| Record Information |
|---|
| Version | 2.0 |
|---|
| Creation Date | 2009-07-21 20:26:12 UTC |
|---|
| Update Date | 2014-12-24 20:25:49 UTC |
|---|
| Accession Number | T3D2703 |
|---|
| Identification |
|---|
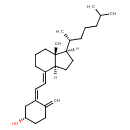
| Common Name | Cholecalciferol |
|---|
| Class | Small Molecule |
|---|
| Description | Cholecalciferol is only found in individuals that have used or taken this drug. It is a derivative of 7-dehydroxycholesterol formed by ultraviolet rays breaking of the C9-C10 bond. It differs from ergocalciferol in having a single bond between C22 and C23 and lacking a methyl group at C24. [PubChem]The first step involved in the activation of vitamin D3 is a 25-hydroxylation which is catalysed by the 25-hydroxylase in the liver and then by other enzymes. The mitochondrial sterol 27-hydroxylase catalyses the first reaction in the oxidation of the side chain of sterol intermediates. The active form of vitamin D3 (calcitriol) binds to intracellular receptors that then function as transcription factors to modulate gene expression. Like the receptors for other steroid hormones and thyroid hormones, the vitamin D receptor has hormone-binding and DNA-binding domains. The vitamin D receptor forms a complex with another intracellular receptor, the retinoid-X receptor, and that heterodimer is what binds to DNA. In most cases studied, the effect is to activate transcription, but situations are also known in which vitamin D suppresses transcription. Calcitriol increases the serum calcium concentrations by: increasing GI absorption of phosphorus and calcium, increasing osteoclastic resorption, and increasing distal renal tubular reabsorption of calcium. Calcitriol appears to promote intestinal absorption of calcium through binding to the vitamin D receptor in the mucosal cytoplasm of the intestine. Subsequently, calcium is absorbed through formation of a calcium-binding protein. |
|---|
| Compound Type | - Antihypocalcemic Agent
- Antihypoparathyroid Agent
- Antithyroid Agent
- Bone Density Conservation Agent
- Drug
- Essential Vitamin
- Food Toxin
- Household Toxin
- Metabolite
- Nutraceutical
- Organic Compound
- Synthetic Compound
- Vitamin
- Vitamin D
|
|---|
| Chemical Structure | |
|---|
| Synonyms | | Synonym | | (+)-vitamin D3 | | (1S,3Z)-3-[(2e)-2-[(1R,3AR,7as)-7a-methyl-1-[(2R)-6-methylheptan-2-yl]-2,3,3a,5,6,7-hexahydro-1H-inden-4-ylidene]ethylidene]-4-methylidene-cyclohexan-1-ol | | (3beta,5Z,7e)-9,10-Secocholesta-5,7,10(19)-trien-3-ol | | (5Z,7e)-(3S)-9,10-Secocholesta-5,7,10(19)-trien-3-ol | | Activated 7-dehydrocholesterol | | Calciol | | CC | | Colecalciferol | | delta-D | | Micro-D | | Oleovitamin D3 | | Optimal-D | | Vigantol | | Vitamin D3 |
|
|---|
| Chemical Formula | C27H44O |
|---|
| Average Molecular Mass | 384.638 g/mol |
|---|
| Monoisotopic Mass | 384.339 g/mol |
|---|
| CAS Registry Number | 67-97-0 |
|---|
| IUPAC Name | (1S,3Z)-3-{2-[(1R,3aS,4E,7aR)-7a-methyl-1-[(2R)-6-methylheptan-2-yl]-octahydro-1H-inden-4-ylidene]ethylidene}-4-methylidenecyclohexan-1-ol |
|---|
| Traditional Name | vitamin D3 |
|---|
| SMILES | [H]\C(\C(\[H])=C1/CCC[C@@]2(C)[C@@]1([H])CC[C@]2([H])[C@]([H])(C)CCCC(C)C)=C1/C[C@@]([H])(O)CCC1=C |
|---|
| InChI Identifier | InChI=1S/C27H44O/c1-19(2)8-6-9-21(4)25-15-16-26-22(10-7-17-27(25,26)5)12-13-23-18-24(28)14-11-20(23)3/h12-13,19,21,24-26,28H,3,6-11,14-18H2,1-2,4-5H3/b22-12+,23-13-/t21-,24+,25-,26+,27-/m1/s1 |
|---|
| InChI Key | InChIKey=QYSXJUFSXHHAJI-YRZJJWOYSA-N |
|---|
| Chemical Taxonomy |
|---|
| Description | belongs to the class of organic compounds known as vitamin d and derivatives. Vitamin D and derivatives are compounds containing a secosteroid backbone, usually secoergostane or secocholestane. |
|---|
| Kingdom | Organic compounds |
|---|
| Super Class | Lipids and lipid-like molecules |
|---|
| Class | Steroids and steroid derivatives |
|---|
| Sub Class | Vitamin D and derivatives |
|---|
| Direct Parent | Vitamin D and derivatives |
|---|
| Alternative Parents | |
|---|
| Substituents | - Triterpenoid
- Cyclic alcohol
- Secondary alcohol
- Organic oxygen compound
- Hydrocarbon derivative
- Organooxygen compound
- Alcohol
- Aliphatic homopolycyclic compound
|
|---|
| Molecular Framework | Aliphatic homopolycyclic compounds |
|---|
| External Descriptors | |
|---|
| Biological Properties |
|---|
| Status | Detected and Not Quantified |
|---|
| Origin | Endogenous |
|---|
