 ]
- General Function:
- Glutathione peroxidase activity
- Specific Function:
- Could play a major role in protecting mammals from the toxicity of ingested organic hydroperoxides. Tert-butyl hydroperoxide, cumene hydroperoxide and linoleic acid hydroperoxide but not phosphatidycholine hydroperoxide, can act as acceptors.
- Gene Name:
- GPX2
- Uniprot ID:
- P18283
- Molecular Weight:
- 21953.835 Da
References
- Kraus RJ, Ganther HE: Reaction of cyanide with glutathione peroxidase. Biochem Biophys Res Commun. 1980 Oct 16;96(3):1116-22. [7437059 ]
- General Function:
- Transcription factor binding
- Specific Function:
- Protects cells and enzymes from oxidative damage, by catalyzing the reduction of hydrogen peroxide, lipid peroxides and organic hydroperoxide, by glutathione.
- Gene Name:
- GPX3
- Uniprot ID:
- P22352
- Molecular Weight:
- 25552.185 Da
References
- Kraus RJ, Ganther HE: Reaction of cyanide with glutathione peroxidase. Biochem Biophys Res Commun. 1980 Oct 16;96(3):1116-22. [7437059 ]
- General Function:
- Peroxidase activity
- Specific Function:
- It protects esophageal epithelia from hydrogen peroxide-induced oxidative stress. It suppresses acidic bile acid-induced reactive oxigen species (ROS) and protects against oxidative DNA damage and double-strand breaks.
- Gene Name:
- GPX7
- Uniprot ID:
- Q96SL4
- Molecular Weight:
- 20995.88 Da
References
- Kraus RJ, Ganther HE: Reaction of cyanide with glutathione peroxidase. Biochem Biophys Res Commun. 1980 Oct 16;96(3):1116-22. [7437059 ]
- General Function:
- Nadp binding
- Specific Function:
- Maintains high levels of reduced glutathione in the cytosol.
- Gene Name:
- GSR
- Uniprot ID:
- P00390
- Molecular Weight:
- 56256.565 Da
References
- Ardelt BK, Borowitz JL, Isom GE: Brain lipid peroxidation and antioxidant protectant mechanisms following acute cyanide intoxication. Toxicology. 1989 Jun 1;56(2):147-54. [2734799 ]
- General Function:
- Phospholipid-hydroperoxide glutathione peroxidase activity
- Specific Function:
- Protects cells against membrane lipid peroxidation and cell death. Required for normal sperm development and male fertility. Could play a major role in protecting mammals from the toxicity of ingested lipid hydroperoxides. Essential for embryonic development. Protects from radiation and oxidative damage.
- Gene Name:
- GPX4
- Uniprot ID:
- P36969
- Molecular Weight:
- 22174.52 Da
References
- Kraus RJ, Ganther HE: Reaction of cyanide with glutathione peroxidase. Biochem Biophys Res Commun. 1980 Oct 16;96(3):1116-22. [7437059 ]
- General Function:
- Ubiquinone binding
- Specific Function:
- Membrane-anchoring subunit of succinate dehydrogenase (SDH) that is involved in complex II of the mitochondrial electron transport chain and is responsible for transferring electrons from succinate to ubiquinone (coenzyme Q).
- Gene Name:
- SDHD
- Uniprot ID:
- O14521
- Molecular Weight:
- 17042.82 Da
References
- Ardelt BK, Borowitz JL, Isom GE: Brain lipid peroxidation and antioxidant protectant mechanisms following acute cyanide intoxication. Toxicology. 1989 Jun 1;56(2):147-54. [2734799 ]
- General Function:
- Succinate dehydrogenase activity
- Specific Function:
- Flavoprotein (FP) subunit of succinate dehydrogenase (SDH) that is involved in complex II of the mitochondrial electron transport chain and is responsible for transferring electrons from succinate to ubiquinone (coenzyme Q). Can act as a tumor suppressor.
