| Record Information |
|---|
| Version | 2.0 |
|---|
| Creation Date | 2014-09-11 02:06:17 UTC |
|---|
| Update Date | 2014-12-24 20:26:56 UTC |
|---|
| Accession Number | T3D4718 |
|---|
| Identification |
|---|
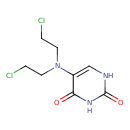
| Common Name | Uracil mustard |
|---|
| Class | Small Molecule |
|---|
| Description | Uracil mustard is only found in individuals that have used or taken this drug. It is a nitrogen mustard derivative of uracil. It is a alkylating antineoplastic agent that is used in lymphatic malignancies, and causes mainly gastrointestinal and bone marrow damage. After activation, uracil mustard binds preferentially to the guanine and cytosine moieties of DNA, leading to cross-linking of DNA, thus inhibiting DNA synthesis and function. |
|---|
| Compound Type | - Amide
- Amine
- Drug
- Metabolite
- Organic Compound
- Organochloride
- Synthetic Compound
|
|---|
| Chemical Structure | |
|---|
| Synonyms | | Synonym | | 5-(Di-2-chloroethyl)aminouracil | | 5-Aminouracil mustard | | 5-N,N-Bis(2-chloroethyl)aminouracil | | 5-[Bis(2-chloroethyl)amino]-2,4(1H,3H)-pyrimidinedione | | 5-[Di(beta-chloroethyl)amino]uracil | | Aminouracil mustard | | Uracil nitrogen mustard |
|
|---|
| Chemical Formula | C8H11Cl2N3O2 |
|---|
| Average Molecular Mass | 252.098 g/mol |
|---|
| Monoisotopic Mass | 251.023 g/mol |
|---|
| CAS Registry Number | 66-75-1 |
|---|
| IUPAC Name | 5-[bis(2-chloroethyl)amino]-1,2,3,4-tetrahydropyrimidine-2,4-dione |
|---|
| Traditional Name | uracil mustard |
|---|
| SMILES | OC1=NC=C(N(CCCl)CCCl)C(O)=N1 |
|---|
| InChI Identifier | InChI=1S/C8H11Cl2N3O2/c9-1-3-13(4-2-10)6-5-11-8(15)12-7(6)14/h5H,1-4H2,(H2,11,12,14,15) |
|---|
| InChI Key | InChIKey=IDPUKCWIGUEADI-UHFFFAOYSA-N |
|---|
| Chemical Taxonomy |
|---|
| Description | belongs to the class of organic compounds known as nitrogen mustard compounds. Nitrogen mustard compounds are compounds having two beta-haloalkyl groups bound to a nitrogen atom. |
|---|
| Kingdom | Organic compounds |
|---|
| Super Class | Organic nitrogen compounds |
|---|
| Class | Organonitrogen compounds |
|---|
| Sub Class | Nitrogen mustard compounds |
|---|
| Direct Parent | Nitrogen mustard compounds |
|---|
| Alternative Parents | |
|---|
| Substituents | - Nitrogen mustard
- Tertiary aliphatic/aromatic amine
- Dialkylarylamine
- Aminopyrimidine
- Pyrimidone
- Hydropyrimidine
- Pyrimidine
- Heteroaromatic compound
- Vinylogous amide
- Lactam
- Tertiary amine
- Urea
- Organoheterocyclic compound
- Azacycle
- Alkyl halide
- Organooxygen compound
- Organochloride
- Organohalogen compound
- Organic oxygen compound
- Hydrocarbon derivative
- Organic oxide
- Alkyl chloride
- Organopnictogen compound
- Amine
- Aromatic heteromonocyclic compound
|
|---|
| Molecular Framework | Aromatic heteromonocyclic compounds |
|---|
| External Descriptors | |
|---|
| Biological Properties |
|---|
| Status | Detected and Not Quantified |
|---|
| Origin | Exogenous |
|---|
| Cellular Locations | |
|---|
| Biofluid Locations | Not Available |
|---|
| Tissue Locations | Not Available |
|---|
| Pathways | Not Available |
|---|
| Applications | Not Available |
|---|
| Biological Roles | Not Available |
|---|
| Chemical Roles | Not Available |
|---|
| Physical Properties |
|---|
| State | Solid |
|---|
| Appearance | White powder. |
|---|
| Experimental Properties | | Property | Value |
|---|
| Melting Point | 206°C | | Boiling Point | Not Available | | Solubility | 1070 mg/L | | LogP | 1.2 |
|
|---|
