| Record Information |
|---|
| Version | 2.0 |
|---|
| Creation Date | 2009-07-23 18:26:13 UTC |
|---|
| Update Date | 2014-12-24 20:25:58 UTC |
|---|
| Accession Number | T3D3094 |
|---|
| Identification |
|---|
| Common Name | Oleandrin |
|---|
| Class | Small Molecule |
|---|
| Description | Oleandrin is a plant toxin found in Oleander (Nerium oleander). It causes both gastrointestinal and cardiac effects. (2) |
|---|
| Compound Type | - Ester
- Ether
- Lachrymator
- Natural Compound
- Organic Compound
- Plant Toxin
|
|---|
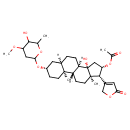
| Chemical Structure | |
|---|
| Synonyms | | Synonym | | Corrigen | | Foliandrin |
|
|---|
| Chemical Formula | C32H48O9 |
|---|
| Average Molecular Mass | 576.718 g/mol |
|---|
| Monoisotopic Mass | 576.330 g/mol |
|---|
| CAS Registry Number | 465-16-7 |
|---|
| IUPAC Name | (1S,2S,5S,7R,10R,11S,13S,14R,15R)-11-hydroxy-5-[(5-hydroxy-4-methoxy-6-methyloxan-2-yl)oxy]-2,15-dimethyl-14-(5-oxo-2,5-dihydrofuran-3-yl)tetracyclo[8.7.0.0²,⁷.0¹¹,¹⁵]heptadecan-13-yl acetate |
|---|
| Traditional Name | (1S,2S,5S,7R,10R,11S,13S,14R,15R)-11-hydroxy-5-[(5-hydroxy-4-methoxy-6-methyloxan-2-yl)oxy]-2,15-dimethyl-14-(5-oxo-2H-furan-3-yl)tetracyclo[8.7.0.0²,⁷.0¹¹,¹⁵]heptadecan-13-yl acetate |
|---|
| SMILES | [H][C@]12CC[C@]3([H])[C@]([H])(CC[C@]4(C)[C@H]([C@H](C[C@]34O)OC(C)=O)C3=CC(=O)OC3)[C@@]1(C)CC[C@@H](C2)OC1CC(OC)C(O)C(C)O1 |
|---|
| InChI Identifier | InChI=1S/C32H48O9/c1-17-29(35)24(37-5)14-27(39-17)41-21-8-10-30(3)20(13-21)6-7-23-22(30)9-11-31(4)28(19-12-26(34)38-16-19)25(40-18(2)33)15-32(23,31)36/h12,17,20-25,27-29,35-36H,6-11,13-16H2,1-5H3/t17?,20-,21+,22+,23-,24?,25+,27?,28+,29?,30+,31-,32+/m1/s1 |
|---|
| InChI Key | InChIKey=JLPDBLFIVFSOCC-UJQZKUKMSA-N |
|---|
| Chemical Taxonomy |
|---|
| Description | belongs to the class of organic compounds known as cardenolide glycosides and derivatives. Cardenolide glycosides and derivatives are compounds containing a carbohydrate glycosidically bound to the cardenolide moiety. |
|---|
| Kingdom | Organic compounds |
|---|
| Super Class | Lipids and lipid-like molecules |
|---|
| Class | Steroids and steroid derivatives |
|---|
| Sub Class | Steroid lactones |
|---|
| Direct Parent | Cardenolide glycosides and derivatives |
|---|
| Alternative Parents | |
|---|
| Substituents | - Cardanolide-glycoside
- Steroidal glycoside
- Steroid ester
- Hydroxysteroid
- 14-hydroxysteroid
- Hexose monosaccharide
- O-glycosyl compound
- Glycosyl compound
- Oxane
- Monosaccharide
- Dicarboxylic acid or derivatives
- 2-furanone
- Alpha,beta-unsaturated carboxylic ester
- Enoate ester
- Tertiary alcohol
- Dihydrofuran
- Cyclic alcohol
- Secondary alcohol
- Lactone
- Carboxylic acid ester
- Oxacycle
- Organoheterocyclic compound
- Ether
- Dialkyl ether
- Carboxylic acid derivative
- Acetal
- Organic oxygen compound
- Organic oxide
- Hydrocarbon derivative
- Organooxygen compound
- Carbonyl group
- Alcohol
- Aliphatic heteropolycyclic compound
|
|---|
| Molecular Framework | Aliphatic heteropolycyclic compounds |
|---|
| External Descriptors | Not Available |
|---|
| Biological Properties |
|---|
| Status | Detected and Not Quantified |
|---|
| Origin | Exogenous |
|---|
| Cellular Locations | |
|---|
| Biofluid Locations | Not Available |
|---|
| Tissue Locations | Not Available |
|---|
| Pathways | Not Available |
|---|
| Applications | Not Available |
|---|
| Biological Roles | Not Available |
|---|
| Chemical Roles | Not Available |
|---|
| Physical Properties |
|---|
| State | Solid |
|---|
| Appearance | White powder. |
|---|
| Experimental Properties | | Property | Value |
|---|
| Melting Point | Not Available | | Boiling Point | Not Available | | Solubility | Not Available | | LogP | Not Available |
