| Record Information |
|---|
| Version | 2.0 |
|---|
| Creation Date | 2014-09-11 05:16:39 UTC |
|---|
| Update Date | 2014-12-24 20:26:57 UTC |
|---|
| Accession Number | T3D4789 |
|---|
| Identification |
|---|
| Common Name | Troglitazone |
|---|
| Class | Small Molecule |
|---|
| Description | Troglitazone was withdrawn in 2000 due to risk of hepatotoxicity. It was superseded by pioglitazone and rosiglitazone. |
|---|
| Compound Type | - Amide
- Amine
- Drug
- Ether
- Organic Compound
- Synthetic Compound
|
|---|
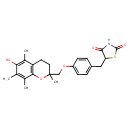
| Chemical Structure | |
|---|
| Synonyms | | Synonym | | Noscal | | Resulin | | Rezulin | | Romozin |
|
|---|
| Chemical Formula | C24H27NO5S |
|---|
| Average Molecular Mass | 441.540 g/mol |
|---|
| Monoisotopic Mass | 441.161 g/mol |
|---|
| CAS Registry Number | 97322-87-7 |
|---|
| IUPAC Name | 5-({4-[(6-hydroxy-2,5,7,8-tetramethyl-3,4-dihydro-2H-1-benzopyran-2-yl)methoxy]phenyl}methyl)-1,3-thiazolidine-2,4-dione |
|---|
| Traditional Name | troglitazone |
|---|
| SMILES | CC1=C(O)C(C)=C2CCC(C)(COC3=CC=C(CC4SC(=O)N=C4O)C=C3)OC2=C1C |
|---|
| InChI Identifier | InChI=1/C24H27NO5S/c1-13-14(2)21-18(15(3)20(13)26)9-10-24(4,30-21)12-29-17-7-5-16(6-8-17)11-19-22(27)25-23(28)31-19/h5-8,19,26H,9-12H2,1-4H3,(H,25,27,28) |
|---|
| InChI Key | InChIKey=GXPHKUHSUJUWKP-UHFFFAOYNA-N |
|---|
| Chemical Taxonomy |
|---|
| Description | belongs to the class of organic compounds known as 1-benzopyrans. These are organic aromatic compounds that 1-benzopyran, a bicyclic compound made up of a benzene ring fused to a pyran, so that the oxygen atom is at the 1-position. |
|---|
| Kingdom | Organic compounds |
|---|
| Super Class | Organoheterocyclic compounds |
|---|
| Class | Benzopyrans |
|---|
| Sub Class | 1-benzopyrans |
|---|
| Direct Parent | 1-benzopyrans |
|---|
| Alternative Parents | |
|---|
| Substituents | - 1-benzopyran
- Phenoxy compound
- Phenol ether
- Alkyl aryl ether
- Thiazolidinedione
- Monocyclic benzene moiety
- Benzenoid
- Dicarboximide
- Thiazolidine
- Carbonic acid derivative
- Thiocarbamic acid derivative
- Carboxylic acid derivative
- Ether
- Oxacycle
- Azacycle
- Organonitrogen compound
- Organic oxide
- Organopnictogen compound
- Organic nitrogen compound
- Organooxygen compound
- Hydrocarbon derivative
- Organic oxygen compound
- Carbonyl group
- Aromatic heteropolycyclic compound
|
|---|
| Molecular Framework | Aromatic heteropolycyclic compounds |
|---|
| External Descriptors | |
|---|
| Biological Properties |
|---|
| Status | Detected and Not Quantified |
|---|
| Origin | Exogenous |
|---|
| Cellular Locations | |
|---|
| Biofluid Locations | Not Available |
|---|
| Tissue Locations | Not Available |
|---|
| Pathways | Not Available |
|---|
| Applications | Not Available |
|---|
| Biological Roles | Not Available |
|---|
| Chemical Roles | Not Available |
|---|
| Physical Properties |
|---|
| State | Solid |
|---|
| Appearance | White powder. |
|---|
| Experimental Properties | | Property | Value |
|---|
| Melting Point | 184-186°C | | Boiling Point | Not Available | | Solubility | Not Available | | LogP | 3.6 |
|
|---|
| Predicted Properties | |
|---|
| Spectra |
|---|
| Spectra | | Spectrum Type | Description | Splash Key | Deposition Date | View |
|---|
| Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | splash10-0a6r-1889600000-e7d440741c30fa701997 | 2021-09-24 | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | Not Available | 2021-10-12 | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_1_1) - 70eV, Positive | Not Available | 2021-11-05 | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_1_2) - 70eV, Positive | Not Available | 2021-11-05 | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TBDMS_1_1) - 70eV, Positive | Not Available | 2021-11-05 | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TBDMS_1_2) - 70eV, Positive | Not Available | 2021-11-05 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - 60V, Negative | splash10-0f6t-1900000000-b41d1d9b00383779a8be | 2021-09-20 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - 45V, Negative | splash10-0f6t-0900000000-ab7c5539034df635ac14 | 2021-09-20 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - 75V, Negative | splash10-0fr2-1900000000-1a1e22916ed5413720eb | 2021-09-20 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - 90V, Negative | splash10-014j-2900000000-52697cbd413c9fbe49f8 | 2021-09-20 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - 30V, Negative | splash10-0005-1507900000-4f7d34b5dc4e0c481b68 | 2021-09-20 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - 15V, Negative | splash10-0006-0000900000-8c86a1f2bb9af73c4ae9 | 2021-09-20 