| Record Information |
|---|
| Version | 2.0 |
|---|
| Creation Date | 2014-08-29 06:51:04 UTC |
|---|
| Update Date | 2014-12-24 20:26:48 UTC |
|---|
| Accession Number | T3D4409 |
|---|
| Identification |
|---|
| Common Name | Xanthine |
|---|
| Class | Small Molecule |
|---|
| Description | Xanthine is a purine base found in most body tissues and fluids, certain plants, and some urinary calculi. It is an intermediate in the degradation of adenosine monophosphate to uric acid, being formed by oxidation of hypoxanthine. The methylated xanthine compounds caffeine, theobromine, and theophylline and their derivatives are used in medicine for their bronchodilator effects. (Dorland, 28th ed.). |
|---|
| Compound Type | - Animal Toxin
- Food Toxin
- Metabolite
- Natural Compound
- Organic Compound
|
|---|
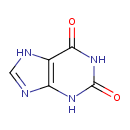
| Chemical Structure | |
|---|
| Synonyms | | Synonym | | 1H-Purine-2,6-diol | | 2,6(1,3)-Purinedion | | 2,6-Dihydroxypurine | | 2,6-Dioxopurine | | 3,7-Dihydro-1H-purine-2,6-dione | | 3,7-Dihydropurine-2,6-dione | | 9H-Purine-2,6(1H,3H)-dione | | 9H-Purine-2,6-diol | | Dioxopurine | | Isoxanthine | | Pseudoxanthine | | Purine-2,6(1H,3H)-dione | | Purine-2,6-diol | | Xanthic oxide | | Xanthin |
|
|---|
| Chemical Formula | C5H4N4O2 |
|---|
| Average Molecular Mass | 152.111 g/mol |
|---|
| Monoisotopic Mass | 152.033 g/mol |
|---|
| CAS Registry Number | 69-89-6 |
|---|
| IUPAC Name | 2,3,6,7-tetrahydro-1H-purine-2,6-dione |
|---|
| Traditional Name | xanthine |
|---|
| SMILES | OC1=NC2=C(N=CN2)C(O)=N1 |
|---|
| InChI Identifier | InChI=1S/C5H4N4O2/c10-4-2-3(7-1-6-2)8-5(11)9-4/h1H,(H3,6,7,8,9,10,11) |
|---|
| InChI Key | InChIKey=LRFVTYWOQMYALW-UHFFFAOYSA-N |
|---|
| Chemical Taxonomy |
|---|
| Description | belongs to the class of organic compounds known as xanthines. These are purine derivatives with a ketone group conjugated at carbons 2 and 6 of the purine moiety. |
|---|
| Kingdom | Organic compounds |
|---|
| Super Class | Organoheterocyclic compounds |
|---|
| Class | Imidazopyrimidines |
|---|
| Sub Class | Purines and purine derivatives |
|---|
| Direct Parent | Xanthines |
|---|
| Alternative Parents | |
|---|
| Substituents | - Xanthine
- 6-oxopurine
- Purinone
- Alkaloid or derivatives
- Pyrimidone
- Pyrimidine
- Azole
- Imidazole
- Heteroaromatic compound
- Vinylogous amide
- Lactam
- Urea
- Azacycle
- Hydrocarbon derivative
- Organic oxide
- Organooxygen compound
- Organonitrogen compound
- Organic nitrogen compound
- Organopnictogen compound
- Organic oxygen compound
- Aromatic heteropolycyclic compound
|
|---|
| Molecular Framework | Aromatic heteropolycyclic compounds |
|---|
| External Descriptors | |
|---|
| Biological Properties |
|---|
| Status | Detected and Not Quantified |
|---|
| Origin | Endogenous |
|---|
| Cellular Locations | - Cytoplasm
- Extracellular
- Peroxisome
|
|---|
| Biofluid Locations | Not Available |
|---|
| Tissue Locations | - Bladder
- Epidermis
- Fibroblasts
- Intestine
- Kidney
- Liver
- Prostate
- Skeletal Muscle
- Testes
|
|---|
| Pathways | | Name | SMPDB Link | KEGG Link |
|---|
| Purine Metabolism | SMP00050 | map00230 | | Xanthinuria type I | SMP00512 | Not Available | | Xanthinuria type II | SMP00513 | Not Available | | Molybdenium Cofactor Deficiency | SMP00203 | Not Available | | Xanthine Dehydrogenase Deficiency (Xanthinuria) | SMP00220 | Not Available |
|
|---|
| Applications | Not Available |
|---|
| Biological Roles | |
|---|