| Cellular Locations | - Cytoplasm
- Extracellular
- Membrane
|
|---|
| Biofluid Locations | Not Available |
|---|
| Tissue Locations | Not Available |
|---|
| Pathways | Not Available |
|---|
| Applications | Not Available |
|---|
| Biological Roles | |
|---|
| Chemical Roles | Not Available |
|---|
| Physical Properties |
|---|
| State | Solid |
|---|
| Appearance | White powder. |
|---|
| Experimental Properties | | Property | Value |
|---|
| Melting Point | 84.5°C | | Boiling Point | 496.4°C | | Solubility | Insoluble | | LogP | 7.5 |
|
|---|
| Predicted Properties | |
|---|
| Spectra |
|---|
| Spectra | | Spectrum Type | Description | Splash Key | Deposition Date | View |
|---|
| Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | splash10-0aou-3029000000-1950c74de34369a70400 | 2017-09-01 | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (1 TMS) - 70eV, Positive | splash10-002f-9207800000-c6f808a014153de38d58 | 2017-10-06 | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | Not Available | 2021-10-12 | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | Not Available | 2021-10-12 | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TBDMS_1_1) - 70eV, Positive | Not Available | 2021-11-06 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-014r-0129000000-80de8aabbfb587d8f7b9 | 2016-08-01 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-0abi-2595000000-1f5db35cc732efef5e27 | 2016-08-01 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-0pb9-7195000000-b742ddd81b2f98395fab | 2016-08-01 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-001i-0009000000-70f22151bd69b3307086 | 2016-08-03 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-001i-0009000000-7ff3c3cf4e98856a166b | 2016-08-03 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-014i-1249000000-8756607b9fd7d6008ac9 | 2016-08-03 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-000i-0459000000-cdb5561f3458bde4c2f1 | 2021-09-22 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-0kmj-7494000000-5a36ac0547956f9ac058 | 2021-09-22 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-0a4i-3930000000-c62b5a8a92e3487b90ed | 2021-09-22 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-001i-0009000000-bfd82bd18d8804cfa1d2 | 2021-09-23 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-001i-0009000000-d168cadb843f05c02d05 | 2021-09-23 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-001i-0539000000-277213cec95d0ba155c5 | 2021-09-23 | View Spectrum | | 1D NMR | 1H NMR Spectrum (1D, 600 MHz, CDCl3, experimental) | Not Available | 2019-05-16 | View Spectrum | | 2D NMR | [1H, 13C]-HSQC NMR Spectrum (2D, 600 MHz, CDCl3, experimental) | Not Available | 2019-05-16 | View Spectrum |
|
|---|
| Toxicity Profile |
|---|
| Route of Exposure | Oral, readily absorbed |
|---|
| Mechanism of Toxicity | The first step involved in the activation of vitamin D3 is a 25-hydroxylation which is catalysed by the 25-hydroxylase in the liver and then by other enzymes. The mitochondrial sterol 27-hydroxylase catalyses the first reaction in the oxidation of the side chain of sterol intermediates. The active form of vitamin D3 (calcitriol) binds to intracellular receptors that then function as transcription factors to modulate gene expression. Like the receptors for other steroid hormones and thyroid hormones, the vitamin D receptor has hormone-binding and DNA-binding domains. The vitamin D receptor forms a complex with another intracellular receptor, the retinoid-X receptor, and that heterodimer is what binds to DNA. In most cases studied, the effect is to activate transcription, but situations are also known in which vitamin D suppresses transcription. Calcitriol increases the serum calcium concentrations by: increasing GI absorption of phosphorus and calcium, increasing osteoclastic resorption, and increasing distal renal tubular reabsorption of calcium. Calcitriol appears to promote intestinal absorption of calcium through binding to the vitamin D receptor in the mucosal cytoplasm of the intestine. Subsequently, calcium is absorbed through formation of a calcium-binding protein. |
|---|
| Metabolism | Within the liver, cholecalciferal is hydroxylated to calcidiol (25-hydroxycholecalciferol) by the enzyme 25-hydroxylase. Within the kidney, calcidiol serves as a substrate for 1-alpha-hydroxylase, yielding calcitriol (1,25-dihydroxycholecalciferol), the biologically active form of vitamin D3.
