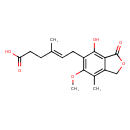
Mycophenolic acid (T3D2960)
| Record Information | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Version | 2.0 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Creation Date | 2009-07-21 20:28:11 UTC | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Update Date | 2014-12-24 20:25:54 UTC | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Accession Number | T3D2960 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Identification | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Common Name | Mycophenolic acid | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Class | Small Molecule | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Description | Mycophenolic acid is an an immunosuppresant drug and potent anti-proliferative, and can be used in place of the older anti-proliferative azathioprine. It is usually used as part of triple therapy including a calcineurin inhibitor (ciclosporin or tacrolimus) and prednisolone. It is also useful in research for the selection of animal cells that express the E. coli gene coding for XGPRT (xanthine guanine phosphoribosyltransferase). | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Compound Type |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Chemical Structure | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Synonyms |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Chemical Formula | C17H20O6 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Average Molecular Mass | 320.337 g/mol | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Monoisotopic Mass | 320.126 g/mol | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| CAS Registry Number | 24280-93-1 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| IUPAC Name | (4E)-6-(4-hydroxy-6-methoxy-7-methyl-3-oxo-1,3-dihydro-2-benzofuran-5-yl)-4-methylhex-4-enoic acid | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Traditional Name | mycophenolic acid | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| SMILES | [H]C(CC1=C(O)C2=C(COC2=O)C(C)=C1OC)=C(C)CCC(O)=O | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| InChI Identifier | InChI=1S/C17H20O6/c1-9(5-7-13(18)19)4-6-11-15(20)14-12(8-23-17(14)21)10(2)16(11)22-3/h4,20H,5-8H2,1-3H3,(H,18,19)/b9-4+ | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| InChI Key | InChIKey=HPNSFSBZBAHARI-RUDMXATFSA-N | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Chemical Taxonomy | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Description | belongs to the class of organic compounds known as phthalides. Phthalides are compounds containing a 3-hydrocarbylidene-2-benzofuran-1(3H)-one moiety,. | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Kingdom | Organic compounds | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Super Class | Organoheterocyclic compounds | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Class | Isocoumarans | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Sub Class | Isobenzofuranones | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Direct Parent | Phthalides | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Alternative Parents |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Substituents |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Molecular Framework | Aromatic heteropolycyclic compounds | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| External Descriptors |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Biological Properties | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Status | Detected and Not Quantified | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Origin | Exogenous | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Cellular Locations |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Biofluid Locations | Not Available | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Tissue Locations |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Pathways | Not Available | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Applications | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Biological Roles | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Chemical Roles | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Physical Properties | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| State | Solid | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Appearance | White powder. | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Experimental Properties |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Predicted Properties |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Spectra | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Spectra |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Toxicity Profile | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Route of Exposure | Bioavailability following oral administration of Myfortic delayed-release tablet ranges from 70-95% | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Mechanism of Toxicity | Mycophenolic acid is a potent, selective, uncompetitive, and reversible inhibitor of inosine monophosphate dehydrogenase (IMPDH), and therefore inhibits the de novo pathway of guanosine nucleotide synthesis without incorporation into DNA. Because T- and B-lymphocytes are critically dependent for their proliferation on de novo synthesis of purines, whereas other cell types can utilize salvage pathways, mycophenolic acid has potent cytostatic effects on lymphocytes. Mycophenolic acid inhibits proliferative responses of T- and B-lymphocytes to both mitogenic and allospecific stimulation. Addition of guanosine or deoxyguanosine reverses the cytostatic effects of mycophenolic acid on lymphocytes. Mycophenolic acid also suppresses antibody formation by B-lymphocytes. Mycophenolic acid prevents the glycosylation of lymphocyte and monocyte glycoproteins that are involved in intercellular adhesion to endothelial cells and may inhibit recruitment of leukocytes into sites of inflammation and graft rejection. | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Metabolism | Mycophenolic acid is metabolized mainly by glucuronyl transferase to glucuronidated metabolites, predominantly the phenolic glucuronide, mycophenolic acid glucuronide (MPAG). MPAG does not manifest pharmacological activity. The acyl glucuronide minor metabolite has pharmacological activity similar to mycophenolic acid. The AUC ratio of Mycophenolic acid:MPAG:acyl glucuronide is approximately 1:24:0.28 at steady state. Half Life: The mean elimination half-life for mycophenolic acid ranges from 8-16 hours, while that of the MPAG metabolite ranges from 13-17 hours. | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Toxicity Values | LD50: 352 mg/kg (Oral, Rat) (2) LD50: 1000 mg/kg (Oral, Mouse) (2) LD50: >6000 mg/kg (Oral, Rabbit) (2) | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Lethal Dose | Not Available | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Carcinogenicity (IARC Classification) | No indication of carcinogenicity to humans (not listed by IARC). | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Uses/Sources | For the prophylaxis of organ rejection in patients receiving allogeneic renal transplants, administered in combination with cyclosporine and corticosteroids. | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Minimum Risk Level | Not Available | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Health Effects | Not Available | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symptoms | Possible signs and symptoms of acute overdose could include the following: hematological abnormalities such as leukopenia and neutropenia, and gastrointestinal symptoms such as abdominal pain, diarrhea, nausea and vomiting, and dyspepsia. | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Treatment | Not Available | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Normal Concentrations | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Abnormal Concentrations | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| External Links | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| DrugBank ID | DB01024 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| HMDB ID | HMDB15159 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PubChem Compound ID | 446541 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| ChEMBL ID | CHEMBL866 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| ChemSpider ID | 393865 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| KEGG ID | C20380 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| UniProt ID | Not Available | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| OMIM ID | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| ChEBI ID | 168396 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| BioCyc ID | Not Available | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| CTD ID | Not Available | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Stitch ID | Mycophenolic acid | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PDB ID | MOA | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| ACToR ID | Not Available | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Wikipedia Link | Mycophenolic_acid | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| References | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Synthesis Reference | Bernard J. Abbott, John G. Whitney, “Method of preparing mycophenolic acid glucoside.” U.S. Patent US4234684, issued January, 1976. | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| MSDS | Link | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| General References |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Gene Regulation | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Up-Regulated Genes |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Down-Regulated Genes | Not Available | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Targets
- General Function:
- Rna binding
- Specific Function:
- Catalyzes the conversion of inosine 5'-phosphate (IMP) to xanthosine 5'-phosphate (XMP), the first committed and rate-limiting step in the de novo synthesis of guanine nucleotides, and therefore plays an important role in the regulation of cell growth. Could also have a single-stranded nucleic acid-binding activity and could play a role in RNA and/or DNA metabolism. It may also have a role in the development of malignancy and the growth progression of some tumors.
- Gene Name:
- IMPDH2
- Uniprot ID:
- P12268
- Molecular Weight:
- 55804.495 Da
Binding/Activity Constants
| Type | Value | Assay Type | Assay Source |
|---|---|---|---|
| Inhibitory | 0.006 uM | Not Available | BindingDB 19264 |
| Inhibitory | 0.007 uM | Not Available | BindingDB 19264 |
| Inhibitory | 0.01 uM | Not Available | BindingDB 19264 |
| IC50 | 0.011 uM | Not Available | BindingDB 19264 |
| IC50 | 0.012 uM | Not Available | BindingDB 19264 |
| IC50 | 0.014 uM | Not Available | BindingDB 19264 |
| IC50 | 0.015 uM | Not Available | BindingDB 19264 |
| IC50 | 0.016 uM | Not Available | BindingDB 19264 |
| IC50 | 0.0248 uM | Not Available | BindingDB 19264 |
| IC50 | 0.0251 uM | Not Available | BindingDB 19264 |
| IC50 | 0.63 uM | Not Available | BindingDB 19264 |
| IC50 | 1.5 uM | Not Available | BindingDB 19264 |
References
- Vannozzi F, Filipponi F, Di Paolo A, Danesi R, Urbani L, Bocci G, Catalano G, De Simone P, Mosca F, Del Tacca M: An exploratory study on pharmacogenetics of inosine-monophosphate dehydrogenase II in peripheral mononuclear cells from liver-transplant recipients. Transplant Proc. 2004 Nov;36(9):2787-90. [15621150 ]
- Wang J, Zeevi A, Webber S, Girnita DM, Addonizio L, Selby R, Hutchinson IV, Burckart GJ: A novel variant L263F in human inosine 5'-monophosphate dehydrogenase 2 is associated with diminished enzyme activity. Pharmacogenet Genomics. 2007 Apr;17(4):283-90. [17496727 ]
- Penuelas S, Noe V, Morales R, Ciudad CJ: Sensitization of human erythroleukemia K562 cells resistant to methotrexate by inhibiting IMPDH. Med Sci Monit. 2005 Jan;11(1):BR6-12. [15614187 ]
