| Record Information |
|---|
| Version | 2.0 |
|---|
| Creation Date | 2014-08-29 06:51:34 UTC |
|---|
| Update Date | 2018-03-21 17:46:24 UTC |
|---|
| Accession Number | T3D4467 |
|---|
| Identification |
|---|
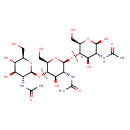
| Common Name | Chitin |
|---|
| Class | Small Molecule |
|---|
| Description | Chitin is one of the main components in the cell walls of fungi, the exoskeletons of insects and other arthropods (such as crustaceans) as well as fish and frogs. It is a polysaccharide that is constructed from units of acetylglucosamine (more completely, N-acetyl-D-glucose-2-amine). These are linked together in beta-1,4 fashion (in a similar manner to the glucose units which form cellulose). In effect, chitin may be described as cellulose with one hydroxyl group on each monomer replaced by an acetylamine group. This allows for increased hydrogen bonding between adjacent polymers, giving the polymer increased strength. Chitin is the second most abundant polysaccharide in the world (after cellulose). Chitinases break down chitin and are a part of the defence mechanism of mammals against chitin-containing parasites in lower life forms. Under certain circumstances, chitin can act as an allergen. Research using murine models has shown that chitin is a size-dependent microbial-associated molecular pattern (MAMP) that can induce an immunological response via pattern recognition receptors. Medium-sized chitin micro-particles (CMPs) have been shown to induce inflammation, while small-sized CMPs reduce inflammation. Additionally, mammalian chitinases may play a key role in mediating the T-helper 2 cell-driven inflammatory response that is commonly associated with asthma. The high prevalence of asthma among people working with chitinous substances, such as crabs and fungi, suggests that chitin might be an allergen playing a significant role in the development of asthma. |
|---|
| Compound Type | - Animal Toxin
- Ether
- Food Toxin
- Household Toxin
- Insect Toxin
- Metabolite
- Natural Compound
- Organic Compound
|
|---|
| Chemical Structure | |
|---|
| Synonyms | | Synonym | | beta-1,4-Poly-N-acetyl-D-glucosamine | | beta-1,4-Poly-N-acetyl-delta-glucosamine | | Poly 2-Acetamido-2-deoxy-D-glucose | | Poly 2-Acetamido-2-deoxy-delta-glucose | | [1,4-(N-Acetyl-beta-D-glucosaminyl)]N | | [1,4-(N-Acetyl-beta-D-glucosaminyl)]N+1 | | [1,4-(N-Acetyl-beta-delta-glucosaminyl)]N | | [1,4-(N-Acetyl-beta-delta-glucosaminyl)]N+1 |
|
|---|
| Chemical Formula | C28H49N3O16 |
|---|
| Average Molecular Mass | 683.699 g/mol |
|---|
| Monoisotopic Mass | 683.311 g/mol |
|---|
| CAS Registry Number | 1398-61-4 |
|---|
| IUPAC Name | Not Available |
|---|
| Traditional Name | Not Available |
|---|
| SMILES | [H][C@@]1(O)OC([H])(CO)[C@@]([H])(COC[C@]2([H])OC([H])(CO)[C@@]([H])(COC[C@]3([H])OC([H])(CO)[C@@]([H])(O)[C@]([H])(O)C3([H])N=C(C)O)[C@]([H])(O)C2([H])N=C(C)O)[C@]([H])(O)C1([H])N=C(C)O |
|---|
| InChI Identifier | InChI=1S/C28H49N3O16/c1-11(35)29-21-19(9-44-8-15-17(5-33)47-28(42)23(25(15)39)31-13(3)37)45-16(4-32)14(24(21)38)7-43-10-20-22(30-12(2)36)27(41)26(40)18(6-34)46-20/h14-28,32-34,38-42H,4-10H2,1-3H3,(H,29,35)(H,30,36)(H,31,37)/t14-,15-,16?,17?,18?,19+,20+,21?,22?,23?,24+,25+,26-,27-,28-/m1/s1 |
|---|
| InChI Key | InChIKey=DJHJJVWPFGHIPH-OODMECLYSA-N |
|---|
| Chemical Taxonomy |
|---|
