| Record Information |
|---|
| Version | 2.0 |
|---|
| Creation Date | 2009-07-21 20:27:42 UTC |
|---|
| Update Date | 2014-12-24 20:25:53 UTC |
|---|
| Accession Number | T3D2898 |
|---|
| Identification |
|---|
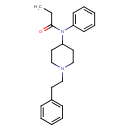
| Common Name | Fentanyl |
|---|
| Class | Small Molecule |
|---|
| Description | A potent narcotic analgesic, abuse of which leads to habituation or addiction. It is primarily a mu-opioid agonist. Fentanyl is also used as an adjunct to general anesthetics, and as an anesthetic for induction and maintenance. (From Martindale, The Extra Pharmacopoeia, 30th ed, p1078) |
|---|
| Compound Type | - Adjuvant
- Adjuvant, Anesthesia
- Amide
- Amine
- Analgesic
- Analgesic, Opioid
- Anesthetic
- Anesthetic, Intravenous
- Drug
- Metabolite
- Narcotic
- Opiate Agonist
- Organic Compound
- Synthetic Compound
|
|---|
| Chemical Structure | |
|---|
| Synonyms | | Synonym | | 1-Phenethyl-4-(N-phenylpropionamido)piperidine | | 1-Phenethyl-4-N-propionylanilinopiperidine | | Abstral | | Actiq | | Duragesic | | Durogesic | | Fentanest | | Fentanil | | Fentanila | | Fentanilo | | Fentanyl citrate | | Fentanylum | | Lazanda | | N-(1-Phenethyl-4-piperidinyl)-N-phenylpropionamide | | N-(1-Phenethyl-4-piperidyl)propionanilide | | N-(1-Phenethyl-piperidin-4-yl)-N-phenyl-propionamide | | N-(1-Phenethylpiperidin-4-yl)-N-phenylpropionamide | | N-Phenethyl-4-(N-propionylanilino)piperidine | | N-Phenyl-N-(1-(2-phenylethyl)-4-piperidinyl)propanamide | | Nasalfent | | Phentanyl | | Rapinyl | | Subsys |
|
|---|
| Chemical Formula | C22H28N2O |
|---|
| Average Molecular Mass | 336.471 g/mol |
|---|
| Monoisotopic Mass | 336.220 g/mol |
|---|
| CAS Registry Number | 437-38-7 |
|---|
| IUPAC Name | N-phenyl-N-[1-(2-phenylethyl)piperidin-4-yl]propanamide |
|---|
| Traditional Name | fentanyl |
|---|
| SMILES | CCC(=O)N(C1CCN(CCC2=CC=CC=C2)CC1)C1=CC=CC=C1 |
|---|
| InChI Identifier | InChI=1S/C22H28N2O/c1-2-22(25)24(20-11-7-4-8-12-20)21-14-17-23(18-15-21)16-13-19-9-5-3-6-10-19/h3-12,21H,2,13-18H2,1H3 |
|---|
| InChI Key | InChIKey=PJMPHNIQZUBGLI-UHFFFAOYSA-N |
|---|
| Chemical Taxonomy |
|---|
| Description | belongs to the class of organic compounds known as fentanyls. Fentanyls are compounds containing the fentanyl moiety or a derivative, which is based on a N-(1-(2-phenylethyl)-4-piperidinyl)-N-phenylpropanamide skeleton. |
|---|
| Kingdom | Organic compounds |
|---|
| Super Class | Organoheterocyclic compounds |
|---|
| Class | Piperidines |
|---|
| Sub Class | Fentanyls |
|---|
| Direct Parent | Fentanyls |
|---|
| Alternative Parents | |
|---|
| Substituents | - Fentanyl
- Phenethylamine
- Anilide
- Aralkylamine
- Monocyclic benzene moiety
- Benzenoid
- Tertiary carboxylic acid amide
- Amino acid or derivatives
- Carboxamide group
- Tertiary amine
- Tertiary aliphatic amine
- Carboxylic acid derivative
- Azacycle
- Organopnictogen compound
- Organooxygen compound
- Organonitrogen compound
- Organic oxygen compound
- Amine
- Carbonyl group
- Organic oxide
- Organic nitrogen compound
- Hydrocarbon derivative
- Aromatic heteromonocyclic compound
|
|---|
| Molecular Framework | Aromatic heteromonocyclic compounds |
|---|
| External Descriptors | |
|---|
| Biological Properties |
|---|
| Status | Detected and Not Quantified |
|---|
| Origin | Exogenous |
|---|
| Cellular Locations | |
|---|
| Biofluid Locations | Not Available |
|---|
| Tissue Locations | Not Available |
|---|
| Pathways | Not Available |
|---|
| Applications | |
|---|
| Biological Roles | |
|---|
| Chemical Roles | Not Available |
|---|
| Physical Properties |
|---|
| State | Solid |
|---|
| Appearance | White powder. |
|---|
| Experimental Properties | | Property | Value |
|---|
| Melting Point | 87.5°C | | Boiling Point | Not Available | | Solubility | 200 mg/L (at 25°C) | | LogP | 4.05 |
|
|---|
| Predicted Properties | |
|---|
| Spectra |
|---|
| Spectra | | Spectrum Type | Description | Splash Key | Deposition Date | View |
|---|
| Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | splash10-000i-5930000000-f7e6ba23728816e17e78 | 2017-09-01 | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | Not Available | 2021-10-12 | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | Not Available | 2021-10-12 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-QTOF , positive | splash10-000i-0009000000-62593b54c6285a566893 | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-QTOF , positive | splash10-000i-0209000000-d550d803c25d31570227 | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-QTOF , positive | splash10-000i-0901000000-09bd125eea134f8811a4 | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-QTOF , positive | splash10-000i-0900000000-2c34993eade063cb426f | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-QTOF , positive | splash10-001r-0900000000-beb72ea99dc9c125d29b | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-QFT , positive | splash10-000i-0009000000-bf70e4d857c26d30032f | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-QFT , positive | splash10-000i-0907000000-a9f5d0c422eff1474d38 | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-QFT , positive | splash10-000i-0900000000-0c03ba9ef576114a9503 | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-QFT , positive | splash10-0a4i-0900000000-d2a36b402c3a7424fab6 | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-QFT , positive | splash10-0a4i-1900000000-e0cbdfad80c306baabc1 | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-QFT , positive | splash10-0a4i-2900000000-ea551550ac6cbe6d3960 | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-QFT , positive | splash10-0pdi-8900000000-89b1a77e558ec74cf3ac | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-QFT , positive | splash10-0kdj-9400000000-358833c028a079789d0b | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-QFT , positive | splash10-0ufs-9200000000-ff2c211a084a5f124781 | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - 40V, Positive | splash10-000i-0900000000-da874163fadaa39d25ad | 2021-09-20 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - 30V, Positive | splash10-000i-0901000000-09bd125eea134f8811a4 | 2021-09-20 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - 10V, Positive | splash10-000i-0009000000-e126a8d88c9106409cac | 2021-09-20 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - 20V, Positive | splash10-000i-0209000000-d550d803c25d31570227 | 2021-09-20 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - 50V, Positive | splash10-001r-0900000000-2ed12cf85c72a16b8cdb | 2021-09-20 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-000i-1239000000-6f316602b5d12ff1b1b9 | 2016-08-03 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-0a4r-4942000000-703ee8a328127a82245c | 2016-08-03 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-0a4i-4900000000-c38eaa80ecf3e4334782 | 2016-08-03 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-000i-0009000000-303253b936dc7fea5dd0 | 2016-08-03 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-0550-3498000000-cbae1ca8da3e5b395fa4 | 2016-08-03 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-05dl-6930000000-8603b3c4374827ab0621 | 2016-08-03 | View Spectrum | | MS | Mass Spectrum (Electron Ionization) | splash10-0002-9850000000-7ce3a6f35f5dee98185f | 2014-09-20 | View Spectrum |
|
|---|
| Toxicity Profile |
|---|
| Route of Exposure | Epidural; parental (transdermal, intramuscular). |
|---|
| Mechanism of Toxicity | Opiate receptors are coupled with G-protein receptors and function as both positive and negative regulators of synaptic transmission via G-proteins that activate effector proteins. Binding of the opiate stimulates the exchange of GTP for GDP on the G-protein complex. As the effector system is adenylate cyclase and cAMP located at the inner surface of the plasma membrane, opioids decrease intracellular cAMP by inhibiting adenylate cyclase. Subsequently, the release of nociceptive neurotransmitters such as substance P, GABA, dopamine, acetylcholine and noradrenaline is inhibited. Opioids also inhibit the release of vasopressin, somatostatin, insulin and glucagon. Fentanyl's analgesic activity is, most likely, due to its conversion to morphine. Opioids close N-type voltage-operated calcium channels (OP2-receptor agonist) and open calcium-dependent inwardly rectifying potassium channels (OP3 and OP1 receptor agonist). This results in hypopolarization and reduced neuronal excitability. |
|---|
| Metabolism | Fentanyl is metabolized primarily via human cytochrome P450 3A4 isoenzyme system.
Route of Elimination: Fentanyl is metabolized primarily via human cytochrome P450 3A4 isoenzyme system and mostly eliminated in urine. Within 72 hours of IV fentanyl administration, approximately 75% of the dose is excreted in urine, mostly as metabolites with less than 10% representing unchanged drug.