- Yam P, Jensen M, Akkina R, Anderson J, Villacres MC, Wu J, Zaia JA, Yee JK: Ex vivo selection and expansion of cells based on expression of a mutated inosine monophosphate dehydrogenase 2 after HIV vector transduction: effects on lymphocytes, monocytes, and CD34+ stem cells. Mol Ther. 2006 Aug;14(2):236-44. Epub 2006 May 2. [16647299 ]
- Dzidic A, Prgomet C, Mohr A, Meyer K, Bauer J, Meyer HH, Pfaffl MW: Effects of mycophenolic acid on inosine monophosphate dehydrogenase I and II mRNA expression in white blood cells and various tissues in sheep. J Vet Med A Physiol Pathol Clin Med. 2006 May;53(4):163-9. [16629948 ]
- Watkins WJ, Chen JM, Cho A, Chong L, Collins N, Fardis M, Huang W, Hung M, Kirschberg T, Lee WA, Liu X, Thomas W, Xu J, Zeynalzadegan A, Zhang J: Phosphonic acid-containing analogues of mycophenolic acid as inhibitors of IMPDH. Bioorg Med Chem Lett. 2006 Jul 1;16(13):3479-83. Epub 2006 Apr 18. [16621550 ]
- Mitsuhashi S, Takenaka J, Iwamori K, Nakajima N, Ubukata M: Structure-activity relationships for inhibition of inosine monophosphate dehydrogenase and differentiation induction of K562 cells among the mycophenolic acid derivatives. Bioorg Med Chem. 2010 Nov 15;18(22):8106-11. doi: 10.1016/j.bmc.2010.09.004. Epub 2010 Sep 18. [20934342 ]
- Dhar TG, Shen Z, Guo J, Liu C, Watterson SH, Gu HH, Pitts WJ, Fleener CA, Rouleau KA, Sherbina NZ, McIntyre KW, Shuster DJ, Witmer MR, Tredup JA, Chen BC, Zhao R, Bednarz MS, Cheney DL, MacMaster JF, Miller LM, Berry KK, Harper TW, Barrish JC, Hollenbaugh DL, Iwanowicz EJ: Discovery of N-[2-[2-[[3-methoxy-4-(5-oxazolyl)phenyl]amino]-5-oxazolyl]phenyl]-N-methyl-4- morpholineacetamide as a novel and potent inhibitor of inosine monophosphate dehydrogenase with excellent in vivo activity. J Med Chem. 2002 May 23;45(11):2127-30. [12014950 ]
- Watterson SH, Carlsen M, Dhar TG, Shen Z, Pitts WJ, Guo J, Gu HH, Norris D, Chorba J, Chen P, Cheney D, Witmer M, Fleener CA, Rouleau K, Townsend R, Hollenbaugh DL, Iwanowicz EJ: Novel inhibitors of IMPDH: a highly potent and selective quinolone-based series. Bioorg Med Chem Lett. 2003 Feb 10;13(3):543-6. [12565968 ]
- Dhar TG, Watterson SH, Chen P, Shen Z, Gu HH, Norris D, Carlsen M, Haslow KD, Pitts WJ, Guo J, Chorba J, Fleener CA, Rouleau KA, Townsend R, Hollenbaugh D, Iwanowicz EJ: Quinolone-based IMPDH inhibitors: introduction of basic residues on ring D and SAR of the corresponding mono, di and benzofused analogues. Bioorg Med Chem Lett. 2003 Feb 10;13(3):547-51. [12565969 ]
- Pitts WJ, Guo J, Dhar TG, Shen Z, Gu HH, Watterson SH, Bednarz MS, Chen BC, Barrish JC, Bassolino D, Cheney D, Fleener CA, Rouleau KA, Hollenbaugh DL, Iwanowicz EJ: Rapid synthesis of triazine inhibitors of inosine monophosphate dehydrogenase. Bioorg Med Chem Lett. 2002 Aug 19;12(16):2137-40. [12127522 ]
- Iwanowicz EJ, Watterson SH, Liu C, Gu HH, Mitt T, Leftheris K, Barrish JC, Fleener CA, Rouleau K, Sherbina NZ, Hollenbaugh DL: Novel guanidine-based inhibitors of inosine monophosphate dehydrogenase. Bioorg Med Chem Lett. 2002 Oct 21;12(20):2931-4. [12270177 ]
- Dhar TG, Liu C, Pitts WJ, Guo J, Watterson SH, Gu H, Fleener CA, Rouleau K, Sherbina NZ, Barrish JC, Hollenbaugh D, Iwanowicz EJ: A survey of cyclic replacements for the central diamide moiety of inhibitors of inosine monophosphate dehydrogenase. Bioorg Med Chem Lett. 2002 Nov 4;12(21):3125-8. [12372516 ]
- Dhar TG, Shen Z, Fleener CA, Rouleau KA, Barrish JC, Hollenbaugh DL, Iwanowicz EJ: The TosMIC approach to 3-(oxazol-5-yl) indoles: application to the synthesis of indole-based IMPDH inhibitors. Bioorg Med Chem Lett. 2002 Nov 18;12(22):3305-8. [12392738 ]
- Dhar TG, Shen Z, Gu HH, Chen P, Norris D, Watterson SH, Ballentine SK, Fleener CA, Rouleau KA, Barrish JC, Townsend R, Hollenbaugh DL, Iwanowicz EJ: 3-cyanoindole-based inhibitors of inosine monophosphate dehydrogenase: synthesis and initial structure-activity relationships. Bioorg Med Chem Lett. 2003 Oct 20;13(20):3557-60. [14505670 ]