| Description | belongs to the class of organic compounds known as acylaminosugars. These are organic compounds containing a sugar linked to a chain through N-acyl group. |
|---|
| Kingdom | Organic compounds |
|---|
| Super Class | Organic oxygen compounds |
|---|
| Class | Organooxygen compounds |
|---|
| Sub Class | Carbohydrates and carbohydrate conjugates |
|---|
| Direct Parent | Acylaminosugars |
|---|
| Alternative Parents | |
|---|
| Substituents | - Oligosaccharide
- Acylaminosugar
- N-acyl-alpha-hexosamine
- Glycosyl compound
- O-glycosyl compound
- Oxane
- Acetamide
- Carboxamide group
- Hemiacetal
- Secondary carboxylic acid amide
- Secondary alcohol
- Acetal
- Carboxylic acid derivative
- Oxacycle
- Organoheterocyclic compound
- Hydrocarbon derivative
- Organic oxide
- Organic nitrogen compound
- Organopnictogen compound
- Carbonyl group
- Alcohol
- Primary alcohol
- Organonitrogen compound
- Aliphatic heteromonocyclic compound
|
|---|
| Molecular Framework | Aliphatic heteromonocyclic compounds |
|---|
| External Descriptors | |
|---|
| Biological Properties |
|---|
| Status | Detected and Not Quantified |
|---|
| Origin | Exogenous |
|---|
| Cellular Locations | |
|---|
| Biofluid Locations | Not Available |
|---|
| Tissue Locations | - Fibroblasts
- Intestine
- Spleen
- Testes
|
|---|
| Pathways | |
|---|
| Applications | |
|---|
| Biological Roles | |
|---|
| Chemical Roles | Not Available |
|---|
| Physical Properties |
|---|
| State | Solid |
|---|
| Appearance | White powder. |
|---|
| Experimental Properties | | Property | Value |
|---|
| Melting Point | Not Available | | Boiling Point | Not Available | | Solubility | Not Available | | LogP | Not Available |
|
|---|
| Predicted Properties | |
|---|
| Spectra |
|---|
| Spectra | | Spectrum Type | Description | Splash Key | Deposition Date | View |
|---|
| Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_1_1) - 70eV, Positive | Not Available | 2021-10-17 | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_1_2) - 70eV, Positive | Not Available | 2021-10-17 | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_1_3) - 70eV, Positive | Not Available | 2021-10-17 | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_1_4) - 70eV, Positive | Not Available | 2021-10-17 | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_1_5) - 70eV, Positive | Not Available | 2021-10-17 | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_1_6) - 70eV, Positive | Not Available | 2021-10-17 | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_1_7) - 70eV, Positive | Not Available | 2021-10-17 | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_1_8) - 70eV, Positive | Not Available | 2021-10-17 | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_1_9) - 70eV, Positive | Not Available | 2021-10-17 | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_1_10) - 70eV, Positive | Not Available | 2021-10-17 | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_1_11) - 70eV, Positive | Not Available | 2021-10-17 | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_2_1) - 70eV, Positive | Not Available | 2021-10-17 | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_2_2) - 70eV, Positive | Not Available | 2021-10-17 | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_2_3) - 70eV, Positive | Not Available | 2021-10-17 | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_2_4) - 