| Record Information |
|---|
| Version | 2.0 |
|---|
| Creation Date | 2014-08-12 17:09:46 UTC |
|---|
| Update Date | 2014-12-24 20:26:34 UTC |
|---|
| Accession Number | T3D3945 |
|---|
| Identification |
|---|
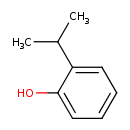
| Common Name | 2-Isopropylphenol |
|---|
| Class | Small Molecule |
|---|
| Description | 2-Isopropylphenol is a flavouring ingredient
2-isopropylphenol belongs to the family of Cumenes. These are aromatic compounds containing a prop-2-ylbenzene moiety. |
|---|
| Compound Type | - Flavouring Agent
- Food Toxin
- Industrial/Workplace Toxin
- Metabolite
- Organic Compound
- Synthetic Compound
|
|---|
| Chemical Structure | |
|---|
| Synonyms | | Synonym | | 1-Hydroxy-2-isopropylbenzene | | 1-Hydroxy-3-isopropylbenzene | | 2-(1-Methylethyl)-Phenol | | 2-(1-Methylethyl)phenol | | 2-(1-Methylethyl)phenol, 9CI | | 2-(Propan-2-yl)phenol | | FEMA 3461 | | Isopropyl-Phenol | | Isopropylphenol, ortho | | O-Cumenol | | O-Hydroxycumene | | O-Isopropyl-Phenol | | O-Isopropylphenol | | Ortho-isopropylphenol | | Prodox 131 |
|
|---|
| Chemical Formula | C9H12O |
|---|
| Average Molecular Mass | 136.191 g/mol |
|---|
| Monoisotopic Mass | 136.089 g/mol |
|---|
| CAS Registry Number | 88-69-7 |
|---|
| IUPAC Name | 2-(propan-2-yl)phenol |
|---|
| Traditional Name | 2-isopropylphenol |
|---|
| SMILES | CC(C)C1=CC=CC=C1O |
|---|
| InChI Identifier | InChI=1S/C9H12O/c1-7(2)8-5-3-4-6-9(8)10/h3-7,10H,1-2H3 |
|---|
| InChI Key | InChIKey=CRBJBYGJVIBWIY-UHFFFAOYSA-N |
|---|
| Chemical Taxonomy |
|---|
| Description | belongs to the class of organic compounds known as cumenes. These are aromatic compounds containing a prop-2-ylbenzene moiety. |
|---|
| Kingdom | Organic compounds |
|---|
| Super Class | Benzenoids |
|---|
| Class | Benzene and substituted derivatives |
|---|
| Sub Class | Cumenes |
|---|
| Direct Parent | Cumenes |
|---|
| Alternative Parents | |
|---|
| Substituents | - Phenylpropane
- Cumene
- 1-hydroxy-4-unsubstituted benzenoid
- 1-hydroxy-2-unsubstituted benzenoid
- Phenol
- Organic oxygen compound
- Hydrocarbon derivative
- Organooxygen compound
- Aromatic homomonocyclic compound
|
|---|
| Molecular Framework | Aromatic homomonocyclic compounds |
|---|
| External Descriptors | |
|---|
| Biological Properties |
|---|
| Status | Detected and Not Quantified |
|---|
| Origin | Exogenous |
|---|
| Cellular Locations | |
|---|
| Biofluid Locations | Not Available |
|---|
| Tissue Locations | Not Available |
|---|
| Pathways | Not Available |
|---|
| Applications | Not Available |
|---|
| Biological Roles | Not Available |
|---|
| Chemical Roles | Not Available |
|---|
| Physical Properties |
|---|
| State | Liquid |
|---|
| Appearance | Not Available |
|---|
| Experimental Properties | | Property | Value |
|---|
| Melting Point | 15 - 16°C | | Boiling Point | Not Available | | Solubility | Insoluble | | LogP | 2.88 |
|
|---|
| Predicted Properties | |
|---|
| Spectra |
|---|
| Spectra | | Spectrum Type | Description | Splash Key | Deposition Date | View |
|---|
