| Record Information |
|---|
| Version | 2.0 |
|---|
| Creation Date | 2009-07-21 20:27:15 UTC |
|---|
| Update Date | 2014-12-24 20:25:52 UTC |
|---|
| Accession Number | T3D2838 |
|---|
| Identification |
|---|
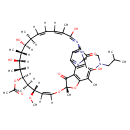
| Common Name | Rifabutin |
|---|
| Class | Small Molecule |
|---|
| Description | Rifabutin is only found in individuals that have used or taken this drug. It is a broad-spectrum antibiotic that is being used as prophylaxis against disseminated Mycobacterium avium complex infection in HIV-positive patients. [PubChem]Rifabutin acts via the inhibition of DNA-dependent RNA polymerase in gram-positive and some gram-negative bacteria, leading to a suppression of RNA synthesis and cell death. |
|---|
| Compound Type | - Amide
- Amine
- Anti-Bacterial Agent
- Antibiotic, Antitubercular
- Drug
- Ester
- Ether
- Metabolite
- Organic Compound
- Synthetic Compound
|
|---|
| Chemical Structure | |
|---|
| Synonyms | | Synonym | | 4-Deoxo-3,4-(2-spiro(N-isobutyl-4-piperidyl)-2,5-dihydro-1H-imidazo)-rifamycin S | | 4-N-Isobutylspiropiperidylrifamycin S | | Ansamicin | | Ansamycin | | Ansatipin | | Ansatipine | | Antibiotic LM 427 | | Mycobutin | | RBT | | Ributin | | Rifabutina | | Rifabutine | | Rifabutinum |
|
|---|
| Chemical Formula | C46H62N4O11 |
|---|
| Average Molecular Mass | 847.005 g/mol |
|---|
| Monoisotopic Mass | 846.442 g/mol |
|---|
| CAS Registry Number | 72559-06-9 |
|---|
| IUPAC Name | (7S,9Z,11S,12R,13S,14R,15R,16R,17S,18S,19Z,21Z)-2,15,17,23-tetrahydroxy-11-methoxy-3,7,12,14,16,18,22-heptamethyl-1'-(2-methylpropyl)-6,32-dioxo-8,33-dioxa-24,27,29-triazaspiro[pentacyclo[23.6.1.1^{4,7}.0^{5,31}.0^{26,30}]tritriacontane-28,4'-piperidine]-1(31),2,4,9,19,21,23,25,29-nonaen-13-yl acetate |
|---|
| Traditional Name | (7S,9Z,11S,12R,13S,14R,15R,16R,17S,18S,19Z,21Z)-2,15,17,23-tetrahydroxy-11-methoxy-3,7,12,14,16,18,22-heptamethyl-1'-(2-methylpropyl)-6,32-dioxo-8,33-dioxa-24,27,29-triazaspiro[pentacyclo[23.6.1.1^{4,7}.0^{5,31}.0^{26,30}]tritriacontane-28,4'-piperidine]-1(31),2,4,9,19,21,23,25,29-nonaen-13-yl acetate |
|---|
| SMILES | [H]\C1=C([H])\[C@]([H])(OC)[C@@]([H])(C)[C@@]([H])(OC(C)=O)[C@]([H])(C)[C@]([H])(O)[C@]([H])(C)[C@@]([H])(O)[C@@]([H])(C)\C([H])=C(\[H])/C(/[H])=C(C)\C(O)=NC2=C3NC4(CCN(CC(C)C)CC4)N=C3C3=C(C(O)=C(C)C4=C3C(=O)[C@](C)(O4)O1)C2=O |
|---|
| InChI Identifier | InChI=1S/C46H62N4O11/c1-22(2)21-50-18-16-46(17-19-50)48-34-31-32-39(54)28(8)42-33(31)43(56)45(10,61-42)59-20-15-30(58-11)25(5)41(60-29(9)51)27(7)38(53)26(6)37(52)23(3)13-12-14-24(4)44(57)47-36(40(32)55)35(34)49-46/h12-15,20,22-23,25-27,30,37-38,41,49,52-54H,16-19,21H2,1-11H3,(H,47,57)/b13-12-,20-15-,24-14-/t23-,25+,26+,27+,30-,37-,38+,41+,45-/m0/s1 |
|---|
| InChI Key | InChIKey=ATEBXHFBFRCZMA-DVAZEERGSA-N |
|---|
| Chemical Taxonomy |
|---|
| Description | belongs to the class of organic compounds known as macrolactams. These are cyclic amides of amino carboxylic acids, having a 1-azacycloalkan-2-one structure, or analogues having unsaturation or heteroatoms replacing one or more carbon atoms of the ring. They are nitrogen analogues (the a nitrogen atom replacing the o atom of the cyclic carboxylic acid group ) of the naturally occurring macrolides. |
