| Record Information |
|---|
| Version | 2.0 |
|---|
| Creation Date | 2009-07-05 03:00:49 UTC |
|---|
| Update Date | 2014-12-24 20:25:42 UTC |
|---|
| Accession Number | T3D2561 |
|---|
| Identification |
|---|
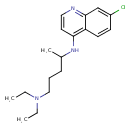
| Common Name | Chloroquine |
|---|
| Class | Small Molecule |
|---|
| Description | Chloroquine is only found in individuals that have used or taken this drug. It is a prototypical antimalarial agent with a mechanism that is not well understood. It has also been used to treat rheumatoid arthritis, systemic lupus erythematosus, and in the systemic therapy of amebic liver abscesses. [PubChem]The mechanism of plasmodicidal action of chloroquine is not completely certain. Like other quinoline derivatives, it is thought to inhibit heme polymerase activity. This results in accumulation of free heme, which is toxic to the parasites. nside red blood cells, the malarial parasite must degrade hemoglobin to acquire essential amino acids, which the parasite requires to construct its own protein and for energy metabolism. Digestion is carried out in a vacuole of the parasite cell.During this process, the parasite produces the toxic and soluble molecule heme. The heme moiety consists of a porphyrin ring called Fe(II)-protoporphyrin IX (FP). To avoid destruction by this molecule, the parasite biocrystallizes heme to form hemozoin, a non-toxic molecule. Hemozoin collects in the digestive vacuole as insoluble crystals.Chloroquine enters the red blood cell, inhabiting parasite cell, and digestive vacuole by simple diffusion. Chloroquine then becomes protonated (to CQ2+), as the digestive vacuole is known to be acidic (pH 4.7); chloroquine then cannot leave by diffusion. Chloroquine caps hemozoin molecules to prevent further biocrystallization of heme, thus leading to heme buildup. Chloroquine binds to heme (or FP) to form what is known as the FP-Chloroquine complex; this complex is highly toxic to the cell and disrupts membrane function. Action of the toxic FP-Chloroquine and FP results in cell lysis and ultimately parasite cell autodigestion. In essence, the parasite cell drowns in its own metabolic products. |
|---|
| Compound Type | - Amebicide
- Amine
- Antimalarial
- Antirheumatic Agent
- Drug
- Metabolite
- Organic Compound
- Organochloride
- Synthetic Compound
|
|---|
| Chemical Structure | |
|---|
| Synonyms | | Synonym | | Aralen | | Artrichin | | Bemaphate | | Capquin | | Chloraquine | | Chlorochin | | Chlorochine | | Chloroquina | | Chloroquinium | | Chloroquinum | | Chlorquin | | Clorochina | | Cloroquina | | Malarex | | N(4)-(7-chloro-4-Quinolinyl)-N(1),N(1)-diethyl-1,4-pentanediamine | | Nivaquine b | | Resochin | | Resoquine | | Reumachlor | | Sanoquin |
|
|---|
| Chemical Formula | C18H26ClN3 |
|---|
| Average Molecular Mass | 319.872 g/mol |
|---|
| Monoisotopic Mass | 319.182 g/mol |
|---|
| CAS Registry Number | 54-05-7 |
|---|
| IUPAC Name | 7-chloro-N-[5-(diethylamino)pentan-2-yl]quinolin-4-amine |
|---|
| Traditional Name | 7-chloro-N-[5-(diethylamino)pentan-2-yl]quinolin-4-amine |
|---|
| SMILES | CCN(CC)CCCC(C)NC1=C2C=CC(Cl)=CC2=NC=C1 |
|---|
| InChI Identifier | InChI=1/C18H26ClN3/c1-4-22(5-2)12-6-7-14(3)21-17-10-11-20-18-13-15(19)8-9-16(17)18/h8-11,13-14H,4-7,12H2,1-3H3,(H,20,21) |
|---|
| InChI Key | InChIKey=WHTVZRBIWZFKQO-UHFFFAOYNA-N |
|---|
| Chemical Taxonomy |
|---|
| Description | belongs to the class of organic compounds known as 4-aminoquinolines. These are organic compounds containing an amino group attached to the 4-position of a quinoline ring system. |
|---|
| Kingdom | Organic compounds |
|---|
| Super Class | Organoheterocyclic compounds |
|---|
| Class | Quinolines and derivatives |
|---|
| Sub Class | Aminoquinolines and derivatives |
|---|
| Direct Parent | 4-aminoquinolines |
|---|
| Alternative Parents | |
|---|
| Substituents | - 4-aminoquinoline
- Haloquinoline
- Chloroquinoline
- Aminopyridine
- Secondary aliphatic/aromatic amine
- Aryl chloride
- Aryl halide
- Pyridine
- Benzenoid
- Heteroaromatic compound
- Tertiary aliphatic amine
- Tertiary amine
- Azacycle
- Secondary amine
- Amine
- Organonitrogen compound
- Organochloride
- Organohalogen compound
- Hydrocarbon derivative
- Organopnictogen compound
- Organic nitrogen compound
- Aromatic heteropolycyclic compound
|
|---|
| Molecular Framework | Aromatic heteropolycyclic compounds |
|---|
| External Descriptors | |
|---|
| Biological Properties |
|---|
| Status | Detected and Not Quantified |
|---|
| Origin | Exogenous |
|---|
| Cellular Locations | - Cytoplasm
- Extracellular
- Membrane
|
|---|
| Biofluid Locations | Not Available |
|---|
| Tissue Locations | Not Available |
|---|
| Pathways | Not Available |
|---|
| Applications | Not Available |
|---|