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - 60V, Positive | splash10-014i-1900000000-c7a1ca757fec1b1a4e5e | 2021-09-20 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - 75V, Positive | splash10-014i-3900000000-c72fbe14217a1770ef4d | 2021-09-20 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-0006-0430900000-ceb18bd79d0f6da79e7d | 2016-08-02 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-014i-0912100000-c1ddd0b2276cc0642c28 | 2016-08-02 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-0bt9-0910000000-70eed845667711a7ff74 | 2016-08-02 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-0006-0133900000-d4c360adbc610d5363cf | 2016-08-03 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-0v4l-4759300000-7eda5e5df7869dbcd569 | 2016-08-03 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-0006-9700000000-bbf919843d75c50ea813 | 2016-08-03 | View Spectrum |
|
|---|
| Toxicity Profile |
|---|
| Route of Exposure | Absorbed rapidly. Food increases the extent of absorption by 30% to 85%. |
|---|
| Mechanism of Toxicity | Troglitazone is a thiazolidinedione antidiabetic agent that lowers blood glucose by improving target cell response to insulin. It has a unique mechanism of action that is dependent on the presence of insulin for activity. Troglitazone decreases hepatic glucose output and increases insulin dependent glucose disposal in skeletal muscle. Its mechanism of action is thought to involve binding to nuclear receptors (PPAR) that regulate the transcription of a number of insulin responsive genes critical for the control of glucose and lipid metabolism. Troglitazone is a ligand to both PPAR‘± and PPAR‘_, with a highter affinity for PPAR‘_. The drug also contains an ‘±-tocopheroyl moiety, potentially giving it vitamin E-like activity. Troglitazone has been shown to reduce inflammation, and is associated with a decrase in nuclear factor kappa-B (NF-‘_B) and a concomitant increase in its inhibitor (I‘_B). NF-‘_B is an important cellular transcription regulator for the immune response. Unlike sulfonylureas, troglitazone is not an insulin secretagogue. |
|---|
| Metabolism | A sulfate conjugate metabolite (Metabolite 1) and a quinone metabolite (Metabolite 3) have been detected in the plasma of healthy males. A glucuronide conjugate (Metabolite 2) has been detected in the urine and also in negligible amounts in the plasma. In healthy volunteers and in patients with type 2 diabetes, the steady-state concentration of Metabolite 1 is six to seven times that of troglitazone and Metabolite 3. In in vivo drug interaction studies, troglitazone has been shown to induce cytochrome P450 CYP3A4 at clinically relevant doses.
Half Life: 16-34 hours |
|---|
| Toxicity Values | Not Available |
|---|
| Lethal Dose | Not Available |
|---|
| Carcinogenicity (IARC Classification) | No indication of carcinogenicity to humans (not listed by IARC). |
|---|
| Uses/Sources | For the treatment of Type II diabetes mellitus. It is used alone or in combination with a sulfonylurea, metformin, or insulin as an adjunct to diet and exercise. |
|---|
| Minimum Risk Level | Not Available |
|---|
| Health Effects | Not Available |
|---|
| Symptoms | Not Available |
|---|
| Treatment | Not Available |
|---|
| Normal Concentrations |
|---|
| Not Available |
|---|
| Abnormal Concentrations |
|---|
| Not Available |
|---|
| External Links |
|---|
| DrugBank ID | DB00197 |
|---|
| HMDB ID | Not Available |
|---|
| PubChem Compound ID | 5591 |
|---|
| ChEMBL ID | Not Available |
|---|
| ChemSpider ID | 5389 |
|---|
| KEGG ID | Not Available |
|---|
| UniProt ID | Not Available |
|---|
| OMIM ID | |
|---|
| ChEBI ID | 9753 |
|---|
| BioCyc ID | Not Available |
|---|
| CTD ID | Not Available |
|---|
| Stitch ID | Not Available |
|---|
| PDB ID | Not Available |
|---|
| ACToR ID | Not Available |
|---|
| Wikipedia Link | Troglitazone |
|---|
| References |
|---|
| Synthesis Reference | Krishnamurthi Vyas, Chebiyyam Prabhakar, Sreenivas Dharmaraja Rao, Mamillapalli Ramabadhra Sarma, Om Gaddam Reddy, Rajagopalan Ramanujam, Ranjan Chakrabarti, “Polymorphic forms of troglitazone having enhanced anti-diabetic activity and a process for their preparation.” U.S. Patent US5700820, issued June, 1992. |
|---|
| MSDS | T3D4789.pdf |
|---|
| General References | - Aljada A, Garg R, Ghanim H, Mohanty P, Hamouda W, Assian E, Dandona P: Nuclear factor-kappaB suppressive and inhibitor-kappaB stimulatory effects of troglitazone in obese patients with type 2 diabetes: evidence of an antiinflammatory action? J Clin Endocrinol Metab. 2001 Jul;86(7):3250-6. [11443197 ]
|
|---|
| Gene Regulation |
|---|
| Up-Regulated Genes | Not Available |
|---|
| Down-Regulated Genes | Not Available |
|---|