- Watterson SH, Chen P, Zhao Y, Gu HH, Dhar TG, Xiao Z, Ballentine SK, Shen Z, Fleener CA, Rouleau KA, Obermeier M, Yang Z, McIntyre KW, Shuster DJ, Witmer M, Dambach D, Chao S, Mathur A, Chen BC, Barrish JC, Robl JA, Townsend R, Iwanowicz EJ: Acridone-based inhibitors of inosine 5'-monophosphate dehydrogenase: discovery and SAR leading to the identification of N-(2-(6-(4-ethylpiperazin-1-yl)pyridin-3-yl)propan-2-yl)-2- fluoro-9-oxo-9,10-dihydroacridine-3-carboxamide (BMS-566419). J Med Chem. 2007 Jul 26;50(15):3730-42. Epub 2007 Jun 22. [17585753 ]
- Nelson PH, Carr SF, Devens BH, Eugui EM, Franco F, Gonzalez C, Hawley RC, Loughhead DG, Milan DJ, Papp E, Patterson JW, Rouhafza S, Sjogren EB, Smith DB, Stephenson RA, Talamas FX, Waltos AM, Weikert RJ, Wu JC: Structure-activity relationships for inhibition of inosine monophosphate dehydrogenase by nuclear variants of mycophenolic acid. J Med Chem. 1996 Oct 11;39(21):4181-96. [8863796 ]
- Chen Z, Zheng Z, Huang H, Song Y, Zhang X, Ma J, Wang B, Zhang C, Ju J: Penicacids A-C, three new mycophenolic acid derivatives and immunosuppressive activities from the marine-derived fungus Penicillium sp. SOF07. Bioorg Med Chem Lett. 2012 May 1;22(9):3332-5. doi: 10.1016/j.bmcl.2012.02.106. Epub 2012 Mar 11. [22464133 ]
- Yang N, Wang QH, Wang WQ, Wang J, Li F, Tan SP, Cheng MS: The design, synthesis and in vitro immunosuppressive evaluation of novel isobenzofuran derivatives. Bioorg Med Chem Lett. 2012 Jan 1;22(1):53-6. doi: 10.1016/j.bmcl.2011.11.078. Epub 2011 Nov 28. [22172700 ]
- Lesiak K, Watanabe KA, Majumdar A, Powell J, Seidman M, Vanderveen K, Goldstein BM, Pankiewicz KW: Synthesis of a methylenebis(phosphonate) analogue of mycophenolic adenine dinucleotide: a glucuronidation-resistant MAD analogue of NAD. J Med Chem. 1998 Feb 12;41(4):618-22. [9484510 ]
- Chen L, Gao G, Felczak K, Bonnac L, Patterson SE, Wilson D, Bennett EM, Jayaram HN, Hedstrom L, Pankiewicz KW: Probing binding requirements of type I and type II isoforms of inosine monophosphate dehydrogenase with adenine-modified nicotinamide adenine dinucleotide analogues. J Med Chem. 2007 Nov 15;50(23):5743-51. Epub 2007 Oct 24. [17958343 ]
- Felczak K, Chen L, Wilson D, Williams J, Vince R, Petrelli R, Jayaram HN, Kusumanchi P, Kumar M, Pankiewicz KW: Cofactor-type inhibitors of inosine monophosphate dehydrogenase via modular approach: targeting the pyrophosphate binding sub-domain. Bioorg Med Chem. 2011 Mar 1;19(5):1594-605. doi: 10.1016/j.bmc.2011.01.042. Epub 2011 Jan 27. [21324702 ]
- Pankiewicz KW, Lesiak-Watanabe KB, Watanabe KA, Patterson SE, Jayaram HN, Yalowitz JA, Miller MD, Seidman M, Majumdar A, Prehna G, Goldstein BM: Novel mycophenolic adenine bis(phosphonate) analogues as potential differentiation agents against human leukemia. J Med Chem. 2002 Jan 31;45(3):703-12. [11806722 ]
- Chen L, Wilson D, Jayaram HN, Pankiewicz KW: Dual inhibitors of inosine monophosphate dehydrogenase and histone deacetylases for cancer treatment. J Med Chem. 2007 Dec 27;50(26):6685-91. Epub 2007 Nov 27. [18038969 ]
- Chen L, Petrelli R, Olesiak M, Wilson DJ, Labello NP, Pankiewicz KW: Bis(sulfonamide) isosters of mycophenolic adenine dinucleotide analogues: inhibition of inosine monophosphate dehydrogenase. Bioorg Med Chem. 2008 Aug 1;16(15):7462-9. doi: 10.1016/j.bmc.2008.06.003. Epub 2008 Jun 10. [18583139 ]
- Chen L, Wilson DJ, Labello NP, Jayaram HN, Pankiewicz KW: Mycophenolic acid analogs with a modified metabolic profile. Bioorg Med Chem. 2008 Oct 15;16(20):9340-5. doi: 10.1016/j.bmc.2008.08.062. Epub 2008 Aug 29. [18809333 ]
- General Function:
- Rna binding
- Specific Function:
- Catalyzes the conversion of inosine 5'-phosphate (IMP) to xanthosine 5'-phosphate (XMP), the first committed and rate-limiting step in the de novo synthesis of guanine nucleotides, and therefore plays an important role in the regulation of cell growth. Could also have a single-stranded nucleic acid-binding activity and could play a role in RNA and/or DNA metabolism. It may also have a role in the development of malignancy and the growth progression of some tumors.
