| Record Information |
|---|
| Version | 2.0 |
|---|
| Creation Date | 2009-07-30 17:59:04 UTC |
|---|
| Update Date | 2014-12-24 20:26:07 UTC |
|---|
| Accession Number | T3D3526 |
|---|
| Identification |
|---|
| Common Name | Pentamidine |
|---|
| Class | Small Molecule |
|---|
| Description | Pentamidine is only found in individuals that have used or taken this drug. It is an antiprotozoal agent effective in trypanosomiasis, leishmaniasis, and some fungal infections; used in treatment of pneumocystis pneumonia in HIV-infected patients. It may cause diabetes mellitus, central nervous system damage, and other toxic effects. [PubChem] The mode of action of pentamidine is not fully understood. It is thought that the drug interferes with nuclear metabolism producing inhibition of the synthesis of DNA, RNA, phospholipids, and proteins. |
|---|
| Compound Type | - Amide
- Amine
- Antifungal Agent
- Antiprotozoal Agent
- Drug
- Ether
- Metabolite
- Organic Compound
- Synthetic Compound
- Trypanocidal Agent
|
|---|
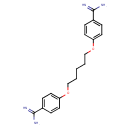
| Chemical Structure | |
|---|
| Synonyms | | Synonym | | 1,3-Bis(4-amidinophenoxy)pentane | | 1,5-Bis(4-amidinophenoxy)pentane | | 4, 4'-Diamidinodiphenoxypentane | | 4,4'-(1,5-Pentanediylbis(oxy))bis-benzenecarboximidamide | | 4,4'-(Pentamethylenedioxy)dibenzamidine | | 4,4'-Diamidinodiphenoxypentane | | NebuPent | | P,P'-(pentamethylenedioxy)dibenzamidine | | Pentacarinat | | Pentacrinat | | Pentam | | Pentam 300 | | Pentamide | | Pentamidin | | Pentamidina | | Pentamidindiisetionat | | Pentamidine Isethionate | | Pentamidinum | | Pneumopent | | PNT |
|
|---|
| Chemical Formula | C19H24N4O2 |
|---|
| Average Molecular Mass | 340.420 g/mol |
|---|
| Monoisotopic Mass | 340.190 g/mol |
|---|
| CAS Registry Number | 100-33-4 |
|---|
| IUPAC Name | 4-{[5-(4-carbamimidoylphenoxy)pentyl]oxy}benzene-1-carboximidamide |
|---|
| Traditional Name | pentamidine |
|---|
| SMILES | NC(=N)C1=CC=C(OCCCCCOC2=CC=C(C=C2)C(N)=N)C=C1 |
|---|
| InChI Identifier | InChI=1S/C19H24N4O2/c20-18(21)14-4-8-16(9-5-14)24-12-2-1-3-13-25-17-10-6-15(7-11-17)19(22)23/h4-11H,1-3,12-13H2,(H3,20,21)(H3,22,23) |
|---|
| InChI Key | InChIKey=XDRYMKDFEDOLFX-UHFFFAOYSA-N |
|---|
| Chemical Taxonomy |
|---|
| Description | belongs to the class of organic compounds known as phenol ethers. These are aromatic compounds containing an ether group substituted with a benzene ring. |
|---|
| Kingdom | Organic compounds |
|---|
| Super Class | Benzenoids |
|---|
| Class | Phenol ethers |
|---|
| Sub Class | Not Available |
|---|
| Direct Parent | Phenol ethers |
|---|
| Alternative Parents | |
|---|
| Substituents | - Phenoxy compound
- Phenol ether
- Alkyl aryl ether
- Monocyclic benzene moiety
- Carboximidamide
- Ether
- Carboxylic acid amidine
- Amidine
- Organic nitrogen compound
- Organic oxygen compound
- Organopnictogen compound
- Hydrocarbon derivative
- Organooxygen compound
- Organonitrogen compound
- Aromatic homomonocyclic compound
|
|---|
| Molecular Framework | Aromatic homomonocyclic compounds |
|---|
| External Descriptors | |
|---|
| Biological Properties |
|---|
| Status | Detected and Not Quantified |
|---|
| Origin | Exogenous |
|---|
| Cellular Locations | |
|---|
| Biofluid Locations | Not Available |
|---|
| Tissue Locations | Not Available |
|---|
| Pathways | Not Available |
|---|
| Applications | |
|---|
| Biological Roles | |
|---|
| Chemical Roles | Not Available |
|---|
| Physical Properties |
|---|
| State | Solid |
|---|
| Appearance | White powder. |
|---|
| Experimental Properties | | Property | Value |
|---|
| Melting Point | 186.0°C (decomposes) | | Boiling Point | Not Available | | Solubility | Complete | | LogP | 4 |
|
|---|
| Predicted Properties | |
|---|
| Spectra |
|---|
| Spectra | | Spectrum Type | Description | Splash Key | Deposition Date | View |
|---|
| Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | splash10-0a4u-0902000000-3844069e5553bbf605a6 | 2017-09-01 | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | Not Available | 2021-10-12 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-QTOF , positive | splash10-0006-0009000000-e30f537a55b02f867429 | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-QTOF , positive | splash10-0006-0009000000-18ffa88d8e83e0c224b3 | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-QTOF , positive | splash10-00di-0309000000-22a206e2fffa6ce5cf2f | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-QTOF , positive | splash10-00dr-0900000000-d135e9c44dbf5da10735 | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-QTOF , positive | splash10-00di-0900000000-d01e29cdf614c3660180 | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-QQ , positive | splash10-0006-0009000000-1499530d0d768e480fd7 | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-QQ , positive | splash10-0006-0009000000-3c60569ced478a95001f | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-QQ , positive | splash10-00di-0209000000-58a168dde59140860009 | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-QQ , positive | splash10-00dr-0901000000-7cd36861b140e4e5a6a2 | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-QQ , positive | splash10-00di-0900000000-60dfcd75868e869bcbf7 | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-IT , positive | splash10-00di-0009000000-6677c6c06aa266d05ff1 | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - 40V, Positive | splash10-00dr-0900000000-210e86ac8fe8716be765 | 2021-09-20 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - 50V, Positive | splash10-00di-0900000000-d01e29cdf614c3660180 | 2021-09-20 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - 10V, Positive | splash10-0006-0009000000-e30f537a55b02f867429 | 2021-09-20 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - 20V, Positive | splash10-0006-0009000000-18ffa88d8e83e0c224b3 | 2021-09-20 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - 40V, Positive | splash10-00dr-0900000000-d135e9c44dbf5da10735 | 2021-09-20 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - 30V, Positive | splash10-00di-0309000000-22a206e2fffa6ce5cf2f | 2021-09-20 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - 50V, Positive | splash10-00di-0900000000-a005235fbf0235ec242a | 2021-09-20 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - 30V, Positive | splash10-00di-0309000000-ba9999e56127535cc0be | 2021-09-20 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-000i-0409000000-15c819a9688c0d5dd79f | 2016-08-03 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-000i-0904000000-cc51d663fb7c9ccf5339 | 2016-08-03 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-000i-4900000000-1a2483dd0fa6cb115ce5 | 2016-08-03 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-000i-0119000000-242a25d74bfc2340c519 | 2021-10-11 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-000i-4239000000-30cf7c8a257fbb023ef8 | 2021-10-11 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-0006-9100000000-9dbfd037d9a4546ce49f | 2021-10-11 | View Spectrum |
|
|---|
| Toxicity Profile |
|---|
| Route of Exposure | Absorbed poorly through the gastrointestinal tract and is usually administered parenterally. |
|---|
| Mechanism of Toxicity | The mode of action of pentamidine is not fully understood. It is thought that the drug interferes with nuclear metabolism producing inhibition of the synthesis of DNA, RNA, phospholipids, and proteins. |
|---|
| Metabolism | Hepatic.
Half Life: 9.1-13.2 hours |
|---|
| Toxicity Values | Not Available |
|---|
| Lethal Dose | Not Available |
|---|
| Carcinogenicity (IARC Classification) | No indication of carcinogenicity to humans (not listed by IARC). |
|---|
| Uses/Sources | For the treatment of pneumonia due to Pneumocystis carinii. |
|---|
| Minimum Risk Level | Not Available |
|---|
| Health Effects | Not Available |
|---|
| Symptoms | Symptoms of overdose include pain, nausea, anorexia, hypotension, fever, rash, bad taste in mouth, confusion/hallucinations, dizziness, and diarrhea. |
|---|
| Treatment | Not Available |
|---|
| Normal Concentrations |
|---|
| Not Available |
|---|
| Abnormal Concentrations |
|---|
| Not Available |
|---|
| External Links |
|---|
| DrugBank ID | DB00738 |
|---|
| HMDB ID | HMDB14876 |
|---|
| PubChem Compound ID | 4735 |
|---|
| ChEMBL ID | CHEMBL55 |
|---|
| ChemSpider ID | 4573 |
|---|
| KEGG ID | C07420 |
|---|
| UniProt ID | Not Available |
|---|
| OMIM ID | |
|---|
| ChEBI ID | 45081 |
|---|
| BioCyc ID | Not Available |
|---|
| CTD ID | Not Available |
|---|
| Stitch ID | Pentamidine |
|---|
| PDB ID | PNT |
|---|
| ACToR ID | Not Available |
|---|
| Wikipedia Link | Pentamidine |
|---|
| References |
|---|
| Synthesis Reference | Not Available |
|---|
| MSDS | Link |
|---|
| General References | - Nguewa PA, Fuertes MA, Cepeda V, Iborra S, Carrion J, Valladares B, Alonso C, Perez JM: Pentamidine is an antiparasitic and apoptotic drug that selectively modifies ubiquitin. Chem Biodivers. 2005 Oct;2(10):1387-400. [17191940 ]
- Drugs.com [Link]
|
|---|
| Gene Regulation |
|---|
| Up-Regulated Genes | | Gene | Gene Symbol | Gene ID | Interaction | Chromosome | Details |
|---|
|
|---|
| Down-Regulated Genes | | Gene | Gene Symbol | Gene ID | Interaction | Chromosome | Details |
|---|
|
|---|