| Predicted Properties | |
|---|
| Spectra |
|---|
| Spectra | | Spectrum Type | Description | Splash Key | Deposition Date | View |
|---|
| Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | splash10-0frf-1930000000-b082a72f43cd6ef8d865 | 2017-09-01 | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | Not Available | 2021-10-12 | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | Not Available | 2021-10-12 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-0udi-0190000000-ee3938fb1cac655b6750 | 2016-08-03 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-0r00-3690000000-e3d7d6c32273a27ea452 | 2016-08-03 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-02t9-4900000000-c141e19ee451b1467b29 | 2016-08-03 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-0k96-6290000000-93d545f0c9fde44350d4 | 2016-08-03 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-0006-9220000000-ee1cf7bef8946f7dc9fe | 2016-08-03 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-0006-9400000000-5577de19673152e348b9 | 2016-08-03 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-0udi-0090000000-5f08b1b5a345ae125a3f | 2021-10-11 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-0udi-0490000000-86e8889e895d31e6b5a4 | 2021-10-11 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-0w4i-3900000000-73fd2207850235223106 | 2021-10-11 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-001i-9000000000-c2fa753da65a4bac80a1 | 2021-10-11 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-001i-9310000000-abaab02ed488d29cc339 | 2021-10-11 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-001i-9500000000-fffdc9aec429e5149380 | 2021-10-11 | View Spectrum | | MS | Mass Spectrum (Electron Ionization) | splash10-0w29-9730000000-047cc283312feb25a4ed | 2014-09-20 | View Spectrum |
|
|---|
| Toxicity Profile |
|---|
| Route of Exposure | Not Available |
|---|
| Mechanism of Toxicity | After activation, it binds preferentially to the guanine and cytosine moieties of DNA, leading to cross-linking of DNA, thus inhibiting DNA synthesis and function. |
|---|
| Metabolism | Not Available |
|---|
| Toxicity Values | Not Available |
|---|
| Lethal Dose | Not Available |
|---|
| Carcinogenicity (IARC Classification) | 2B, possibly carcinogenic to humans. (1) |
|---|
| Uses/Sources | Used for its antineoplastic properties. |
|---|
| Minimum Risk Level | Not Available |
|---|
| Health Effects | Not Available |
|---|
| Symptoms | Not Available |
|---|
| Treatment | Not Available |
|---|
| Normal Concentrations |
|---|
| Not Available |
|---|
| Abnormal Concentrations |
|---|
| Not Available |
|---|
| External Links |
|---|
| DrugBank ID | DB00791 |
|---|
| HMDB ID | HMDB14929 |
|---|
| PubChem Compound ID | 6194 |
|---|
| ChEMBL ID | CHEMBL1488 |
|---|
| ChemSpider ID | 5959 |
|---|
| KEGG ID | C11686 |
|---|
| UniProt ID | Not Available |
|---|
| OMIM ID | |
|---|
| ChEBI ID | 9884 |
|---|
| BioCyc ID | Not Available |
|---|
| CTD ID | Not Available |
|---|
| Stitch ID | Not Available |
|---|
| PDB ID | Not Available |
|---|
| ACToR ID | Not Available |
|---|
| Wikipedia Link | Uracil_mustard |
|---|
| References |
|---|
| Synthesis Reference | Not Available |
|---|
| MSDS | Not Available |
|---|
| General References | - International Agency for Research on Cancer (2014). IARC Monographs on the Evaluation of Carcinogenic Risks to Humans. [Link]
|
|---|
| Gene Regulation |
|---|
| Up-Regulated Genes | Not Available |
|---|
| Down-Regulated Genes | Not Available |
|---|