| Record Information |
|---|
| Version | 2.0 |
|---|
| Creation Date | 2009-06-08 16:35:18 UTC |
|---|
| Update Date | 2014-12-24 20:22:52 UTC |
|---|
| Accession Number | T3D0825 |
|---|
| Identification |
|---|
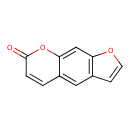
| Common Name | Psoralen |
|---|
| Class | Small Molecule |
|---|
| Description | Psoralen is found in carrot. Psoralen is found in common vegetables, e.g. parsnip, celery especially if diseased or `spoiled' Psoralen is a significant mutagen and is used for this purpose in molecular biology research.
Psoralen has been shown to exhibit anti-proliferative, anti-allergenic and anti-histamine functions (1, 2, 2).
Psoralen belongs to the family of Furanocoumarins. These are polycyclic aromatic compounds containing a furan ring fused to a coumarin moeity. |
|---|
| Compound Type | - Aromatic Hydrocarbon
- Ester
- Food Toxin
- Furocoumarin
- Industrial/Workplace Toxin
- Metabolite
- Natural Compound
- Organic Compound
- Plant Toxin
|
|---|
| Chemical Structure | |
|---|
| Synonyms | | Synonym | | 2H-Furo[3,2-g]chromen-2-one | | 3-(6-Hydroxy-5-benzofuranyl)-2-propenoic acid delta-lactone | | 5-Benzofuranacrylic acid, 6-hydroxy-, delta-lactone | | 6,7-Furanocoumarin | | 6-Hydroxy-5-benzofuranacrylic acid beta-lactone | | 6-Hydroxy-5-benzofuranacrylic acid delta-lactone | | 6-Hydroxy-5-benzofuranacrylic acid gamma-lactone | | 7H-Furo(3,2-g)(1)benzopyran-7-one | | 7H-Furo[3,2-g]benzopyran-7-one | | 7H-Furo[3,2-g]chromen-7-one | | 7H-Furo[3,2-g][1]benzopyran-7-one | | 7H-Furo[3,2-g][1]benzopyran-7-one, 9CI | | Ficusin | | Furo(2',3',7,6)coumarin | | Furo(3,2-g)-coumarin | | Furo(4',5',6,7)coumarin | | Furocoumarin | | Furo[2',3':7,6]coumarin | | Furo[2'.3':7.6]coumarin | | Furo[3,2-g]coumarin | | Furo[4',5':6,7]coumarin | | Phytoalexin | | Phytoalexin-CMPD | | Psoralene | | Psorline-p |
|
|---|
| Chemical Formula | C11H6O3 |
|---|
| Average Molecular Mass | 186.164 g/mol |
|---|
| Monoisotopic Mass | 186.032 g/mol |
|---|
| CAS Registry Number | 66-97-7 |
|---|
| IUPAC Name | 7H-furo[3,2-g]chromen-7-one |
|---|
| Traditional Name | psoralen |
|---|
| SMILES | O=C1OC2=C(C=C1)C=C1C=COC1=C2 |
|---|
| InChI Identifier | InChI=1S/C11H6O3/c12-11-2-1-7-5-8-3-4-13-9(8)6-10(7)14-11/h1-6H |
|---|
| InChI Key | InChIKey=ZCCUUQDIBDJBTK-UHFFFAOYSA-N |
|---|
| Chemical Taxonomy |
|---|
| Description | belongs to the class of organic compounds known as psoralens. These are organic compounds containing a psoralen moiety, which consists of a furan fused to a chromenone to for 7H-furo[3,2-g]chromen-7-one. |
|---|
| Kingdom | Organic compounds |
|---|
| Super Class | Phenylpropanoids and polyketides |
|---|
| Class | Coumarins and derivatives |
|---|
| Sub Class | Furanocoumarins |
|---|
| Direct Parent | Psoralens |
|---|
| Alternative Parents | |
|---|
| Substituents | - Psoralen
- Benzopyran
- 1-benzopyran
- Benzofuran
- Pyranone
- Benzenoid
- Pyran
- Furan
- Heteroaromatic compound
- Lactone
- Oxacycle
- Organoheterocyclic compound
- Organooxygen compound
- Hydrocarbon derivative
- Organic oxide
- Organic oxygen compound
- Aromatic heteropolycyclic compound
|
|---|
| Molecular Framework | Aromatic heteropolycyclic compounds |
|---|
| External Descriptors | |
|---|
| Biological Properties |
|---|
| Status | Detected and Not Quantified |
|---|
| Origin | Exogenous |
|---|
| Cellular Locations | |
|---|
| Biofluid Locations | Not Available |
|---|
| Tissue Locations | Not Available |
|---|
| Pathways | Not Available |
|---|
| Applications | Not Available |
|---|
| Biological Roles | |
|---|
| Chemical Roles | Not Available |
|---|
| Physical Properties |
|---|
| State | Solid |
|---|
| Appearance | White powder. |
|---|
| Experimental Properties | | Property | Value |
|---|
| Melting Point | 171°C | | Boiling Point | Not Available | | Solubility | Not Available | | LogP | 1.67 |
|
|---|
| Predicted Properties | |
|---|
| Spectra |
|---|
| Spectra | | Spectrum Type | Description | Splash Key | Deposition Date | View |
|---|