|---|
| Kingdom | Organic compounds |
|---|
| Super Class | Phenylpropanoids and polyketides |
|---|
| Class | Macrolactams |
|---|
| Sub Class | Not Available |
|---|
| Direct Parent | Macrolactams |
|---|
| Alternative Parents | |
|---|
| Substituents | - Naphthofuran
- Macrolactam
- Azaspirodecane
- Naphthalene
- Benzofuran
- Coumaran
- Aryl ketone
- Aryl alkyl ketone
- Ketal
- Piperidine
- Benzenoid
- 3-imidazoline
- Vinylogous amide
- Vinylogous acid
- Lactam
- Ketimine
- Amino acid or derivatives
- Tertiary aliphatic amine
- Tertiary amine
- Secondary carboxylic acid amide
- Carboxylic acid ester
- Ketone
- Carboxamide group
- Secondary alcohol
- Acetal
- Carboxylic acid derivative
- Dialkyl ether
- Secondary aliphatic amine
- Enamine
- Ether
- Oxacycle
- Azacycle
- Organoheterocyclic compound
- Organic 1,3-dipolar compound
- Propargyl-type 1,3-dipolar organic compound
- Monocarboxylic acid or derivatives
- Polyol
- Secondary amine
- Organic oxygen compound
- Organopnictogen compound
- Imine
- Carbonyl group
- Organic oxide
- Alcohol
- Hydrocarbon derivative
- Amine
- Organic nitrogen compound
- Organonitrogen compound
- Organooxygen compound
- Aromatic heteropolycyclic compound
|
|---|
| Molecular Framework | Aromatic heteropolycyclic compounds |
|---|
| External Descriptors | Not Available |
|---|
| Biological Properties |
|---|
| Status | Detected and Not Quantified |
|---|
| Origin | Exogenous |
|---|
| Cellular Locations | |
|---|
| Biofluid Locations | Not Available |
|---|
| Tissue Locations | Not Available |
|---|
| Pathways | Not Available |
|---|
| Applications | |
|---|
| Biological Roles | |
|---|
| Chemical Roles | Not Available |
|---|
| Physical Properties |
|---|
| State | Solid |
|---|
| Appearance | White powder. |
|---|
| Experimental Properties | | Property | Value |
|---|
| Melting Point | Not Available | | Boiling Point | Not Available | | Solubility | Minimally soluble (0.19 mg/mL) | | LogP | 4.1 |
|
|---|
| Predicted Properties | |
|---|
| Spectra |
|---|
| Spectra | | Spectrum Type | Description | Splash Key | Deposition Date | View |
|---|
| Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-06vi-2000000290-b6c2e05a4e08c21c8009 | 2016-08-03 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-0a4i-8000002970-8db040b5d2c5d1c7f870 | 2016-08-03 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-0a4i-9000003500-61568975d849f4e01490 | 2016-08-03 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-0002-1000000390-013052b1c69581676709 | 2016-08-04 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-0kbs-2000000790-222783e8f7fc477df855 | 2016-08-04 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-000m-3000001910-593444223c16d943f93e | 2016-08-04 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-05pa-0000000970-2dfaa272c475120f1997 | 2021-10-11 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-00kr-0000000930-3f004468a35f4bfc7d81 | 2021-10-11 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-00kk-0000000950-e0fa551433e76d0694bb | 2021-10-11 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-0002-1000000290-6b79ec41174e5c65892f | 2021-10-11 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-0a4i-6000000930-138088289c4e164844b3 | 2021-10-11 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-000l-2000000940-3d9bc441c2d70d547151 | 2021-10-11 | View Spectrum |