70eV, Positive | Not Available | 2021-10-17 | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_2_5) - 70eV, Positive | Not Available | 2021-10-17 | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_2_6) - 70eV, Positive | Not Available | 2021-10-17 | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_2_7) - 70eV, Positive | Not Available | 2021-10-17 | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_2_8) - 70eV, Positive | Not Available | 2021-10-17 | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_2_9) - 70eV, Positive | Not Available | 2021-10-17 | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_2_10) - 70eV, Positive | Not Available | 2021-10-17 | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_2_11) - 70eV, Positive | Not Available | 2021-10-17 | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_2_12) - 70eV, Positive | Not Available | 2021-10-17 | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_2_13) - 70eV, Positive | Not Available | 2021-10-17 | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_2_14) - 70eV, Positive | Not Available | 2021-10-17 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-01si-0020195000-2e6a56a958abe902a3e9 | 2017-07-25 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-11ec-0273492000-ace7f79ec49825bd5fd6 | 2017-07-25 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-0zmr-6594462000-c3586d5d836795b9e53a | 2017-07-25 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-006w-1352911000-863eeeb6aa19c4b1a13e | 2017-07-26 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-0a4i-2344295000-9c0b132184373172725c | 2017-07-26 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-0zfr-4957010000-d6d15cd685b32cc6a8cd | 2017-07-26 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-00ai-0000190000-6edcc24d4bf1dd7ae865 | 2021-09-22 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-0a4i-5001391000-7e5b14aac0963d66ab7e | 2021-09-22 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-0abc-9022240000-288bdbd056f808dedbb7 | 2021-09-22 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-116r-0251519000-1653bc016143a1446468 | 2021-09-22 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-0m99-3941354000-8656850f46179cee5b75 | 2021-09-22 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-01qi-7900000000-d7d93fd040c9f9381f77 | 2021-09-22 | View Spectrum | | 1D NMR | 1H NMR Spectrum (1D, 100 MHz, D2O, predicted) | Not Available | 2021-09-25 | View Spectrum | | 1D NMR | 13C NMR Spectrum (1D, 100 MHz, D2O, predicted) | Not Available | 2021-09-25 | View Spectrum | | 1D NMR | 1H NMR Spectrum (1D, 1000 MHz, D2O, predicted) | Not Available | 2021-09-25 | View Spectrum | | 1D NMR | 13C NMR Spectrum (1D, 1000 MHz, D2O, predicted) | Not Available | 2021-09-25 | View Spectrum | | 1D NMR | 1H NMR Spectrum (1D, 200 MHz, D2O, predicted) | Not Available | 2021-09-25 | View Spectrum | | 1D NMR | 13C NMR Spectrum (1D, 200 MHz, D2O, predicted) | Not Available | 2021-09-25 | View Spectrum | | 1D NMR | 1H NMR Spectrum (1D, 300 MHz, D2O, predicted) | Not Available | 2021-09-25 | View Spectrum | | 1D NMR | 13C NMR Spectrum (1D, 300 MHz, D2O, predicted) | Not Available | 2021-09-25 | View Spectrum | | 1D NMR | 1H NMR Spectrum (1D, 400 MHz, D2O, predicted) | Not Available | 2021-09-25 | View Spectrum | | 1D NMR | 13C NMR