| GC-MS | GC-MS Spectrum - GC-EI-TOF (Non-derivatized) | splash10-0pbi-0900000000-f4158c6b7c0ecf20d9fe | 2017-09-12 | View Spectrum | | GC-MS | GC-MS Spectrum - GC-EI-TOF (Non-derivatized) | splash10-0r2i-6900000000-276c6db470fc13777449 | 2017-09-12 | View Spectrum | | GC-MS | GC-MS Spectrum - GC-EI-TOF (Non-derivatized) | splash10-0pbi-0900000000-f4158c6b7c0ecf20d9fe | 2018-05-18 | View Spectrum | | GC-MS | GC-MS Spectrum - GC-EI-TOF (Non-derivatized) | splash10-0r2i-6900000000-276c6db470fc13777449 | 2018-05-18 | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | splash10-052u-1900000000-900e13490ddfba7e89ef | 2017-07-27 | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | Not Available | 2021-10-12 | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | Not Available | 2021-10-12 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-qTof , Positive | splash10-00lr-1900000000-ac91ad62c36a29665a71 | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-QTOF , positive | splash10-000i-0900000000-75211365e04f08273288 | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-QTOF , positive | splash10-00lr-0900000000-fa9d08e8d5421ebeaa77 | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-QTOF , positive | splash10-0f89-0900000000-bacd49139ec536fb55e7 | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - , positive | splash10-00lr-1900000000-ac91ad62c36a29665a71 | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - , positive | splash10-001r-0900000000-195d9e7e3b29cf95c327 | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - 10V, Positive | splash10-000i-0900000000-75211365e04f08273288 | 2021-09-20 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - 20V, Positive | splash10-00lr-0900000000-fa9d08e8d5421ebeaa77 | 2021-09-20 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - 40V, Positive | splash10-0f89-0900000000-bacd49139ec536fb55e7 | 2021-09-20 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-000i-0900000000-287480067e742345fcb9 | 2015-05-27 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-000i-0900000000-2bf4a5ffe9d6883b015e | 2015-05-27 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-0006-1900000000-519db61c8cb6a8a31549 | 2015-05-27 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-000i-0900000000-559cba64d14574cb0041 | 2015-05-27 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-000i-0900000000-503a6a3d5ba551e5c0f5 | 2015-05-27 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-0006-0900000000-857f114f045cf9aa2b39 | 2015-05-27 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-000i-0900000000-f3f07adc83a68e9e76ff | 2021-09-23 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-000i-0900000000-f3f07adc83a68e9e76ff | 2021-09-23 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-0a4i-0900000000-a2c2c14d0cf6b23ada92 | 2021-09-23 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-000i-0900000000-89e4df10d3690e5a6d15 | 2021-09-24 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-000i-0900000000-89e4df10d3690e5a6d15 | 2021-09-24 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-000i-0900000000-89e4df10d3690e5a6d15 | 2021-09-24 | View Spectrum | | MS | Mass Spectrum (Electron Ionization) | splash10-052r-3900000000-eff7b2146482de5c7740 | 2014-09-20 | View Spectrum | | 1D NMR | 1H NMR Spectrum (1D, 300 MHz, CDCl3, experimental) | Not Available | 2014-09-20 | View Spectrum |
|
|---|
| Toxicity Profile |
|---|
| Route of Exposure | Not Available |
|---|
| Mechanism of Toxicity | The mechanism of action many furocoumarins is based on their ability to form photoadducts with DNA and other cellular components such as RNA, proteins, and several proteins found in the membrane such as phospholipases A2 and C, Ca-dependent and cAMPdependent protein-kinase and epidermal growth factor. Furocoumarins intercalate between base pairs of DNA and after ultraviolet-A irradiation, giving cycloadducts. (7). |
|---|
| Metabolism | Not Available |
|---|
| Toxicity Values | Not Available |
|---|
| Lethal Dose | Not Available |
|---|
| Carcinogenicity (IARC Classification) | Not listed by IARC. IARC has assessed other furocoumarins, classifying 8-methoxypsoralen as carcinogenic to humans (Group 1), 5-methoxypsoralen as possibly carcinogenic to humans (Group 2A), and certain other furocoumarins as not being classifiable as to their carcinogenicity to humans (Group 3). (8) |