| Biological Roles | Not Available |
|---|
| Chemical Roles | Not Available |
|---|
| Physical Properties |
|---|
| State | Solid |
|---|
| Appearance | White to slightly yellow, crystalline powder (2). |
|---|
| Experimental Properties | | Property | Value |
|---|
| Melting Point | 289°C | | Boiling Point | Not Available | | Solubility | 10.6 mg/L | | LogP | 4.63 |
|
|---|
| Predicted Properties | |
|---|
| Spectra |
|---|
| Spectra | | Spectrum Type | Description | Splash Key | Deposition Date | View |
|---|
| GC-MS | GC-MS Spectrum - CI-B (Non-derivatized) | splash10-00di-0009000000-d54119d64cfc341cee7d | 2017-09-12 | View Spectrum | | GC-MS | GC-MS Spectrum - CI-B (Non-derivatized) | splash10-00di-0009000000-d54119d64cfc341cee7d | 2018-05-18 | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | splash10-0pbi-9242000000-6cd79ce1c8a9ada4550d | 2017-09-01 | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | Not Available | 2021-10-12 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - 40V, Positive | splash10-002p-8920000000-552012cb889bf65320d8 | 2021-09-20 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - 20V, Positive | splash10-0002-0391000000-7360fd3b6728bd6574b8 | 2021-09-20 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - 10V, Positive | splash10-00di-0029000000-3ea9f3105801d7abf080 | 2021-09-20 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-00di-0119000000-8f9d4513bf5993f3c9c4 | 2016-08-03 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-00dl-3957000000-e533e26da609c9329dae | 2016-08-03 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-00bc-9530000000-646244109f4c47e0c2a6 | 2016-08-03 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-014i-0009000000-d45e56a4f82a2d58657d | 2016-08-03 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-014i-1229000000-09769326d92e55dd78aa | 2016-08-03 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-00fu-9630000000-f2986e805921cc8d3a37 | 2016-08-03 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-00di-0049000000-7e55e664b21e89ddbe44 | 2021-10-11 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-00dj-0198000000-64a0ed0e3e428bf65cbb | 2021-10-11 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-004r-6950000000-75bff8c32e9dcca35dbf | 2021-10-11 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-014i-0009000000-b550e787bd639403cc76 | 2021-10-11 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-014i-0109000000-3494d09f71ff43a3ce73 | 2021-10-11 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-004i-2910000000-efba909b8046e2bf2c67 | 2021-10-11 | View Spectrum | | MS | Mass Spectrum (Electron Ionization) | splash10-000i-9320000000-2663c398ede2e502ca34 | 2014-09-20 | View Spectrum |
|
|---|
| Toxicity Profile |
|---|
| Route of Exposure | Inhalation
Completely absorbed from gastrointestinal tract |
|---|
| Mechanism of Toxicity | The mechanism of plasmodicidal action of chloroquine is not completely certain. Like other quinoline derivatives, it is thought to inhibit heme polymerase activity. This results in accumulation of free heme, which is toxic to the parasites. nside red blood cells, the malarial parasite must degrade hemoglobin to acquire essential amino acids, which the parasite requires to construct its own protein and for energy metabolism. Digestion is carried out in a vacuole of the parasite cell.
During this process, the parasite produces the toxic and soluble molecule heme. The heme moiety consists of a porphyrin ring called Fe(II)-protoporphyrin IX (FP). To avoid destruction by this molecule, the parasite biocrystallizes heme to form hemozoin, a non-toxic molecule. Hemozoin collects in the digestive vacuole as insoluble crystals.
Chloroquine enters the red blood cell, inhabiting parasite cell, and digestive vacuole by simple diffusion. Chloroquine then becomes protonated (to CQ2+), as the digestive vacuole is known to be acidic (pH 4.7); chloroquine then cannot leave by diffusion. Chloroquine caps hemozoin molecules to prevent further biocrystallization of heme, thus leading to heme buildup. Chloroquine binds to heme (or FP) to form what is known as the FP-Chloroquine complex; this complex is highly toxic to the cell and disrupts membrane function. Action of the toxic FP-Chloroquine and FP results in cell lysis and ultimately parasite cell autodigestion. In essence, the parasite cell drowns in its own metabolic products. |
|---|
| Metabolism | Completely absorbed from gastrointestinal tract. Chloroquine is partially metabolized; the major metabolite is desethylchloroquine. Desethylchloroquine also has antiplasmodial activity, but is slightly less active than chloroquine. Bisdesethylchloroquine, which is a carboxylic acid derivative, and several other unidentified metabolites are also formed in small amounts (3).