Half Life: 7 hours (range 3-12) |
|---|
| Toxicity Values | LD50: 3.1 mg/kg (rat)
LD50: 0.03 mg/kg (monkeys) |
|---|
| Lethal Dose | Not Available |
|---|
| Carcinogenicity (IARC Classification) | No indication of carcinogenicity to humans (not listed by IARC). |
|---|
| Uses/Sources | For the treatment of cancer patients with severe pain that breaks through their regular narcotic therapy. Used as an inhalation anesthetic. (6) |
|---|
| Minimum Risk Level | Not Available |
|---|
| Health Effects | Medical problems can include congested lungs, liver disease, tetanus, infection of the heart valves, skin abscesses, anemia and pneumonia. Death can occur from overdose. |
|---|
| Symptoms | More common symptoms are dizziness, light-headedness, or feeling faint; drowsiness; nausea or vomiting; unusual tiredness or weakness. Less common or rare ones are blurred or double vision or other vision problems; confusion; constipation; convulsions (seizures); difficult or painful urination; mental depression; shortness of breath, trouble in breathing, tightness in the chest, or wheezing; skin rash, hives, or itching; unusual excitement (6) |
|---|
| Treatment | For the management of hypoventilation, immediate countermeasures include removing the Fentanyl and physically or verbally stimulating the patient. These actions can be followed by administration of a specific narcotic antagonist such as naloxone. The duration of hypoventilation following an overdose may be longer than the effects of the narcotic antagonist's action (the half-life of naloxone ranges from 30 to 81 minutes). The interval between IV antagonist doses should be carefully chosen because of the possibility of re-narcotization after system removal; repeated administration of naloxone may be necessary. Reversal of the narcotic effect may result in acute onset of pain and the release of catecholamines. Always ensure a patent airway is established and maintained, administer oxygen and assist or control respiration as indicated and use an oropharyngeal airway or endotracheal tube if necessary. Adequate body temperature and fluid intake should be maintained. (8) |
|---|
| Normal Concentrations |
|---|
| Not Available |
|---|
| Abnormal Concentrations |
|---|
| Not Available |
|---|
| External Links |
|---|
| DrugBank ID | DB00813 |
|---|
| HMDB ID | HMDB14951 |
|---|
| PubChem Compound ID | 3345 |
|---|
| ChEMBL ID | CHEMBL596 |
|---|
| ChemSpider ID | 3228 |
|---|
| KEGG ID | Not Available |
|---|
| UniProt ID | Not Available |
|---|
| OMIM ID | |
|---|
| ChEBI ID | 119915 |
|---|
| BioCyc ID | Not Available |
|---|
| CTD ID | Not Available |
|---|
| Stitch ID | Fentanyl |
|---|
| PDB ID | Not Available |
|---|
| ACToR ID | Not Available |
|---|
| Wikipedia Link | Fentanyl |
|---|
| References |
|---|
| Synthesis Reference | Mark Rubino, “Process of making fentanyl intermediates.” U.S. Patent US20060100438, issued May 11, 2006. |
|---|
| MSDS | Link |
|---|
| General References | - Van Bever WF, Niemegeers CJ, Janssen PA: Synthetic analgesics. Synthesis and pharmacology of the diastereoisomers of N-(3-methyl-1-(2-phenylethyl)-4-piperidyl)-N-phenylpropanamide and N-(3-methyl-1-(1-methyl-2-phenylethyl)-4-piperidyl)-N-phenylpropanamide. J Med Chem. 1974 Oct;17(10):1047-51. [4420811 ]
- Messina J, Darwish M, Fine PG: Fentanyl buccal tablet. Drugs Today (Barc). 2008 Jan;44(1):41-54. doi: 10.1358/dot.2008.44.1.1178469. [18301803 ]
- Taylor DR: Fentanyl buccal tablet: rapid relief from breakthrough pain. Expert Opin Pharmacother. 2007 Dec;8(17):3043-51. [18001263 ]
- Simpson DM, Messina J, Xie F, Hale M: Fentanyl buccal tablet for the relief of breakthrough pain in opioid-tolerant adult patients with chronic neuropathic pain: a multicenter, randomized, double-blind, placebo-controlled study. Clin Ther. 2007 Apr;29(4):588-601. [17617282 ]
- Martindale. The Extra Pharmacopoeia, 30th ed.
- Drugs.com [Link]
- Drugs.com [Link]
- RxList: The Internet Drug Index (2009). [Link]
|
|---|
| Gene Regulation |
|---|
| Up-Regulated Genes | | Gene | Gene Symbol | Gene ID | Interaction | Chromosome | Details |
|---|
|
|---|
| Down-Regulated Genes | | Gene | Gene Symbol | Gene ID | Interaction | Chromosome | Details |
|---|
|
|---|