- Gene Name:
- SDHA
- Uniprot ID:
- P31040
- Molecular Weight:
- 72690.975 Da
References
- Ardelt BK, Borowitz JL, Isom GE: Brain lipid peroxidation and antioxidant protectant mechanisms following acute cyanide intoxication. Toxicology. 1989 Jun 1;56(2):147-54. [2734799 ]
- General Function:
- Ubiquinone binding
- Specific Function:
- Iron-sulfur protein (IP) subunit of succinate dehydrogenase (SDH) that is involved in complex II of the mitochondrial electron transport chain and is responsible for transferring electrons from succinate to ubiquinone (coenzyme Q).
- Gene Name:
- SDHB
- Uniprot ID:
- P21912
- Molecular Weight:
- 31629.365 Da
References
- Ardelt BK, Borowitz JL, Isom GE: Brain lipid peroxidation and antioxidant protectant mechanisms following acute cyanide intoxication. Toxicology. 1989 Jun 1;56(2):147-54. [2734799 ]
- General Function:
- Succinate dehydrogenase activity
- Specific Function:
- Membrane-anchoring subunit of succinate dehydrogenase (SDH) that is involved in complex II of the mitochondrial electron transport chain and is responsible for transferring electrons from succinate to ubiquinone (coenzyme Q).
- Gene Name:
- SDHC
- Uniprot ID:
- Q99643
- Molecular Weight:
- 18610.03 Da
References
- Ardelt BK, Borowitz JL, Isom GE: Brain lipid peroxidation and antioxidant protectant mechanisms following acute cyanide intoxication. Toxicology. 1989 Jun 1;56(2):147-54. [2734799 ]
- General Function:
- Zinc ion binding
- Specific Function:
- Destroys radicals which are normally produced within the cells and which are toxic to biological systems.
- Gene Name:
- SOD1
- Uniprot ID:
- P00441
- Molecular Weight:
- 15935.685 Da
References
- Lee WG, Hwang JH, Na BK, Cho JH, Lee HW, Cho SH, Kong Y, Song CY, Kim TS: Functional expression of a recombinant copper/zinc superoxide dismutase of filarial nematode, Brugia malayi. J Parasitol. 2005 Feb;91(1):205-8. [15856906 ]
- General Function:
- Protein homodimerization activity
- Specific Function:
- This is a copper-containing oxidase that functions in the formation of pigments such as melanins and other polyphenolic compounds. Catalyzes the rate-limiting conversions of tyrosine to DOPA, DOPA to DOPA-quinone and possibly 5,6-dihydroxyindole to indole-5,6 quinone.
- Gene Name:
- TYR
- Uniprot ID:
- P14679
- Molecular Weight:
- 60392.69 Da
References
- Laufer Z, Beckett RP, Minibayeva FV: Co-occurrence of the multicopper oxidases tyrosinase and laccase in lichens in sub-order peltigerineae. Ann Bot. 2006 Nov;98(5):1035-42. Epub 2006 Sep 1. [16950829 ]
- General Function:
- Cholesterol binding
- Specific Function:
- Can bind protoporphyrin IX and may play a role in the transport of porphyrins and heme (By similarity). Promotes the transport of cholesterol across mitochondrial membranes and may play a role in lipid metabolism (PubMed:24814875), but its precise physiological role is controversial. It is apparently not required for steroid hormone biosynthesis. Was initially identified as peripheral-type benzodiazepine receptor; can also bind isoquinoline carboxamides (PubMed:1847678).