|
|---|
| Toxicity Profile |
|---|
| Route of Exposure | Rifabutin is readily absorbed from the gastrointestinal tract, with an absolute bioavailability averaging 20%. |
|---|
| Mechanism of Toxicity | Rifabutin acts via the inhibition of DNA-dependent RNA polymerase in gram-positive and some gram-negative bacteria, leading to a suppression of RNA synthesis and cell death. |
|---|
| Metabolism | Hepatic. Of the five metabolites that have been identified, 25-O-desacetyl and 31-hydroxy are the most predominant. The former metabolite has an activity equal to the parent drug and contributes up to 10% to the total antimicrobial activity.
Route of Elimination: A mass-balance study in three healthy adult volunteers with 14C-labeled rifabutin showed that 53% of the oral dose was excreted in the urine, primarily as metabolites. About 30% of the dose is excreted in the feces.
Half Life: 45 (± 17) hours |
|---|
| Toxicity Values | LD50: 4.8 g/kg (male mouse) |
|---|
| Lethal Dose | Not Available |
|---|
| Carcinogenicity (IARC Classification) | No indication of carcinogenicity to humans (not listed by IARC). |
|---|
| Uses/Sources | For the prevention of disseminated Mycobacterium avium complex (MAC) disease in patients with advanced HIV infection. Used in the treatment of tuberculosis. It has also found to be useful in the treatment of (Chlamydia) Chlamydophila pneumoniae (Cpn) Infection.[Wikipedia] |
|---|
| Minimum Risk Level | Not Available |
|---|
| Health Effects | Not Available |
|---|
| Symptoms | Symptoms include diarrhea, eructation, abdominal pain, vomitting, nausea, and insomnia. |
|---|
| Treatment | Clinical experience with rifamycins suggests that gastric lavage to evacuate gastric contents (within a few hours of overdose), followed by instillation of an activated charcoal slurry into the stomach, may help absorb any remaining drug from the gastrointestinal tract. (2) |
|---|
| Normal Concentrations |
|---|
| Not Available |
|---|
| Abnormal Concentrations |
|---|
| Not Available |
|---|
| External Links |
|---|
| DrugBank ID | DB00615 |
|---|
| HMDB ID | HMDB14753 |
|---|
| PubChem Compound ID | 6323490 |
|---|
| ChEMBL ID | CHEMBL444633 |
|---|
| ChemSpider ID | 4883445 |
|---|
| KEGG ID | C07235 |
|---|
| UniProt ID | Not Available |
|---|
| OMIM ID | |
|---|
| ChEBI ID | 8857 |
|---|
| BioCyc ID | Not Available |
|---|
| CTD ID | Not Available |
|---|
| Stitch ID | Rifabutin |
|---|
| PDB ID | RBT |
|---|
| ACToR ID | Not Available |
|---|
| Wikipedia Link | Rifabutin |
|---|
| References |
|---|
| Synthesis Reference | Not Available |
|---|
| MSDS | Link |
|---|
| General References | - Drugs.com [Link]
- RxList: The Internet Drug Index (2009). [Link]
|
|---|
| Gene Regulation |
|---|
| Up-Regulated Genes | | Gene | Gene Symbol | Gene ID | Interaction | Chromosome | Details |
|---|
|
|---|
| Down-Regulated Genes | Not Available |
|---|