|
|---|
| Predicted Properties | |
|---|
| Spectra |
|---|
| Spectra | | Spectrum Type | Description | Splash Key | Deposition Date | View |
|---|
| Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-014j-0003490000-5a2b48a3b1da1591116b | 2016-06-03 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-014j-0309620000-44ad0bc476f043dbc31f | 2016-06-03 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-0udj-3709000000-4e1f0232082d24073d3f | 2016-06-03 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-0059-1000290000-16abcd2098c038fce667 | 2016-08-03 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-0a59-4101930000-02f6f045a6ce32b65410 | 2016-08-03 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-090u-4001900000-19864fff559b4884dbf4 | 2016-08-03 | View Spectrum |
|
|---|
| Toxicity Profile |
|---|
| Route of Exposure | Oral (ingestion) (4) ; dermal (4) |
|---|
| Mechanism of Toxicity | Oleandrin likely exerts its toxic effects by inhibiting Na+,K+-ATPase. (1) |
|---|
| Metabolism | Not Available |
|---|
| Toxicity Values | Not Available |
|---|
| Lethal Dose | Not Available |
|---|
| Carcinogenicity (IARC Classification) | No indication of carcinogenicity to humans (not listed by IARC). |
|---|
| Uses/Sources | Oleandrin is a plant toxin found in Oleander (Nerium oleander). (2) |
|---|
| Minimum Risk Level | Not Available |
|---|
| Health Effects | Oleandrin causes both gastrointestinal and cardiac effects. Reactions to poisonings from this plant can also affect the central nervous system, possibly resulting in coma that can lead to death. (2) |
|---|
| Symptoms | The gastrointestinal effects can consist of nausea and vomiting, excess salivation, abdominal pain, diarrhea, and colic. Cardiac reactions consist of irregular heart rate, and extremities may become pale and cold due to poor or irregular circulation. Reactions to poisonings from this plant can also affect the central nervous system. These symptoms can include drowsiness, tremors or shaking of the muscles, seizures, collapse, and even coma that can lead to death. Oleander sap can cause skin irritations, severe eye inflammation and irritation, and allergy reactions characterized by dermatitis. (2) |
|---|
| Treatment | Oleandrin poisoning is treated by induced vomiting, gastric lavage, and/or administration of activated charcoal to prevent further absorption. In severe cases, the antidote Digoxin Immune Fab may be used. (3) |
|---|
| Normal Concentrations |
|---|
| Not Available |
|---|
| Abnormal Concentrations |
|---|
| Not Available |
|---|
| External Links |
|---|
| DrugBank ID | Not Available |
|---|
| HMDB ID | Not Available |
|---|
| PubChem Compound ID | 261943 |
|---|
| ChEMBL ID | Not Available |
|---|
| ChemSpider ID | 229992 |
|---|
| KEGG ID | Not Available |
|---|
| UniProt ID | Not Available |
|---|
| OMIM ID | |
|---|
| ChEBI ID | Not Available |
|---|
| BioCyc ID | Not Available |
|---|
| CTD ID | Not Available |
|---|
| Stitch ID | Oleandrin |
|---|
| PDB ID | Not Available |
|---|
| ACToR ID | Not Available |
|---|
| Wikipedia Link | Not Available |
|---|
| References |
|---|
| Synthesis Reference | Not Available |
|---|
| MSDS | T3D3094.pdf |
|---|
| General References | - Jortani SA, Helm RA, Valdes R Jr: Inhibition of Na,K-ATPase by oleandrin and oleandrigenin, and their detection by digoxin immunoassays. Clin Chem. 1996 Oct;42(10):1654-8. [8855150 ]
- Wikipedia. Oleandrin. Last Updated 8 July 2009. [Link]
- Wikipedia. Nerium oleander. Last Updated 9 July 2009. [Link]
- Wikipedia. Phytotoxin. Last Updated 7 August 2009. [Link]
|
|---|
| Gene Regulation |
|---|
| Up-Regulated Genes | Not Available |
|---|
| Down-Regulated Genes | Not Available |
|---|