- Gene Name:
- TSPO
- Uniprot ID:
- P30536
- Molecular Weight:
- 18827.81 Da
Binding/Activity Constants
| Type | Value | Assay Type | Assay Source |
|---|---|---|---|
| AC50 | 0.29 uM | NVS_MP_hPBR | Novascreen |
References
- Sipes NS, Martin MT, Kothiya P, Reif DM, Judson RS, Richard AM, Houck KA, Dix DJ, Kavlock RJ, Knudsen TB: Profiling 976 ToxCast chemicals across 331 enzymatic and receptor signaling assays. Chem Res Toxicol. 2013 Jun 17;26(6):878-95. doi: 10.1021/tx400021f. Epub 2013 May 16. [23611293 ]
- General Function:
- Zinc ion binding
- Specific Function:
- Nuclear hormone receptor. The steroid hormones and their receptors are involved in the regulation of eukaryotic gene expression and affect cellular proliferation and differentiation in target tissues. Ligand-dependent nuclear transactivation involves either direct homodimer binding to a palindromic estrogen response element (ERE) sequence or association with other DNA-binding transcription factors, such as AP-1/c-Jun, c-Fos, ATF-2, Sp1 and Sp3, to mediate ERE-independent signaling. Ligand binding induces a conformational change allowing subsequent or combinatorial association with multiprotein coactivator complexes through LXXLL motifs of their respective components. Mutual transrepression occurs between the estrogen receptor (ER) and NF-kappa-B in a cell-type specific manner. Decreases NF-kappa-B DNA-binding activity and inhibits NF-kappa-B-mediated transcription from the IL6 promoter and displace RELA/p65 and associated coregulators from the promoter. Recruited to the NF-kappa-B response element of the CCL2 and IL8 promoters and can displace CREBBP. Present with NF-kappa-B components RELA/p65 and NFKB1/p50 on ERE sequences. Can also act synergistically with NF-kappa-B to activate transcription involving respective recruitment adjacent response elements; the function involves CREBBP. Can activate the transcriptional activity of TFF1. Also mediates membrane-initiated estrogen signaling involving various kinase cascades. Isoform 3 is involved in activation of NOS3 and endothelial nitric oxide production. Isoforms lacking one or several functional domains are thought to modulate transcriptional activity by competitive ligand or DNA binding and/or heterodimerization with the full length receptor. Essential for MTA1-mediated transcriptional regulation of BRCA1 and BCAS3. Isoform 3 can bind to ERE and inhibit isoform 1.
- Gene Name:
- ESR1
- Uniprot ID:
- P03372
- Molecular Weight:
- 66215.45 Da
Binding/Activity Constants
| Type | Value | Assay Type | Assay Source |
|---|---|---|---|
| AC50 | 0.67 uM | Tox21_ERa_BLA_Antagonist_ratio | Tox21/NCGC |
References
- Sipes NS, Martin MT, Kothiya P, Reif DM, Judson RS, Richard AM, Houck KA, Dix DJ, Kavlock RJ, Knudsen TB: Profiling 976 ToxCast chemicals across 331 enzymatic and receptor signaling assays. Chem Res Toxicol. 2013 Jun 17;26(6):878-95. doi: 10.1021/tx400021f. Epub 2013 May 16. [23611293 ]
- General Function:
- Transcriptional activator activity, rna polymerase ii distal enhancer sequence-specific binding
- Specific Function:
- Transcription activator that binds to antioxidant response (ARE) elements in the promoter regions of target genes. Important for the coordinated up-regulation of genes in response to oxidative stress. May be involved in the transcriptional activation of genes of the beta-globin cluster by mediating enhancer activity of hypersensitive site 2 of the beta-globin locus control region.
- Gene Name:
- NFE2L2
- Uniprot ID:
- Q16236
- Molecular Weight:
- 67825.9 Da
Binding/Activity Constants
| Type | Value | Assay Type | Assay Source |
|---|---|---|---|
| AC50 | 1.28 uM | ATG_NRF2_ARE_CIS | Attagene |
References
- Sipes NS, Martin MT, Kothiya P, Reif DM, Judson RS, Richard AM, Houck KA, Dix DJ, Kavlock RJ, Knudsen TB: Profiling 976 ToxCast chemicals across 331 enzymatic and receptor signaling assays. Chem Res Toxicol. 2013 Jun 17;26(6):878-95. doi: 10.1021/tx400021f. Epub 2013 May 16. [23611293 ]
- General Function:
- Oxidoreductase activity, acting on paired donors, with incorporation or reduction of molecular oxygen, reduced flavin or flavoprotein as one donor, and incorporation of one atom of oxygen
- Specific Function:
- Cytochromes P450 are a group of heme-thiolate monooxygenases. In liver microsomes, this enzyme is involved in an NADPH-dependent electron transport pathway. It oxidizes a variety of structurally unrelated compounds, including steroids, fatty acids, and xenobiotics. Most active in catalyzing 2-hydroxylation. Caffeine is metabolized primarily by cytochrome CYP1A2 in the liver through an initial N3-demethylation. Also acts in the metabolism of aflatoxin B1 and acetaminophen. Participates in the bioactivation of carcinogenic aromatic and heterocyclic amines. Catalizes the N-hydroxylation of heterocyclic amines and the O-deethylation of phenacetin.