| GC-MS | GC-MS Spectrum - EI-B (Non-derivatized) | splash10-00dr-9700000000-0a3ec6f0b6db49b1bb8c | 2017-09-12 | View Spectrum | | GC-MS | GC-MS Spectrum - EI-B (Non-derivatized) | splash10-00dr-9700000000-0a3ec6f0b6db49b1bb8c | 2018-05-18 | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | splash10-00dr-6900000000-65b25162f6cb1b887f17 | 2017-09-01 | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (1 TMS) - 70eV, Positive | splash10-006x-8900000000-6f77c31ece141c6bd323 | 2017-10-06 | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | Not Available | 2021-10-12 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-000i-0900000000-f27e4c86fd02f3a4d9f5 | 2016-08-03 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-000i-7900000000-b30b63f3decaee02aa55 | 2016-08-03 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-0uxu-9400000000-6296af1e3299f7217b19 | 2016-08-03 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-000i-0900000000-69d37dcc656ad26252a0 | 2016-08-03 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-000i-0900000000-36ddf0e1e27114cf431a | 2016-08-03 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-00ku-7900000000-512c84878dbff828e6bf | 2016-08-03 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-000i-0900000000-3d153ee27562ca42735c | 2021-09-22 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-0019-0900000000-197285ccea74149669c9 | 2021-09-22 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-014l-9100000000-8b244056e4ef8843629a | 2021-09-22 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-0002-9300000000-3a595b114a7498fb8470 | 2021-09-23 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-00ku-6900000000-4dd9eeb2be33c550227c | 2021-09-23 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-00mo-9000000000-66881aa4860995f95bad | 2021-09-23 | View Spectrum | | MS | Mass Spectrum (Electron Ionization) | splash10-00di-4900000000-0ab562a07073aa1cb86a | 2014-09-20 | View Spectrum | | 1D NMR | 1H NMR Spectrum (1D, 400 MHz, CDCl3, experimental) | Not Available | 2014-09-20 | View Spectrum | | 1D NMR | 13C NMR Spectrum (1D, 25.16 MHz, CDCl3, experimental) | Not Available | 2014-09-23 | View Spectrum | | 1D NMR | 13C NMR Spectrum (1D, 100 MHz, D2O, predicted) | Not Available | 2021-09-25 | View Spectrum | | 1D NMR | 1H NMR Spectrum (1D, 100 MHz, D2O, predicted) | Not Available | 2021-09-25 | View Spectrum | | 1D NMR | 13C NMR Spectrum (1D, 1000 MHz, D2O, predicted) | Not Available | 2021-09-25 | View Spectrum | | 1D NMR | 1H NMR Spectrum (1D, 1000 MHz, D2O, predicted) | Not Available | 2021-09-25 | View Spectrum | | 1D NMR | 13C NMR Spectrum (1D, 200 MHz, D2O, predicted) | Not Available | 2021-09-25 | View Spectrum | | 1D NMR | 1H NMR Spectrum (1D, 200 MHz, D2O, predicted) | Not Available | 2021-09-25 | View Spectrum | | 1D NMR | 13C NMR Spectrum (1D, 300 MHz, D2O, predicted) | Not Available | 2021-09-25 | View Spectrum | | 1D NMR | 1H NMR Spectrum (1D, 300 MHz, D2O, predicted) | Not Available | 2021-09-25 | View Spectrum | | 1D NMR | 13C NMR Spectrum (1D, 400 MHz, D2O, predicted) | Not Available | 2021-09-25 | View Spectrum | | 1D NMR | 1H NMR Spectrum (1D, 400 MHz, D2O, predicted) | Not Available | 2021-09-25 | View Spectrum | | 1D NMR | 13C NMR Spectrum (1D, 500 MHz, D2O, predicted) | Not Available | 2021-09-25 | View Spectrum | | 1D NMR | 1H NMR Spectrum (1D, 500 MHz, D2O, predicted) | Not