- Gene Name:
- IMPDH1
- Uniprot ID:
- P20839
- Molecular Weight:
- 55405.365 Da
Binding/Activity Constants
| Type | Value | Assay Type | Assay Source |
|---|---|---|---|
| Inhibitory | 0.033 uM | Not Available | BindingDB 19264 |
| Inhibitory | 0.04 uM | Not Available | BindingDB 19264 |
| IC50 | 0.019 uM | Not Available | BindingDB 19264 |
| IC50 | 0.02 uM | Not Available | BindingDB 19264 |
| IC50 | 0.032 uM | Not Available | BindingDB 19264 |
| IC50 | 0.055 uM | Not Available | BindingDB 19264 |
References
- Dzidic A, Prgomet C, Mohr A, Meyer K, Bauer J, Meyer HH, Pfaffl MW: Effects of mycophenolic acid on inosine monophosphate dehydrogenase I and II mRNA expression in white blood cells and various tissues in sheep. J Vet Med A Physiol Pathol Clin Med. 2006 May;53(4):163-9. [16629948 ]
- Mitsuhashi S, Takenaka J, Iwamori K, Nakajima N, Ubukata M: Structure-activity relationships for inhibition of inosine monophosphate dehydrogenase and differentiation induction of K562 cells among the mycophenolic acid derivatives. Bioorg Med Chem. 2010 Nov 15;18(22):8106-11. doi: 10.1016/j.bmc.2010.09.004. Epub 2010 Sep 18. [20934342 ]
- Nelson PH, Eugui E, Wang CC, Allison AC: Synthesis and immunosuppressive activity of some side-chain variants of mycophenolic acid. J Med Chem. 1990 Feb;33(2):833-8. [1967654 ]
- Watkins WJ, Chen JM, Cho A, Chong L, Collins N, Fardis M, Huang W, Hung M, Kirschberg T, Lee WA, Liu X, Thomas W, Xu J, Zeynalzadegan A, Zhang J: Phosphonic acid-containing analogues of mycophenolic acid as inhibitors of IMPDH. Bioorg Med Chem Lett. 2006 Jul 1;16(13):3479-83. Epub 2006 Apr 18. [16621550 ]
- Watterson SH, Carlsen M, Dhar TG, Shen Z, Pitts WJ, Guo J, Gu HH, Norris D, Chorba J, Chen P, Cheney D, Witmer M, Fleener CA, Rouleau K, Townsend R, Hollenbaugh DL, Iwanowicz EJ: Novel inhibitors of IMPDH: a highly potent and selective quinolone-based series. Bioorg Med Chem Lett. 2003 Feb 10;13(3):543-6. [12565968 ]
- Lesiak K, Watanabe KA, Majumdar A, Powell J, Seidman M, Vanderveen K, Goldstein BM, Pankiewicz KW: Synthesis of a methylenebis(phosphonate) analogue of mycophenolic adenine dinucleotide: a glucuronidation-resistant MAD analogue of NAD. J Med Chem. 1998 Feb 12;41(4):618-22. [9484510 ]
- Chen L, Gao G, Felczak K, Bonnac L, Patterson SE, Wilson D, Bennett EM, Jayaram HN, Hedstrom L, Pankiewicz KW: Probing binding requirements of type I and type II isoforms of inosine monophosphate dehydrogenase with adenine-modified nicotinamide adenine dinucleotide analogues. J Med Chem. 2007 Nov 15;50(23):5743-51. Epub 2007 Oct 24. [17958343 ]
- Felczak K, Chen L, Wilson D, Williams J, Vince R, Petrelli R, Jayaram HN, Kusumanchi P, Kumar M, Pankiewicz KW: Cofactor-type inhibitors of inosine monophosphate dehydrogenase via modular approach: targeting the pyrophosphate binding sub-domain. Bioorg Med Chem. 2011 Mar 1;19(5):1594-605. doi: 10.1016/j.bmc.2011.01.042. Epub 2011 Jan 27. [21324702 ]
- Pankiewicz KW, Lesiak-Watanabe KB, Watanabe KA, Patterson SE, Jayaram HN, Yalowitz JA, Miller MD, Seidman M, Majumdar A, Prehna G, Goldstein BM: Novel mycophenolic adenine bis(phosphonate) analogues as potential differentiation agents against human leukemia. J Med Chem. 2002 Jan 31;45(3):703-12. [11806722 ]
- Chen L, Wilson D, Jayaram HN, Pankiewicz KW: Dual inhibitors of inosine monophosphate dehydrogenase and histone deacetylases for cancer treatment. J Med Chem. 2007 Dec 27;50(26):6685-91. Epub 2007 Nov 27. [18038969 ]