- Gene Name:
- CYP1A2
- Uniprot ID:
- P05177
- Molecular Weight:
- 58293.76 Da
Binding/Activity Constants
| Type | Value | Assay Type | Assay Source |
|---|---|---|---|
| AC50 | 1.35 uM | NVS_ADME_hCYP1A2 | Novascreen |
References
- Sipes NS, Martin MT, Kothiya P, Reif DM, Judson RS, Richard AM, Houck KA, Dix DJ, Kavlock RJ, Knudsen TB: Profiling 976 ToxCast chemicals across 331 enzymatic and receptor signaling assays. Chem Res Toxicol. 2013 Jun 17;26(6):878-95. doi: 10.1021/tx400021f. Epub 2013 May 16. [23611293 ]
- General Function:
- Zinc ion binding
- Specific Function:
- Receptor for retinoic acid. Retinoic acid receptors bind as heterodimers to their target response elements in response to their ligands, all-trans or 9-cis retinoic acid, and regulate gene expression in various biological processes. The RAR/RXR heterodimers bind to the retinoic acid response elements (RARE) composed of tandem 5'-AGGTCA-3' sites known as DR1-DR5. The high affinity ligand for RXRs is 9-cis retinoic acid. RXRA serves as a common heterodimeric partner for a number of nuclear receptors. The RXR/RAR heterodimers bind to the retinoic acid response elements (RARE) composed of tandem 5'-AGGTCA-3' sites known as DR1-DR5. In the absence of ligand, the RXR-RAR heterodimers associate with a multiprotein complex containing transcription corepressors that induce histone acetylation, chromatin condensation and transcriptional suppression. On ligand binding, the corepressors dissociate from the receptors and associate with the coactivators leading to transcriptional activation. The RXRA/PPARA heterodimer is required for PPARA transcriptional activity on fatty acid oxidation genes such as ACOX1 and the P450 system genes.
- Gene Name:
- RXRA
- Uniprot ID:
- P19793
- Molecular Weight:
- 50810.835 Da
Binding/Activity Constants
| Type | Value | Assay Type | Assay Source |
|---|---|---|---|
| AC50 | 3.28 uM | OT_NURR1_NURR1RXRa_1440 | Odyssey Thera |
References
- Sipes NS, Martin MT, Kothiya P, Reif DM, Judson RS, Richard AM, Houck KA, Dix DJ, Kavlock RJ, Knudsen TB: Profiling 976 ToxCast chemicals across 331 enzymatic and receptor signaling assays. Chem Res Toxicol. 2013 Jun 17;26(6):878-95. doi: 10.1021/tx400021f. Epub 2013 May 16. [23611293 ]
- General Function:
- Zinc ion binding
- Specific Function:
- Ligand-activated transcription factor. Receptor for bile acids such as chenodeoxycholic acid, lithocholic acid and deoxycholic acid. Represses the transcription of the cholesterol 7-alpha-hydroxylase gene (CYP7A1) through the induction of NR0B2 or FGF19 expression, via two distinct mechanisms. Activates the intestinal bile acid-binding protein (IBABP). Activates the transcription of bile salt export pump ABCB11 by directly recruiting histone methyltransferase CARM1 to this locus.
- Gene Name:
- NR1H4
- Uniprot ID:
- Q96RI1
- Molecular Weight:
- 55913.915 Da
Binding/Activity Constants
| Type | Value | Assay Type | Assay Source |
|---|---|---|---|
| AC50 | 7.06 uM | OT_SRC1_SRC1FXR_0480 | Odyssey Thera |
References
- Sipes NS, Martin MT, Kothiya P, Reif DM, Judson RS, Richard AM, Houck KA, Dix DJ, Kavlock RJ, Knudsen TB: Profiling 976 ToxCast chemicals across 331 enzymatic and receptor signaling assays. Chem Res Toxicol. 2013 Jun 17;26(6):878-95. doi: 10.1021/tx400021f. Epub 2013 May 16. [23611293 ]