| Chemical Roles | Not Available |
|---|
| Physical Properties |
|---|
| State | Solid |
|---|
| Appearance | White powder. |
|---|
| Experimental Properties | | Property | Value |
|---|
| Melting Point | > 300°C | | Boiling Point | Not Available | | Solubility | 0.069 mg/mL at 16°C; 9.5 mg/mL (sodium salt) | | LogP | -0.73 |
|
|---|
| Predicted Properties | |
|---|
| Spectra |
|---|
| Spectra | | Spectrum Type | Description | Splash Key | Deposition Date | View |
|---|
| GC-MS | GC-MS Spectrum - GC-EI-TOF (Pegasus III TOF-MS system, Leco; GC 6890, Agilent Technologies) (Non-derivatized) | splash10-0f6t-0924000000-9b80e0a2a60c73ca0180 | 2014-06-16 | View Spectrum | | GC-MS | GC-MS Spectrum - GC-EI-TOF (Non-derivatized) | splash10-0f6t-0924000000-9b80e0a2a60c73ca0180 | 2017-09-12 | View Spectrum | | GC-MS | GC-MS Spectrum - GC-EI-TOF (Non-derivatized) | splash10-0f6t-0924000000-30dc5892eecde860846a | 2017-09-12 | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | splash10-0kai-7900000000-2dc30b0fc4cff2239dbe | 2016-09-22 | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | Not Available | 2021-10-12 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - Quattro_QQQ 10V, Positive (Annotated) | splash10-0udi-0900000000-a70539989d121bfacee0 | 2012-07-24 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - Quattro_QQQ 25V, Positive (Annotated) | splash10-0a4i-6900000000-b047b06406308dbaeda8 | 2012-07-24 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - Quattro_QQQ 40V, Positive (Annotated) | splash10-0a4i-9300000000-ed480ed920c3e9b576ec | 2012-07-24 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-QTOF (UPLC Q-Tof Premier, Waters) , Negative | splash10-0zfr-0900000000-efb049914c9bce596267 | 2012-08-31 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-QTOF , negative | splash10-0zfr-0900000000-efb049914c9bce596267 | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - , negative | splash10-0udi-0900000000-5fee91293851bb02193e | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-IT , positive | splash10-000i-0900000000-4568a814903ff411923a | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - 40V, Negative | splash10-00kf-9000000000-e078a358156f5a04d4c1 | 2021-09-20 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - 20V, Negative | splash10-0a4i-3900000000-250f7dc30d6fd96a4ef6 | 2021-09-20 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - 20V, Positive | splash10-0ik9-0900000000-3a6dad2473c1654789f6 | 2021-09-20 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - 10V, Negative | splash10-0udi-0900000000-a4e9443b51c3ac2fc58b | 2021-09-20 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - 40V, Negative | splash10-00kf-9000000000-d23801a49488b1a5f0a2 | 2021-09-20 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - 20V, Negative | splash10-0a4i-4900000000-b177a2a36043e5e87efc | 2021-09-20 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - 35V, Negative | splash10-0udi-0900000000-cb3b35c3117b7f36bf5c | 2021-09-20 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - 10V, Negative | splash10-0zfr-0900000000-8d2a19e7c37d2cb7658f | 2021-09-20 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - 30V, Negative | splash10-05mo-9300000000-33d00ab2288aaa055517 | 2021-09-20 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - 20V, Positive | splash10-0ik9-1900000000-3a6dad2473c1654789f6 | 2021-09-20 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - 40V, Positive | splash10-001i-9000000000-37e76df2401a85967caa | 2021-09-20 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - 35V, Negative | splash10-0udi-0900000000-4132da06dda895afc3ab | 2021-09-20 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-0udi-0900000000-2e9e069e2df414aed037 | 2015-05-26 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-0w29-0900000000-fa52193346bc456d89e8 | 2015-05-26 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-0a5i-9400000000-bbf70998e8b7515cb440 | 2015-05-26 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-0udi-0900000000-566d663553ce4f0ec207 | 2015-05-27 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-0zfr-1900000000-d0a5d2c0f89f8d42d903 | 2015-05-27 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-0006-9000000000-351d9f8ee3470f911829 | 2015-05-27 | View Spectrum | | MS | Mass Spectrum (Electron Ionization) | splash10-0udi-7900000000-2d5ab5d5db8ff4981467 | 2014-09-20 | View Spectrum | | 1D NMR | 1H NMR Spectrum (1D, 500 MHz, H2O, experimental) | Not Available | 2012-12-04 | View Spectrum | | 1D NMR | 13C NMR Spectrum (1D, 100.7 MHz, DMSO-d6, experimental) | Not Available | 2014-09-23 | View Spectrum | | 1D NMR | 1H NMR Spectrum (1D, D2O, experimental) | Not Available | 2016-10-22 | View Spectrum | | 1D NMR | 13C NMR Spectrum (1D, D2O, experimental) | Not Available | 2016-10-22 | View Spectrum | | 1D NMR | 13C NMR Spectrum (1D, 100 MHz, D2O, predicted) | Not Available | 2021-09-29 | View Spectrum | | 1D NMR | 1H NMR Spectrum (1D, 100 MHz, D2O, predicted) | Not Available | 2021-09-29 | View Spectrum | | 1D NMR | 13C NMR Spectrum (1D, 1000 MHz, D2O, predicted) | Not Available | 2021-09-29 | View Spectrum | | 1D NMR | 1H NMR Spectrum (1D, 1000 MHz, D2O, predicted) | Not Available | 2021-09-29 | View Spectrum | | 1D NMR | 13C NMR Spectrum (1D, 200 MHz, D2O, predicted) | Not Available | 2021-09-29 | View Spectrum | | 1D NMR | 1H NMR Spectrum (1D, 200 MHz, D2O, predicted) | Not Available | 2021-09-29 | View Spectrum | | 1D NMR | 13C NMR Spectrum (1D, 300 MHz, D2O, predicted) | Not Available | 2021-09-29 | View Spectrum | | 1D NMR | 1H NMR Spectrum (1D, 300 MHz, D2O, predicted) | Not Available | 2021-09-29 | View Spectrum | | 1D NMR | 13C NMR Spectrum (1D, 400 MHz, D2O, predicted) | Not Available | 2021-09-29 | View Spectrum | | 1D NMR | 1H NMR Spectrum (1D, 400 MHz, D2O, predicted) | Not Available | 2021-09-29 | View Spectrum | | 1D NMR | 13C NMR Spectrum (1D, 500 MHz, D2O, predicted) | Not Available | 2021-09-29 | View Spectrum | | 1D NMR | 1H NMR Spectrum (1D, 500 MHz, D2O, predicted) | Not Available | 2021-09-29 | View Spectrum | | 1D NMR | 13C NMR Spectrum (1D, 600 MHz, D2O, predicted) | Not Available | 2021-09-29 | View Spectrum | | 1D NMR | 1H NMR Spectrum (1D, 600 MHz, D2O, predicted) | Not Available | 2021-09-29 | View Spectrum | | 1D NMR | 13C NMR Spectrum (1D, 700 MHz, D2O, predicted) | Not Available | 2021-09-29 | View Spectrum | | 1D NMR | 1H NMR Spectrum (1D, 700 MHz, D2O, predicted) | Not Available | 2021-09-29 | View Spectrum | | 1D NMR | 13C NMR Spectrum (1D, 800 MHz, D2O, predicted) | Not Available | 2021-09-29 | View Spectrum | | 1D NMR | 1H NMR Spectrum (1D, 800 MHz, D2O, predicted) | Not Available | 2021-09-29 | View Spectrum | | 1D NMR | 13C NMR Spectrum (1D, 900 MHz, D2O, predicted) | Not Available | 2021-09-29 | View Spectrum | | 1D NMR | 1H NMR Spectrum (1D, 900 MHz, D2O, predicted) | Not Available | 2021-09-29 | View Spectrum | | 2D NMR | [1H, 1H]-TOCSY. Unexported temporarily by An Chi on Oct 15, 2021 until json or nmrML file is generated. 2D NMR Spectrum (experimental) | Not Available | 2012-12-04 | View Spectrum | | 2D NMR | [1H, 13C]-HSQC NMR Spectrum (2D, 600 MHz, 100%_DMSO, experimental) | Not Available | 2012-12-05 | View Spectrum |
