| Record Information |
|---|
| Version | 2.0 |
|---|
| Creation Date | 2014-10-14 21:18:37 UTC |
|---|
| Update Date | 2014-12-24 20:27:01 UTC |
|---|
| Accession Number | T3D4981 |
|---|
| Identification |
|---|
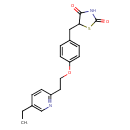
| Common Name | Pioglitazone |
|---|
| Class | Small Molecule |
|---|
| Description | Pioglitazone is used for the treatment of diabetes mellitus type 2. Pioglitazone selectively stimulates nuclear receptor peroxisone proliferator-activated receptor gamma (PPAR-gamma). It modulates the transcription of the insulin-sensitive genes involved in the control of glucose and lipid metabolism in the lipidic, muscular tissues and in the liver. |
|---|
| Compound Type | - Drug
- Hypoglycemic Agent
- Metabolite
- Synthetic Compound
|
|---|
| Chemical Structure | |
|---|
| Synonyms | | Synonym | | (+-)-5-((4-(2-(5-Ethyl-2-pyridinyl)ethoxy)phenyl)methyl)-2,4-thiazolidinedione | | Actos | | Actost | | Glustin | | Pioglitazona | | Pioglitazone HCl | | Pioglitazone Hydrochloride | | Pioglitazonum |
|
|---|
| Chemical Formula | C19H20N2O3S |
|---|
| Average Molecular Mass | 356.439 g/mol |
|---|
| Monoisotopic Mass | 356.119 g/mol |
|---|
| CAS Registry Number | 111025-46-8 |
|---|
| IUPAC Name | 5-({4-[2-(5-ethylpyridin-2-yl)ethoxy]phenyl}methyl)-1,3-thiazolidine-2,4-dione |
|---|
| Traditional Name | pioglitazone |
|---|
| SMILES | CCC1=CN=C(CCOC2=CC=C(CC3SC(=O)N=C3O)C=C2)C=C1 |
|---|
| InChI Identifier | InChI=1/C19H20N2O3S/c1-2-13-3-6-15(20-12-13)9-10-24-16-7-4-14(5-8-16)11-17-18(22)21-19(23)25-17/h3-8,12,17H,2,9-11H2,1H3,(H,21,22,23) |
|---|
| InChI Key | InChIKey=HYAFETHFCAUJAY-UHFFFAOYNA-N |
|---|
| Chemical Taxonomy |
|---|
| Description | belongs to the class of organic compounds known as phenol ethers. These are aromatic compounds containing an ether group substituted with a benzene ring. |
|---|
| Kingdom | Organic compounds |
|---|
| Super Class | Benzenoids |
|---|
| Class | Phenol ethers |
|---|
| Sub Class | Not Available |
|---|
| Direct Parent | Phenol ethers |
|---|
| Alternative Parents | |
|---|
| Substituents | - Phenoxy compound
- Phenol ether
- Alkyl aryl ether
- Thiazolidinedione
- Monocyclic benzene moiety
- Pyridine
- Dicarboximide
- Heteroaromatic compound
- Thiazolidine
- Carbonic acid derivative
- Thiocarbamic acid derivative
- Carboxylic acid derivative
- Ether
- Azacycle
- Organoheterocyclic compound
- Organonitrogen compound
- Organic oxide
- Organopnictogen compound
- Organic nitrogen compound
- Organooxygen compound
- Hydrocarbon derivative
- Organic oxygen compound
- Carbonyl group
- Aromatic heteromonocyclic compound
|
|---|
| Molecular Framework | Aromatic heteromonocyclic compounds |
|---|
| External Descriptors | |
|---|
| Biological Properties |
|---|
| Status | Detected and Not Quantified |
|---|
| Origin | Exogenous |
|---|
| Cellular Locations | |
|---|
| Biofluid Locations | Not Available |
|---|
| Tissue Locations | Not Available |
|---|
| Pathways | Not Available |
|---|
| Applications | Not Available |
|---|
| Biological Roles | Not Available |
|---|
| Chemical Roles | Not Available |
|---|
| Physical Properties |
|---|
| State | Solid |
|---|
| Appearance | Not Available |
|---|
| Experimental Properties | | Property | Value |
|---|
| Melting Point | 183-184 °C | | Boiling Point | Not Available | | Solubility | mg/mL | | LogP | 2.3 |
|
|---|
| Predicted Properties | |
|---|
| Spectra |
|---|
| Spectra | | Spectrum Type | Description | Splash Key | Deposition Date | View |
|---|
| Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | splash10-05fr-0982000000-bc531fd621e090712033 | 2017-09-01 | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | Not Available | 2021-10-12 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-QFT , negative | splash10-0a4i-0009000000-070cf69939743719671d | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-QFT , negative | splash10-0bt9-0209000000-d9d34be41f49a9ffbdef | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-QFT , negative | splash10-0udi-0900000000-ef25b6f93ae4fcee23c3 | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-QFT , negative | splash10-0udi-0900000000-3011f00e839bfc06d89d | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-QFT , negative | splash10-0gba-0900000000-b0488e0d155d1587bbb5 | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-QFT , negative | splash10-014j-0900000000-b32570ad6af58a4a6c72 | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-QFT , negative | splash10-0a4i-0009000000-c67370f6cb2ad36cab3e | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-QTOF , positive | splash10-0a59-0908000000-aab2ee9addb1334a7064 | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-QFT , positive | splash10-0a4i-0009000000-fd4bc4d9d4bb45e91295 | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-QFT , positive | splash10-0a4i-0409000000-115c6889c4e9205b3028 | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-QFT , positive | splash10-001i-0900000000-a2e878da4d2f6a4bb717 | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-QFT , positive | splash10-001i-0900000000-031fa259cd3b26217dad | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-QFT , positive | splash10-00lr-0900000000-e2cd3c294a2e53e1a17e | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-QFT , positive | splash10-0159-1900000000-aacbd8dce006b5ef5938 | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - , positive | splash10-0a4i-1619000000-4b248e31af7f50e87108 | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-QFT , positive | splash10-053r-0906000000-a73152b364036b45dfeb | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - 45V, Negative | splash10-0udi-0900000000-4cbb6d60c5b330f46e83 | 2021-09-20 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - 30V, Negative | splash10-0bt9-0009000000-e15320a688303951fc88 | 2021-09-20 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - 60V, Negative | splash10-0udi-0900000000-83eb5dacf9c310e76b61 | 2021-09-20 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-0a4i-0219000000-989038e0dc6e9a9ecf80 | 2016-08-01 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-0540-0984000000-b1e6f3241976f6936760 | 2016-08-01 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-0kai-1900000000-8b2d810f6330f368aa47 | 2016-08-01 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-0a4i-0159000000-0b74a48747ffb1b4f2d2 | 2016-08-03 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-0uec-3692000000-141c6eb446379ead16bb | 2016-08-03 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-0f6x-9700000000-a10d6ccf2ad58937dfa5 | 2016-08-03 | View Spectrum |
|
|---|
| Toxicity Profile |
|---|
| Route of Exposure | Following oral administration, in the fasting state, pioglitazone is first measurable in serum within 30 minutes, with peak concentrations observed within 2 hours. Food slightly delays the time to peak serum concentration to 3 to 4 hours, but does not alter the extent of absorption. |
|---|
| Mechanism of Toxicity | Pioglitazone acts as an agonist at peroxisome proliferator activated receptors (PPAR) in target tissues for insulin action such as adipose tissue, skeletal muscle, and liver. Activation of PPAR-gamma receptors increases the transcription of insulin-responsive genes involved in the control of glucose production, transport, and utilization. In this way, pioglitazone both enhances tissue sensitivity to insulin and reduces hepatic gluconeogenesis. Thus, insulin resistance associated with type 2 diabetes mellitus is improved without an increase in insulin secretion by pancreatic β cells. |
|---|
| Metabolism | Hepatic |
|---|
| Toxicity Values | Hypogycemia; LD50=mg/kg (orally in rat) |
|---|
| Lethal Dose | Not Available |
|---|
| Carcinogenicity (IARC Classification) | 2A, probably carcinogenic to humans. (4) |
|---|
| Uses/Sources | Treatment of Type II diabetes mellitus |
|---|
| Minimum Risk Level | Not Available |
|---|
| Health Effects | Not Available |
|---|
| Symptoms | Not Available |
|---|
| Treatment | Not Available |
|---|
| Normal Concentrations |
|---|
| Not Available |
|---|
| Abnormal Concentrations |
|---|
| Not Available |
|---|
| External Links |
|---|
| DrugBank ID | DB01132 |
|---|
| HMDB ID | HMDB15264 |
|---|
| PubChem Compound ID | 4829 |
|---|
| ChEMBL ID | CHEMBL595 |
|---|
| ChemSpider ID | 4663 |
|---|
| KEGG ID | C07675 |
|---|
| UniProt ID | Not Available |
|---|
| OMIM ID | |
|---|
| ChEBI ID | 8228 |
|---|
| BioCyc ID | Not Available |
|---|
| CTD ID | C060836 |
|---|
| Stitch ID | Not Available |
|---|
| PDB ID | Not Available |
|---|
| ACToR ID | Not Available |
|---|
| Wikipedia Link | Pioglitazone |
|---|
| References |
|---|
| Synthesis Reference | Chandra Khanduri, Yatendra Kumar, Atulya Panda, Suchitra Chakraborty, Mukesh Sharma, “Process for the preparation of pioglitazone.” U.S. Patent US20070078170, issued April 05, 2007. |
|---|
| MSDS | T3D4981.pdf |
|---|
| General References | - Colca JR, McDonald WG, Waldon DJ, Leone JW, Lull JM, Bannow CA, Lund ET, Mathews WR: Identification of a novel mitochondrial protein ("mitoNEET") cross-linked specifically by a thiazolidinedione photoprobe. Am J Physiol Endocrinol Metab. 2004 Feb;286(2):E252-60. Epub 2003 Oct 21. [14570702 ]
- Paddock ML, Wiley SE, Axelrod HL, Cohen AE, Roy M, Abresch EC, Capraro D, Murphy AN, Nechushtai R, Dixon JE, Jennings PA: MitoNEET is a uniquely folded 2Fe 2S outer mitochondrial membrane protein stabilized by pioglitazone. Proc Natl Acad Sci U S A. 2007 Sep 4;104(36):14342-7. Epub 2007 Aug 31. [17766440 ]
- Lincoff AM, Wolski K, Nicholls SJ, Nissen SE: Pioglitazone and risk of cardiovascular events in patients with type 2 diabetes mellitus: a meta-analysis of randomized trials. JAMA. 2007 Sep 12;298(10):1180-8. [17848652 ]
- International Agency for Research on Cancer (2014). IARC Monographs on the Evaluation of Carcinogenic Risks to Humans. [Link]
|
|---|
| Gene Regulation |
|---|
| Up-Regulated Genes | Not Available |
|---|
| Down-Regulated Genes | Not Available |
|---|