Half Life: Several weeks |
|---|
| Toxicity Values | Not Available |
|---|
| Lethal Dose | Not Available |
|---|
| Carcinogenicity (IARC Classification) | No indication of carcinogenicity to humans (not listed by IARC). |
|---|
| Uses/Sources | For the treatment of vitamin D deficiency or insufficiency, refractory rickets (vitamin D resistant rickets), familial hypophosphatemia and hypoparathyroidism, and in the management of hypocalcemia and renal osteodystrophy in patients with chronic renal failure undergoing dialysis. Also used in conjunction with calcium in the management and prevention of primary or corticosteroid-induced osteoporosis. |
|---|
| Minimum Risk Level | Not Available |
|---|
| Health Effects | Not Available |
|---|
| Symptoms | Hypercalcemia - Early symptoms of hypercalcemia, include nausea and vomiting, weakness, headache, somnolence, dry mouth, constipation, metallic taste, muscle pain and bone pain. Late symptoms and signs of hypercalcemia, include polyuria, polydipsia, anorexia, weight loss, nocturia, conjunctivitis, pancreatitis, photophobia, rhinorrhea, pruritis, hyperthermia, decreased libido, elevated BUN, albuminuria, hypercholesterolemia, elevated ALT (SGPT) and AST (SGOT), ectopic calcification, nephrocalcinosis, hypertension and cardiac arrhythmias. |
|---|
| Treatment | Not Available |
|---|
| Normal Concentrations |
|---|
| Not Available |
|---|
| Abnormal Concentrations |
|---|
| Not Available |
|---|
| External Links |
|---|
| DrugBank ID | DB00169 |
|---|
| HMDB ID | HMDB14315 |
|---|
| PubChem Compound ID | 5280795 |
|---|
| ChEMBL ID | CHEMBL1042 |
|---|
| ChemSpider ID | 4444353 |
|---|
| KEGG ID | C05443 |
|---|
| UniProt ID | Not Available |
|---|
| OMIM ID | 270400 |
|---|
| ChEBI ID | 28940 |
|---|
| BioCyc ID | VITAMIN_D_%7B3%7D |
|---|
| CTD ID | Not Available |
|---|
| Stitch ID | Cholecalciferol |
|---|
| PDB ID | Not Available |
|---|
| ACToR ID | 7582 |
|---|
| Wikipedia Link | Cholecalciferol |
|---|
| References |
|---|
| Synthesis Reference | Jean Jolly, Primo Rizzi, Jean Taillardat, “1.alpha.,25.alpha.-Dihydroxy-cholecalciferol and methods for the production thereof.” U.S. Patent US4435325, issued May, 1977. |
|---|
| MSDS | Link |
|---|
| General References | - Armas LA, Hollis BW, Heaney RP: Vitamin D2 is much less effective than vitamin D3 in humans. J Clin Endocrinol Metab. 2004 Nov;89(11):5387-91. [15531486 ]
|
|---|
| Gene Regulation |
|---|
| Up-Regulated Genes | | Gene | Gene Symbol | Gene ID | Interaction | Chromosome | Details |
|---|
|
|---|
| Down-Regulated Genes | | Gene | Gene Symbol | Gene ID | Interaction | Chromosome | Details |
|---|
|
|---|