Available | 2021-09-25 | View Spectrum | | 1D NMR | 13C NMR Spectrum (1D, 600 MHz, D2O, predicted) | Not Available | 2021-09-25 | View Spectrum | | 1D NMR | 1H NMR Spectrum (1D, 600 MHz, D2O, predicted) | Not Available | 2021-09-25 | View Spectrum | | 1D NMR | 13C NMR Spectrum (1D, 700 MHz, D2O, predicted) | Not Available | 2021-09-25 | View Spectrum | | 1D NMR | 1H NMR Spectrum (1D, 700 MHz, D2O, predicted) | Not Available | 2021-09-25 | View Spectrum | | 1D NMR | 13C NMR Spectrum (1D, 800 MHz, D2O, predicted) | Not Available | 2021-09-25 | View Spectrum | | 1D NMR | 1H NMR Spectrum (1D, 800 MHz, D2O, predicted) | Not Available | 2021-09-25 | View Spectrum | | 1D NMR | 13C NMR Spectrum (1D, 900 MHz, D2O, predicted) | Not Available | 2021-09-25 | View Spectrum | | 1D NMR | 1H NMR Spectrum (1D, 900 MHz, D2O, predicted) | Not Available | 2021-09-25 | View Spectrum |
|
|---|
| Toxicity Profile |
|---|
| Route of Exposure | Not Available |
|---|
| Mechanism of Toxicity | Not Available |
|---|
| Metabolism | Not Available |
|---|
| Toxicity Values | Not Available |
|---|
| Lethal Dose | Not Available |
|---|
| Carcinogenicity (IARC Classification) | No indication of carcinogenicity to humans (not listed by IARC). |
|---|
| Uses/Sources | Not Available |
|---|
| Minimum Risk Level | Not Available |
|---|
| Health Effects | Not Available |
|---|
| Symptoms | Not Available |
|---|
| Treatment | Not Available |
|---|
| Normal Concentrations |
|---|
| Not Available |
|---|
| Abnormal Concentrations |
|---|
| Not Available |
|---|
| External Links |
|---|
| DrugBank ID | Not Available |
|---|
| HMDB ID | HMDB32029 |
|---|
| PubChem Compound ID | 6943 |
|---|
| ChEMBL ID | CHEMBL30018 |
|---|
| ChemSpider ID | 6677 |
|---|
| KEGG ID | Not Available |
|---|
| UniProt ID | Not Available |
|---|
| OMIM ID | |
|---|
| ChEBI ID | 38506 |
|---|
| BioCyc ID | Not Available |
|---|
| CTD ID | Not Available |
|---|
| Stitch ID | Not Available |
|---|
| PDB ID | IP0 |
|---|
| ACToR ID | Not Available |
|---|
| Wikipedia Link | Not Available |
|---|
| References |
|---|
| Synthesis Reference | Not Available |
|---|
| MSDS | T3D3945.pdf |
|---|
| General References | - Chiha M, Merouani S, Hamdaoui O, Baup S, Gondrexon N, Petrier C: Modeling of ultrasonic degradation of non-volatile organic compounds by Langmuir-type kinetics. Ultrason Sonochem. 2010 Jun;17(5):773-82. doi: 10.1016/j.ultsonch.2010.03.007. Epub 2010 Mar 27. [20388590 ]
- Toyama T, Momotani N, Ogata Y, Miyamori Y, Inoue D, Sei K, Mori K, Kikuchi S, Ike M: Isolation and characterization of 4-tert-butylphenol-utilizing Sphingobium fuliginis strains from Phragmites australis rhizosphere sediment. Appl Environ Microbiol. 2010 Oct;76(20):6733-40. doi: 10.1128/AEM.00258-10. Epub 2010 Aug 27. [20802076 ]
- Li J, Ma M, Wang Z: In vitro profiling of endocrine disrupting effects of phenols. Toxicol In Vitro. 2010 Feb;24(1):201-7. doi: 10.1016/j.tiv.2009.09.008. Epub 2009 Sep 16. [19765641 ]
- Toyama T, Maeda N, Murashita M, Chang YC, Kikuchi S: Isolation and characterization of a novel 2-sec-butylphenol-degrading bacterium Pseudomonas sp. strain MS-1. Biodegradation. 2010 Apr;21(2):157-65. doi: 10.1007/s10532-009-9290-y. Epub 2009 Aug 25. [19705287 ]
