| Record Information |
|---|
| Version | 2.0 |
|---|
| Creation Date | 2014-09-11 02:04:51 UTC |
|---|
| Update Date | 2014-12-24 20:26:55 UTC |
|---|
| Accession Number | T3D4684 |
|---|
| Identification |
|---|
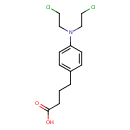
| Common Name | Chlorambucil |
|---|
| Class | Small Molecule |
|---|
| Description | A nitrogen mustard alkylating agent used as antineoplastic agent for the treatment of various malignant and nonmalignant diseases. Although it is less toxic than most other nitrogen mustards, it has been listed as a known carcinogen in the Fourth Annual Report on Carcinogens (NTP 85-002, 1985). |
|---|
| Compound Type | - Amine
- Antineoplastic Agent, Alkylating
- Drug
- Metabolite
- Organic Compound
- Organochloride
- Synthetic Compound
|
|---|
| Chemical Structure | |
|---|
| Synonyms | | Synonym | | 4-(P-Bis(beta-chloroethyl)aminophenyl)butyric acid | | 4-(p-bis(β-chloroethyl)aminophenyl)butyric acid | | 4-[p-[bis(2-chloroethyl)amino]phenyl]butyric acid | | Ambochlorin | | Celkeran | | Chlocambucil | | CHLORAMBUCIL | | Chloraminophen | | Chloraminophene | | Chlorbutin | | Chlorbutine | | Chloroambucil | | Chlorobutin | | Chlorobutine | | gamma-[P-Di(2-chloroethyl)aminophenyl]butyric acid | | Leukeran | | N,N-Di-2-chloroethyl-gamma-P-aminophenylbutyric acid | | N,N-di-2-chloroethyl-γ-p-aminophenylbutyric acid | | Phenylbutyric acid nitrogen mustard | | γ-[p-di(2-chloroethyl)aminophenyl]butyric acid |
|
|---|
| Chemical Formula | C14H19Cl2NO2 |
|---|
| Average Molecular Mass | 304.212 g/mol |
|---|
| Monoisotopic Mass | 303.079 g/mol |
|---|
| CAS Registry Number | 305-03-3 |
|---|
| IUPAC Name | 4-{4-[bis(2-chloroethyl)amino]phenyl}butanoic acid |
|---|
| Traditional Name | chlorambucil |
|---|
| SMILES | OC(=O)CCCC1=CC=C(C=C1)N(CCCl)CCCl |
|---|
| InChI Identifier | InChI=1S/C14H19Cl2NO2/c15-8-10-17(11-9-16)13-6-4-12(5-7-13)2-1-3-14(18)19/h4-7H,1-3,8-11H2,(H,18,19) |
|---|
| InChI Key | InChIKey=JCKYGMPEJWAADB-UHFFFAOYSA-N |
|---|
| Chemical Taxonomy |
|---|
| Description | belongs to the class of organic compounds known as nitrogen mustard compounds. Nitrogen mustard compounds are compounds having two beta-haloalkyl groups bound to a nitrogen atom. |
|---|
| Kingdom | Organic compounds |
|---|
| Super Class | Organic nitrogen compounds |
|---|
| Class | Organonitrogen compounds |
|---|
| Sub Class | Nitrogen mustard compounds |
|---|
| Direct Parent | Nitrogen mustard compounds |
|---|
| Alternative Parents | |
|---|
| Substituents | - Aniline or substituted anilines
- Dialkylarylamine
- Tertiary aliphatic/aromatic amine
- Nitrogen mustard
- Monocyclic benzene moiety
- Benzenoid
- Amino acid or derivatives
- Amino acid
- Tertiary amine
- Carboxylic acid derivative
- Carboxylic acid
- Monocarboxylic acid or derivatives
- Alkyl chloride
- Organohalogen compound
- Organochloride
- Organooxygen compound
- Hydrocarbon derivative
- Organic oxide
- Carbonyl group
- Organopnictogen compound
- Organic oxygen compound
- Amine
- Alkyl halide
- Aromatic homomonocyclic compound
|
|---|
| Molecular Framework | Aromatic homomonocyclic compounds |
|---|
| External Descriptors | |
|---|
| Biological Properties |
|---|
| Status | Detected and Not Quantified |
|---|
| Origin | Exogenous |
|---|
| Cellular Locations | |
|---|
| Biofluid Locations | Not Available |
|---|
| Tissue Locations | Not Available |
|---|
| Pathways | Not Available |
|---|
| Applications | Not Available |
|---|
| Biological Roles | Not Available |
|---|
| Chemical Roles | Not Available |
|---|
| Physical Properties |
|---|
| State | Solid |
|---|
| Appearance | White powder. |
|---|
| Experimental Properties | | Property | Value |
|---|
| Melting Point | 65°C | | Boiling Point | Not Available | | Solubility | 1.24E+004 mg/L | | LogP | 1.7 |
|
|---|
| Predicted Properties | |
|---|
| Spectra |
|---|
| Spectra | | Spectrum Type | Description | Splash Key | Deposition Date | View |
|---|
| Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | splash10-066u-2790000000-bf4786edba412b2ccaae | 2017-09-01 | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (1 TMS) - 70eV, Positive | splash10-024i-5393000000-3be6fe379684458bfa85 | 2017-10-06 | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | Not Available | 2021-10-12 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-qTof , Positive | splash10-004i-0002900000-472e293a011d716bb5a8 | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - , positive | splash10-0zfr-0519000000-1bca00472dfc8c30a465 | 2017-09-14 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-0udr-0096000000-3899963778fe2e6e88de | 2016-08-03 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-08g0-3591000000-6ae3349b3f490c58310d | 2016-08-03 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-0j4i-6920000000-15483700b73ee5d5c44b | 2016-08-03 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-0udi-0039000000-b8ff9ca35aa22b00bd35 | 2016-08-03 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-0uxr-1193000000-7606e0823853c0c258ee | 2016-08-03 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-052f-9540000000-f5720e555315984712a6 | 2016-08-03 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-001i-9004000000-788a1a6f740556f91441 | 2021-09-23 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-001i-9000000000-724cdb1db65352c21f6c | 2021-09-23 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-001i-9100000000-58ace4115f9b048a722e | 2021-09-23 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-0udr-0098000000-d1dce37c9e184d7829de | 2021-09-24 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-00kr-0091000000-f4d0cc5c505d1414f7cd | 2021-09-24 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-014m-1920000000-59f8073bfe41e45f2bdb | 2021-09-24 | View Spectrum | | MS | Mass Spectrum (Electron Ionization) | splash10-0udi-2491000000-ddb7d5da8316562dba36 | 2014-09-20 | View Spectrum |
|
|---|
| Toxicity Profile |
|---|
| Route of Exposure | Not Available |
|---|
| Mechanism of Toxicity | Alkylating agents work by three different mechanisms: 1) attachment of alkyl groups to DNA bases, resulting in the DNA being fragmented by repair enzymes in their attempts to replace the alkylated bases, preventing DNA synthesis and RNA transcription from the affected DNA, 2) DNA damage via the formation of cross-links (bonds between atoms in the DNA) which prevents DNA from being separated for synthesis or transcription, and 3) the induction of mispairing of the nucleotides leading to mutations. |
|---|
| Metabolism | Route of Elimination: Chlorambucil is extensively metabolized in the liver primarily to phenylacetic acid mustard. The pharmacokinetic data suggests that oral chlorambucil undergoes rapid gastrointestinal absorption and plasma clearance and that it is almost completely metabolized, having extremely low urinary excretion.
Half Life: 1.5 hours |
|---|
| Toxicity Values | Not Available |
|---|
| Lethal Dose | Not Available |
|---|
| Carcinogenicity (IARC Classification) | 1, carcinogenic to humans. (3) |
|---|
| Uses/Sources | For treatment of chronic lymphatic (lymphocytic) leukemia, childhood minimal-change nephrotic syndrome, and malignant lymphomas including lymphosarcoma, giant follicular lymphoma, Hodgkin's disease, non-Hodgkin's lymphomas, and Waldenstrн_mду»s Macroglobulinemia. |
|---|
| Minimum Risk Level | Not Available |
|---|
| Health Effects | Not Available |
|---|
| Symptoms | Not Available |
|---|
| Treatment | Not Available |
|---|
| Normal Concentrations |
|---|
| Not Available |
|---|
| Abnormal Concentrations |
|---|
| Not Available |
|---|
| External Links |
|---|
| DrugBank ID | DB00291 |
|---|
| HMDB ID | HMDB14436 |
|---|
| PubChem Compound ID | 2708 |
|---|
| ChEMBL ID | CHEMBL515 |
|---|
| ChemSpider ID | 2607 |
|---|
| KEGG ID | C06900 |
|---|
| UniProt ID | Not Available |
|---|
| OMIM ID | |
|---|
| ChEBI ID | 28830 |
|---|
| BioCyc ID | Not Available |
|---|
| CTD ID | Not Available |
|---|
| Stitch ID | Not Available |
|---|
| PDB ID | CBL |
|---|
| ACToR ID | Not Available |
|---|
| Wikipedia Link | Chlorambucil |
|---|
| References |
|---|
| Synthesis Reference | Phillips, A. P. and Mentha, J.W.; U.S.Patent 3,046,301; July 24, 1962; assigned to Burroughs
Wellcome & Co. (U.S.A.) Inc. |
|---|
| MSDS | Link |
|---|
| General References | - Rai KR, Peterson BL, Appelbaum FR, Kolitz J, Elias L, Shepherd L, Hines J, Threatte GA, Larson RA, Cheson BD, Schiffer CA: Fludarabine compared with chlorambucil as primary therapy for chronic lymphocytic leukemia. N Engl J Med. 2000 Dec 14;343(24):1750-7. [11114313 ]
- Yang K, Tan J, Wu T: Alkylating agents for Waldenstrom's macroglobulinaemia. Cochrane Database Syst Rev. 2009 Jan 21;(1):CD006719. doi: 10.1002/14651858.CD006719.pub3. [19160296 ]
- International Agency for Research on Cancer (2014). IARC Monographs on the Evaluation of Carcinogenic Risks to Humans. [Link]
|
|---|
| Gene Regulation |
|---|
| Up-Regulated Genes | Not Available |
|---|
| Down-Regulated Genes | Not Available |
|---|