|
|---|
| Toxicity Profile |
|---|
| Route of Exposure | Endogenous, Ingestion, Dermal (contact) |
|---|
| Mechanism of Toxicity | Xanthine is a poorly soluble compound. As a result high concentrations of serum xanthine can lead to the formation of kidney stones (xanthine kidney stones) which can, over the long term, induce kidney failure. |
|---|
| Metabolism | Xanthine is readily converted to uric acid. The enzyme xanthine oxidase makes uric acid from xanthine and hypoxanthine, which in turn are produced from other purines. In humans and higher primates, uric acid is the final oxidation (breakdown) product of purine metabolism and is excreted in urine. |
|---|
| Toxicity Values | Not Available |
|---|
| Lethal Dose | Not Available |
|---|
| Carcinogenicity (IARC Classification) | No indication of carcinogenicity to humans (not listed by IARC). |
|---|
| Uses/Sources | Naturally produced by the body (endogenous). |
|---|
| Minimum Risk Level | Not Available |
|---|
| Health Effects | Chronically high concentrations of xanthine can lead to health problems such as renal failure and xanthine kidney stones, one of the rarest types of kidney stones. Chronically high levels of xanthine are associated with at least 4 inborn errors of metabolism including: Xanthinuria type I, Xanthuria type II, Molybdenium Cofactor Deficiency, and Xanthinuria. |
|---|
| Symptoms | May lead to arthropathy, myopathy, crystal nephropathy, urolithiasis, or renal failure. |
|---|
| Treatment | Chronic Exposure: Kidney dialysis is usually needed to relieve the symptoms of xanthine toxicity until normal kidney function can be restored.
Acute Exposure: EYES: irrigate opened eyes for several minutes under running water. INGESTION: do not induce vomiting. Rinse mouth with water (never give anything by mouth to an unconscious person). Seek immediate medical advice. |
|---|
| Normal Concentrations |
|---|
| Not Available |
|---|
| Abnormal Concentrations |
|---|
| Not Available |
|---|
| External Links |
|---|
| DrugBank ID | DB02134 |
|---|
| HMDB ID | HMDB00292 |
|---|
| PubChem Compound ID | 1188 |
|---|
| ChEMBL ID | CHEMBL1424 |
|---|
| ChemSpider ID | 1151 |
|---|
| KEGG ID | C00385 |
|---|
| UniProt ID | Not Available |
|---|
| OMIM ID | |
|---|
| ChEBI ID | 17712 |
|---|
| BioCyc ID | XANTHINE |
|---|
| CTD ID | Not Available |
|---|
| Stitch ID | Not Available |
|---|
| PDB ID | XAN |
|---|
| ACToR ID | Not Available |
|---|
| Wikipedia Link | Xanthine |
|---|
| References |
|---|
| Synthesis Reference | John P. Zikakis, “Preparation of high purity xanthine oxidase from bovine milk.” U.S. Patent US4172763, issued October 30, 1979. |
|---|
| MSDS | Link |
|---|
| General References | - Ihara H, Shino Y, Morita Y, Kawaguchi E, Hashizume N, Yoshida M: Is skeletal muscle damaged by the oxidative stress following anaerobic exercise? J Clin Lab Anal. 2001;15(5):239-43. [11574951 ]
- Niklasson F: Simultaneous liquid-chromatographic determination of hypoxanthine, xanthine, urate, and creatinine in cerebrospinal fluid, with direct injection. Clin Chem. 1983 Aug;29(8):1543-6. [6872216 ]
- Teeuwen HW, Elbers EL, van Rossum JM: Rapid and sensitive gas-chromatographic determination of caffeine in blood plasma, saliva, and xanthine beverages. Mol Biol Rep. 1991 Feb;15(1):1-7. [1875916 ]