Route of Elimination: Excretion of chloroquine is quite slow, but is increased by acidification of the urine.
Half Life: 1-2 months |
|---|
| Toxicity Values | Not Available |
|---|
| Lethal Dose | Not Available |
|---|
| Carcinogenicity (IARC Classification) | 3, not classifiable as to its carcinogenicity to humans. (6) |
|---|
| Uses/Sources | For the suppressive treatment and for acute attacks of malaria due to P. vivax, P.malariae, P. ovale, and susceptible strains of P. falciparum, Second-line agent in treatment of Rheumatoid Arthritis (1). |
|---|
| Minimum Risk Level | Not Available |
|---|
| Health Effects | Possible heath effects include an irreversible retinal damage, visual disturbances, nyctalopia; scotomatous vision with field defects of paracentral, pericentral ring types, and typically temporal scotomas, nerve type deafness; tinnitus, reduced hearing in patients with preexisting auditory damage. Other effects include pleomorphic skin eruptions, skeletal muscle myopathy, hypotension, electrocardiographic change as well as neuropsychiatric changes including psychosis, delirium, personality changes and depression (RxList, A308). |
|---|
| Symptoms | Convulsive seizures. Mild and transient headache. |
|---|
| Treatment | Treatment is symptomatic and must be prompt with immediate evacuation of the stomach by emesis (at home, before transportation to the hospital) or gastric lavage until the stomach is completely emptied. If finely powdered, activated charcoal is introduced by stomach tube, after lavage, and within 30 minutes after ingestion of the antimalarial, it may inhibit further intestinal absorption of the drug. To be effective, the dose of activated charcoal should be at least five times the estimated dose of chloroquine ingested. Convulsions, if present, should be controlled before attempting gastric lavage. (8) |
|---|
| Normal Concentrations |
|---|
| Not Available |
|---|
| Abnormal Concentrations |
|---|
| Not Available |
|---|
| External Links |
|---|
| DrugBank ID | DB00608 |
|---|
| HMDB ID | HMDB14746 |
|---|
| PubChem Compound ID | 2719 |
|---|
| ChEMBL ID | CHEMBL76 |
|---|
| ChemSpider ID | 2618 |
|---|
| KEGG ID | C07625 |
|---|
| UniProt ID | Not Available |
|---|
| OMIM ID | |
|---|
| ChEBI ID | 3638 |
|---|
| BioCyc ID | Not Available |
|---|
| CTD ID | D002738 |
|---|
| Stitch ID | Chloroquine |
|---|
| PDB ID | CLQ |
|---|
| ACToR ID | Not Available |
|---|
| Wikipedia Link | Chloroquine |
|---|
| References |
|---|
| Synthesis Reference | Andersag, H., Breitner, S.and Jung, H.; U S . Patent 2,233,970; March 4,1941; assigned to
Winthrop Chemical Company, Inc. |
|---|
| MSDS | Link |
|---|
| General References | - Wishart DS, Knox C, Guo AC, Cheng D, Shrivastava S, Tzur D, Gautam B, Hassanali M: DrugBank: a knowledgebase for drugs, drug actions and drug targets. Nucleic Acids Res. 2008 Jan;36(Database issue):D901-6. Epub 2007 Nov 29. [18048412 ]
- Flanagan JN, Young MV, Persons KS, Wang L, Mathieu JS, Whitlatch LW, Holick MF, Chen TC: Vitamin D metabolism in human prostate cells: implications for prostate cancer chemoprevention by vitamin D. Anticancer Res. 2006 Jul-Aug;26(4A):2567-72. [16886665 ]
- Kojima T, Matsumoto M, Togashi H, Tachibana K, Kemmotsu O, Yoshioka M: Fluvoxamine suppresses the long-term potentiation in the hippocampal CA1 field of anesthetized rats: an effect mediated via 5-HT1A receptors. Brain Res. 2003 Jan 3;959(1):165-8. [12480170 ]
- Osol A, Hoover JE, et al. (eds) (1975). Remington's Pharmaceutical Sciences. 15th ed. Easton, Pennsylvania: Mack Publishing Co.
- McEvoy GK (ed) (2006). American Hospital Formulary Service - Drug Information 2006. Bethesda, MD: American Society of Health-System Pharmacists.
- International Agency for Research on Cancer (2014). IARC Monographs on the Evaluation of Carcinogenic Risks to Humans. [Link]
- Drugs.com [Link]
- RxList: The Internet Drug Index (2009). [Link]
|
|---|
| Gene Regulation |
|---|
| Up-Regulated Genes | | Gene | Gene Symbol | Gene ID | Interaction | Chromosome | Details |
|---|
|
|---|
| Down-Regulated Genes | | Gene | Gene Symbol | Gene ID | Interaction | Chromosome | Details |
|---|
|
|---|