Spectrum (1D, 400 MHz, D2O, predicted) | Not Available | 2021-09-25 | View Spectrum | | 1D NMR | 1H NMR Spectrum (1D, 500 MHz, D2O, predicted) | Not Available | 2021-09-25 | View Spectrum | | 1D NMR | 13C NMR Spectrum (1D, 500 MHz, D2O, predicted) | Not Available | 2021-09-25 | View Spectrum | | 1D NMR | 1H NMR Spectrum (1D, 600 MHz, D2O, predicted) | Not Available | 2021-09-25 | View Spectrum | | 1D NMR | 13C NMR Spectrum (1D, 600 MHz, D2O, predicted) | Not Available | 2021-09-25 | View Spectrum | | 1D NMR | 1H NMR Spectrum (1D, 700 MHz, D2O, predicted) | Not Available | 2021-09-25 | View Spectrum | | 1D NMR | 13C NMR Spectrum (1D, 700 MHz, D2O, predicted) | Not Available | 2021-09-25 | View Spectrum | | 1D NMR | 1H NMR Spectrum (1D, 800 MHz, D2O, predicted) | Not Available | 2021-09-25 | View Spectrum | | 1D NMR | 13C NMR Spectrum (1D, 800 MHz, D2O, predicted) | Not Available | 2021-09-25 | View Spectrum | | 1D NMR | 1H NMR Spectrum (1D, 900 MHz, D2O, predicted) | Not Available | 2021-09-25 | View Spectrum | | 1D NMR | 13C NMR Spectrum (1D, 900 MHz, D2O, predicted) | Not Available | 2021-09-25 | View Spectrum |
|
|---|
| Toxicity Profile |
|---|
| Route of Exposure | Not Available |
|---|
| Mechanism of Toxicity | Not Available |
|---|
| Metabolism | Not Available |
|---|
| Toxicity Values | Not Available |
|---|
| Lethal Dose | Not Available |
|---|
| Carcinogenicity (IARC Classification) | No indication of carcinogenicity to humans (not listed by IARC). |
|---|
| Uses/Sources | If chitin is detected they then produce enzymes to digest the chitin by reducing it to simple sugars and ammonia. It is the second most abundant biopolymer on earth, found especially in insects and fungi. |
|---|
| Minimum Risk Level | Not Available |
|---|
| Health Effects | Not Available |
|---|
| Symptoms | Not Available |
|---|
| Treatment | Not Available |
|---|
| Normal Concentrations |
|---|
| Not Available |
|---|
| Abnormal Concentrations |
|---|
| Not Available |
|---|
| External Links |
|---|
| DrugBank ID | Not Available |
|---|
| HMDB ID | HMDB03362 |
|---|
| PubChem Compound ID | 21252321 |
|---|
| ChEMBL ID | Not Available |
|---|
| ChemSpider ID | 399508 |
|---|
| KEGG ID | C00461 |
|---|
| UniProt ID | Not Available |
|---|
| OMIM ID | |
|---|
| ChEBI ID | 17029 |
|---|
| BioCyc ID | 14-N-ACETYL-BETA-D-GLUCOSAMINYLN1 |
|---|
| CTD ID | Not Available |
|---|
| Stitch ID | Not Available |
|---|
| PDB ID | Not Available |
|---|
| ACToR ID | Not Available |
|---|
| Wikipedia Link | Chitin |
|---|
| References |
|---|
| Synthesis Reference | Kurita, Keisuke; Koyama, Yoshiyuki; Nishimura, Shinichiro; Kamiya, Mami. Facile preparation of water-soluble chitin from chitosan. Chemistry Letters (1989), (9), 1597-8. |
|---|
| MSDS | Link |
|---|
| General References | - Nakajima M, Atsumi K, Kifune K, Miura K, Kanamaru H: Chitin is an effective material for sutures. Jpn J Surg. 1986 Nov;16(6):418-24. [3820865 ]
- Soule JB, Halverson AL, Becker RB, Pistole MC, Orenstein JM: A patient with acquired immunodeficiency syndrome and untreated Encephalitozoon (Septata) intestinalis microsporidiosis leading to small bowel perforation. Response to albendazole. Arch Pathol Lab Med. 1997 Aug;121(8):880-7. [9278619 ]