- Castro-Gago M, Rodriguez IN, Rodriguez-Nunez A, Guitian JP, Rocamonde SL, Rodriguez-Segade S: Therapeutic criteria in hydrocephalic children. Childs Nerv Syst. 1989 Dec;5(6):361-3. [2611770 ]
- Kaya M, Moriwaki Y, Ka T, Inokuchi T, Yamamoto A, Takahashi S, Tsutsumi Z, Tsuzita J, Oku Y, Yamamoto T: Plasma concentrations and urinary excretion of purine bases (uric acid, hypoxanthine, and xanthine) and oxypurinol after rigorous exercise. Metabolism. 2006 Jan;55(1):103-7. [16324927 ]
- Liu Z, Li T, Wang E: Simultaneous determination of guanine, uric acid, hypoxanthine and xanthine in human plasma by reversed-phase high-performance liquid chromatography with amperometric detection. Analyst. 1995 Aug;120(8):2181-4. [7677251 ]
- Becker MA, Kisicki J, Khosravan R, Wu J, Mulford D, Hunt B, MacDonald P, Joseph-Ridge N: Febuxostat (TMX-67), a novel, non-purine, selective inhibitor of xanthine oxidase, is safe and decreases serum urate in healthy volunteers. Nucleosides Nucleotides Nucleic Acids. 2004 Oct;23(8-9):1111-6. [15571211 ]
- Kawasaki N, Tanimoto T, Tanaka A, Hayakawa T, Miyasaka N: Determination of non-protein-bound iron in human synovial fluid by high-performance liquid chromatography with electrochemical detection. J Chromatogr B Biomed Appl. 1994 Jun 17;656(2):436-40. [7987499 ]
- Cooper N, Khosravan R, Erdmann C, Fiene J, Lee JW: Quantification of uric acid, xanthine and hypoxanthine in human serum by HPLC for pharmacodynamic studies. J Chromatogr B Analyt Technol Biomed Life Sci. 2006 Jun 6;837(1-2):1-10. Epub 2006 May 2. [16631418 ]
- Eells JT, Spector R: Purine and pyrimidine base and nucleoside concentrations in human cerebrospinal fluid and plasma. Neurochem Res. 1983 Nov;8(11):1451-7. [6656991 ]
- Kiss A, Barenyi M, Csontai A: Xanthine stone in the urinary bladder of a male child. Urol Int. 1999;63(4):242-4. [10743702 ]
- Kjaergaard N, Moller-Petersen JF, Kristiansen FV, Petersen PL, Ekelund S, Skovbo P: Xanthine and hypoxanthine in amniotic fluid during pregnancy. Dan Med Bull. 1990 Dec;37(6):559-60. [2127397 ]
- Wiley DM, Szabo I, Maguire MH, Finley BE, Bennett TL: Measurement of hypoxanthine and xanthine in late-gestation human amniotic fluid by reversed-phase high-performance liquid chromatography with photodiode-array detection. J Chromatogr. 1990 Nov 30;533:73-86. [2081781 ]
- Gudbjornsson B, Zak A, Niklasson F, Hallgren R: Hypoxanthine, xanthine, and urate in synovial fluid from patients with inflammatory arthritides. Ann Rheum Dis. 1991 Oct;50(10):669-72. [1958086 ]
- Ginsburg I: Could synergistic interactions among reactive oxygen species, proteinases, membrane-perforating enzymes, hydrolases, microbial hemolysins and cytokines be the main cause of tissue damage in infectious and inflammatory conditions? Med Hypotheses. 1998 Oct;51(4):337-46. [9824842 ]
- Sreekumar A, Poisson LM, Rajendiran TM, Khan AP, Cao Q, Yu J, Laxman B, Mehra R, Lonigro RJ, Li Y, Nyati MK, Ahsan A, Kalyana-Sundaram S, Han B, Cao X, Byun J, Omenn GS, Ghosh D, Pennathur S, Alexander DC, Berger A, Shuster JR, Wei JT, Varambally S, Beecher C, Chinnaiyan AM: Metabolomic profiles delineate potential role for sarcosine in prostate cancer progression. Nature. 2009 Feb 12;457(7231):910-4. doi: 10.1038/nature07762. [19212411 ]
|
|---|
| Gene Regulation |
|---|
| Up-Regulated Genes | Not Available |
|---|
| Down-Regulated Genes | Not Available |
|---|