|---|
| Uses/Sources | Anthelmintics, cross-linking reagent, photosensitising agent. Photochemical reagent for the investigation of nucleic acid structure and function. (7) |
|---|
| Minimum Risk Level | Not Available |
|---|
| Health Effects | The furocoumarin 8-methoxypsoralen is carcinogenic to humans, and possibly 5-methoxypsoralen as well (8). There is some evidence from mouse studies that other furocoumarins are carcinogenic when combined with exposure to UVA radiation (3). The SKLM regards the additional risk of skin cancer arising from the consumption of typical quantities of furocoumarin-containing foods, which remain significantly below the range of phototoxic doses, as insignificant. However, the consumption of phototoxic quantities cannot be ruled out for certain foods, particularly celery and parsnips, that may lead to significant increases in furocoumarin concentrations, depending on the storage, processing and production conditions. (9) Furocoumarin photochemotherapy is known to induce a number of side-effects including erythema, edema, hyperpigmentation, and premature aging of skin. All photobiological effects of furocoumarins result from their photochemical reactions. Because many dietary or water soluble furocoumarins are strong inhibitors of cytochrome P450s, they will also cause adverse drug reactions when taken with other drugs. |
|---|
| Symptoms | Harmful if swallowed. Irritating to eyes, respiratory system and skin. (7) |
|---|
| Treatment | In case of contact with eyes, rinse immediately with plenty of water and seek medical advice. (7) |
|---|
| Normal Concentrations |
|---|
| Not Available |
|---|
| Abnormal Concentrations |
|---|
| Not Available |
|---|
| External Links |
|---|
| DrugBank ID | Not Available |
|---|
| HMDB ID | HMDB34272 |
|---|
| PubChem Compound ID | 6199 |
|---|
| ChEMBL ID | CHEMBL164660 |
|---|
| ChemSpider ID | 5964 |
|---|
| KEGG ID | C09305 |
|---|
| UniProt ID | Not Available |
|---|
| OMIM ID | |
|---|
| ChEBI ID | 27616 |
|---|
| BioCyc ID | PHYTOALEXIN-CMPD |
|---|
| CTD ID | D005363 |
|---|
| Stitch ID | Psoralen |
|---|
| PDB ID | Not Available |
|---|
| ACToR ID | Not Available |
|---|
| Wikipedia Link | Psoralen |
|---|
| References |
|---|
| Synthesis Reference | Not Available |
|---|
| MSDS | T3D0825.pdf |
|---|
| General References | - Rodighiero P, Guiotto A, Pastorini G, Manzini P, Bordin F, Baccichetti F, Carlassare F, Vedaldi D, Dall'Acqua F: Photochemical and photobiological properties of 4,5'-dimethylpsoralen, a bifunctional contaminant of synthetic 4,5'-dimethylangelicin. Farmaco Sci. 1981 Jul;36(7):648-62. [7274446 ]
- Insawang M, Wongpraparut C: Recalcitrant solar urticaria induced by UVA and visible light: a case report. J Med Assoc Thai. 2010 Oct;93(10):1238-41. [20973330 ]
- Mullen MP, Pathak MA, West JD, Harrist TJ, Dall'Acqua F: Carcinogenic effects of monofunctional and bifunctional furocoumarins. Natl Cancer Inst Monogr. 1984 Dec;66:205-10. [6531030 ]
- Ostertag E, Becker T, Ammon J, Bauer-Aymanns H, Schrenk D: Effects of storage conditions on furocoumarin levels in intact, chopped, or homogenized parsnips. J Agric Food Chem. 2002 Apr 24;50(9):2565-70. [11958623 ]
- Santana L, Uriarte E, Roleira F, Milhazes N, Borges F: Furocoumarins in medicinal chemistry. Synthesis, natural occurrence and biological activity. Curr Med Chem. 2004 Dec;11(24):3239-61. [15579011 ]
- Yannai, Shmuel. (2004) Dictionary of food compounds with CD-ROM: Additives, flavors, and ingredients. Boca Raton: Chapman & Hall/CRC.
- Herboreal Ltd - Manufacturer of rare phytochemicals (2009). [Link]
- International Agency for Research on Cancer (2014). IARC Monographs on the Evaluation of Carcinogenic Risks to Humans. [Link]
- DFG Senate Commission on Food Safety (SKLM): Toxicological Assessment of Furocoumarins in Foodstuffs (2006) [Link]
|
|---|
| Gene Regulation |
|---|
| Up-Regulated Genes | Not Available |
|---|
| Down-Regulated Genes | Not Available |
|---|