- Cho S, Choi Y, Park S, Park T: Carvacrol prevents diet-induced obesity by modulating gene expressions involved in adipogenesis and inflammation in mice fed with high-fat diet. J Nutr Biochem. 2012 Feb;23(2):192-201. doi: 10.1016/j.jnutbio.2010.11.016. Epub 2011 Mar 29. [21447440 ]
- Glenn GM, Klamczynski AP, Woods DF, Chiou B, Orts WJ, Imam SH: Encapsulation of plant oils in porous starch microspheres. J Agric Food Chem. 2010 Apr 14;58(7):4180-4. doi: 10.1021/jf9037826. [20196603 ]
- Shiizaki K, Asai S, Ebata S, Kawanishi M, Yagi T: Establishment of yeast reporter assay systems to detect ligands of thyroid hormone receptors alpha and beta. Toxicol In Vitro. 2010 Mar;24(2):638-44. doi: 10.1016/j.tiv.2009.10.001. Epub 2009 Oct 22. [19853653 ]
- Krcmar S: Responses of Tabanidae (Diptera) to canopy traps baited with 4-methylphenol, 3-isopropylphenol, and naphthalene. J Vector Ecol. 2007 Dec;32(2):188-92. [18260506 ]
- Harvey KA, Xu Z, Whitley P, Davisson VJ, Siddiqui RA: Characterization of anticancer properties of 2,6-diisopropylphenol-docosahexaenoate and analogues in breast cancer cells. Bioorg Med Chem. 2010 Mar 1;18(5):1866-74. doi: 10.1016/j.bmc.2010.01.045. Epub 2010 Jan 25. [20153203 ]
- Tsuchiya H, Ueno T, Tanaka T, Matsuura N, Mizogami M: Comparative study on determination of antioxidant and membrane activities of propofol and its related compounds. Eur J Pharm Sci. 2010 Jan 31;39(1-3):97-102. doi: 10.1016/j.ejps.2009.11.001. Epub 2009 Nov 6. [19897032 ]
- Leon I, Cocinero EJ, Millan J, Rijs AM, Usabiaga I, Lesarri A, Castano F, Fernandez JA: A combined spectroscopic and theoretical study of propofol.(H2O)3. J Chem Phys. 2012 Aug 21;137(7):074303. [22920116 ]
- Li J, Ma M, Wang Z: A two-hybrid yeast assay to quantify the effects of xenobiotics on retinoid X receptor-mediated gene expression. Toxicol Lett. 2008 Feb 15;176(3):198-206. doi: 10.1016/j.toxlet.2007.11.006. Epub 2007 Dec 3. [18207673 ]
- Feng Y, Colosi LM, Gao S, Huang Q, Mao L: Transformation and removal of tetrabromobisphenol A from water in the presence of natural organic matter via laccase-catalyzed reactions: reaction rates, products, and pathways. Environ Sci Technol. 2013 Jan 15;47(2):1001-8. doi: 10.1021/es302680c. Epub 2013 Jan 7. [23256593 ]
- Li J, Ma M, Wang Z: A two-hybrid yeast assay to quantify the effects of xenobiotics on thyroid hormone-mediated gene expression. Environ Toxicol Chem. 2008 Jan;27(1):159-67. [18092857 ]
- Alessio RJ, Li X, Martin DF: Removal of BPA model compounds and related substances by means of column chromatography using Octolig(R). J Environ Sci Health A Tox Hazard Subst Environ Eng. 2012;47(14):2198-204. doi: 10.1080/10934529.2012.707535. [22934990 ]
- Novak M, Brinster AM, Dickhoff JN, Erb JM, Jones MP, Leopold SH, Vollman AT, Wang YT, Glover SA: Chemistry of 4-alkylaryloxenium ion "precursors": sound and fury signifying something? J Org Chem. 2007 Dec 21;72(26):9954-62. Epub 2007 Nov 21. [18027966 ]
- Yannai, Shmuel. (2004) Dictionary of food compounds with CD-ROM: Additives, flavors, and ingredients. Boca Raton: Chapman & Hall/CRC.
|
|---|
| Gene Regulation |
|---|
| Up-Regulated Genes | Not Available |
|---|
| Down-Regulated Genes | Not Available |
|---|