- Chen L, Petrelli R, Olesiak M, Wilson DJ, Labello NP, Pankiewicz KW: Bis(sulfonamide) isosters of mycophenolic adenine dinucleotide analogues: inhibition of inosine monophosphate dehydrogenase. Bioorg Med Chem. 2008 Aug 1;16(15):7462-9. doi: 10.1016/j.bmc.2008.06.003. Epub 2008 Jun 10. [18583139 ]
- Chen L, Wilson DJ, Labello NP, Jayaram HN, Pankiewicz KW: Mycophenolic acid analogs with a modified metabolic profile. Bioorg Med Chem. 2008 Oct 15;16(20):9340-5. doi: 10.1016/j.bmc.2008.08.062. Epub 2008 Aug 29. [18809333 ]
- General Function:
- Rna binding
- Specific Function:
- Catalyzes the conversion of inosine 5'-phosphate (IMP) to xanthosine 5'-phosphate (XMP), the first committed and rate-limiting step in the de novo synthesis of guanine nucleotides, and therefore plays an important role in the regulation of cell growth. Could also have a single-stranded nucleic acid-binding activity and could play a role in RNA and/or DNA metabolism. It may also have a role in the development of malignancy and the growth progression of some tumors.
- Gene Name:
- IMPDH2
- Uniprot ID:
- P12268
- Molecular Weight:
- 55804.495 Da
References
- Wishart DS, Knox C, Guo AC, Cheng D, Shrivastava S, Tzur D, Gautam B, Hassanali M: DrugBank: a knowledgebase for drugs, drug actions and drug targets. Nucleic Acids Res. 2008 Jan;36(Database issue):D901-6. Epub 2007 Nov 29. [18048412 ]
- General Function:
- Zinc ion binding
- Specific Function:
- Nuclear receptor that binds peroxisome proliferators such as hypolipidemic drugs and fatty acids. Once activated by a ligand, the nuclear receptor binds to DNA specific PPAR response elements (PPRE) and modulates the transcription of its target genes, such as acyl-CoA oxidase. It therefore controls the peroxisomal beta-oxidation pathway of fatty acids. Key regulator of adipocyte differentiation and glucose homeostasis. ARF6 acts as a key regulator of the tissue-specific adipocyte P2 (aP2) enhancer. Acts as a critical regulator of gut homeostasis by suppressing NF-kappa-B-mediated proinflammatory responses. Plays a role in the regulation of cardiovascular circadian rhythms by regulating the transcription of ARNTL/BMAL1 in the blood vessels (By similarity).
- Gene Name:
- PPARG
- Uniprot ID:
- P37231
- Molecular Weight:
- 57619.58 Da
Binding/Activity Constants
| Type | Value | Assay Type | Assay Source |
|---|---|---|---|
| Dissociation | 110 uM | Not Available | BindingDB 19264 |
References
- Ubukata M, Takamori H, Ohashi M, Mitsuhashi S, Yamashita K, Asada T, Nakajima N, Matsuura N, Tsuruga M, Taki K, Magae J: Mycophenolic acid as a latent agonist of PPARgamma. Bioorg Med Chem Lett. 2007 Sep 1;17(17):4767-70. Epub 2007 Jun 26. [17618115 ]
- General Function:
- Rna binding
- Specific Function:
- Catalyzes the conversion of inosine 5'-phosphate (IMP) to xanthosine 5'-phosphate (XMP), the first committed and rate-limiting step in the de novo synthesis of guanine nucleotides, and therefore plays an important role in the regulation of cell growth. Could also have a single-stranded nucleic acid-binding activity and could play a role in RNA and/or DNA metabolism. It may also have a role in the development of malignancy and the growth progression of some tumors.
- Gene Name:
- IMPDH2
- Uniprot ID:
- P12268
- Molecular Weight:
- 55804.495 Da