- Gheri G, Sgambati E, Thyrion GD, Vichi D, Orlandini GE: The oligosaccharidic content of the glycoconjugates of the prepubertal descended and undescended testis: lectin histochemical study. Ital J Anat Embryol. 2004 Apr-Jun;109(2):69-84. [15481156 ]
- Hatcher VB, Schwarzmann GO, Jeanloz RW, McArthur JW: Changes in the sialic acid concentration in the major cervical glycoprotein from the bonnet monkey (Macaca radiata) during a hormonally induced cycle. Fertil Steril. 1977 Jun;28(6):682-8. [405259 ]
- Piskun RP, Pentiuk AA, Serkova VK, Polesia TL, Savitskaia EA: [Enterosorbents in the treatment of atherosclerosis]. Eksp Klin Farmakol. 1998 Mar-Apr;61(2):69-74. [9621181 ]
- Kawashita M, Nakao M, Minoda M, Kim HM, Beppu T, Miyamoto T, Kokubo T, Nakamura T: Apatite-forming ability of carboxyl group-containing polymer gels in a simulated body fluid. Biomaterials. 2003 Jun;24(14):2477-84. [12695074 ]
- Howling GI, Dettmar PW, Goddard PA, Hampson FC, Dornish M, Wood EJ: The effect of chitin and chitosan on the proliferation of human skin fibroblasts and keratinocytes in vitro. Biomaterials. 2001 Nov;22(22):2959-66. [11575470 ]
- Vasstrand EN, Jensen HB: Affinity chromatography of human saliva lysozyme and effect of pH and ionic strength on lytic activity. Scand J Dent Res. 1980 Jun;88(3):219-28. [6932088 ]
- Collard CD, Montalto MC, Reenstra WR, Buras JA, Stahl GL: Endothelial oxidative stress activates the lectin complement pathway: role of cytokeratin 1. Am J Pathol. 2001 Sep;159(3):1045-54. [11549596 ]
- Shibuya N, Nakamura K, Ogoshi K, Ohta T, Hori Y, Kodama K, Yamamoto M: Modification of mutagenic activities of pro-mutagens by glyco-ursodeoxycholic acid in the Ames assay. Tohoku J Exp Med. 1999 Sep;189(1):1-9. [10622203 ]
- Ishihara C, Yoshimatsu K, Tsuji M, Arikawa J, Saiki I, Tokura S, Azuma I: Anti-viral activity of sulfated chitin derivatives against Friend murine leukaemia and herpes simplex type-1 viruses. Vaccine. 1993;11(6):670-4. [8391740 ]
- Zampini M, Pruzzo C, Bondre VP, Tarsi R, Cosmo M, Bacciaglia A, Chhabra A, Srivastava R, Srivastava BS: Vibrio cholerae persistence in aquatic environments and colonization of intestinal cells: involvement of a common adhesion mechanism. FEMS Microbiol Lett. 2005 Mar 15;244(2):267-73. [15766778 ]
- Xu Y, Olman V, Xu D: Clustering gene expression data using a graph-theoretic approach: an application of minimum spanning trees. Bioinformatics. 2002 Apr;18(4):536-45. [12016051 ]
- Sharaev PN, Strelkov NS, Sannikova AA, Zvorykin IA, Men'shikova NN, Sakhabutdinova EP, Gabdrakhmanova NK: [The method for detection of urinary lysozyme]. Klin Lab Diagn. 2001 Apr;(4):52-3. [11393034 ]
- Farnia P, Mohammadi F, Zarifi Z, Tabatabee DJ, Ganavi J, Ghazisaeedi K, Farnia PK, Gheydi M, Bahadori M, Masjedi MR, Velayati AA: Improving sensitivity of direct microscopy for detection of acid-fast bacilli in sputum: use of chitin in mucus digestion. J Clin Microbiol. 2002 Feb;40(2):508-11. [11825964 ]
- Sano H, Matsukubo T, Shibasaki K, Itoi H, Takaesu Y: Inhibition of adsorption of oral streptococci to saliva treated hydroxyapatite by chitin derivatives. Bull Tokyo Dent Coll. 1991 Feb;32(1):9-17. [1668072 ]
- Johnson SM, Pappagianis D: The coccidioidal complement fixation and immunodiffusion-complement fixation antigen is a chitinase. Infect Immun. 1992 Jul;60(7):2588-92. [1612728 ]
|
|---|
| Gene Regulation |
|---|
| Up-Regulated Genes | Not Available |
|---|
| Down-Regulated Genes | Not Available |
|---|