| Record Information |
|---|
| Version | 2.0 |
|---|
| Creation Date | 2009-07-21 20:26:38 UTC |
|---|
| Update Date | 2014-12-24 20:25:51 UTC |
|---|
| Accession Number | T3D2756 |
|---|
| Identification |
|---|
| Common Name | Cetirizine |
|---|
| Class | Small Molecule |
|---|
| Description | Cetirizine is a medication used for the treatment of allergies, hay fever, angioedema, and hives. It is a second-generation H1-receptor antagonist antihistamine and works by blocking H1 histamine receptors. It is a major metabolite of hydroxyzine, and has the same basic side effects, including dry mouth. A potent second-generation histamine H1 antagonist that is effective in the treatment of allergic rhinitis, chronic urticaria, and pollen-induced asthma. Unlike many traditional antihistamines, it does not cause drowsiness or anticholinergic side effects. Cetirizine hydrochloride is a medication used for the treatment of allergies, hay fever, angioedema, and hives. It is a second-generation H1-receptor antagonist antihistamine and works by blocking H1 histamine receptors. It is a major metabolite of hydroxyzine, and has the same basic side effects, including dry mouth. |
|---|
| Compound Type | - Amine
- Anti-Allergic Agent
- Drug
- Ether
- Food Toxin
- Histamine Antagonist
- Histamine H1 Antagonist, Non-Sedating
- Metabolite
- Organic Compound
- Organochloride
- Synthetic Compound
|
|---|
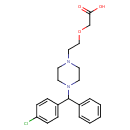
| Chemical Structure | |
|---|
| Synonyms | | Synonym | | Alerlisin | | Benaday | | Cetirizin | | Cetirizina | | Cetirizine hydrochloride | | Cetirizinum | | Cetryn | | Formistin | | Hitrizin | | Humex | | Reactine | | Virlix | | Zirtek | | Zyrlex | | Zyrtec |
|
|---|
| Chemical Formula | C21H25ClN2O3 |
|---|
| Average Molecular Mass | 388.888 g/mol |
|---|
| Monoisotopic Mass | 388.155 g/mol |
|---|
| CAS Registry Number | 83881-51-0 |
|---|
| IUPAC Name | 2-(2-{4-[(4-chlorophenyl)(phenyl)methyl]piperazin-1-yl}ethoxy)acetic acid |
|---|
| Traditional Name | cetirizine |
|---|
| SMILES | OC(=O)COCCN1CCN(CC1)C(C1=CC=CC=C1)C1=CC=C(Cl)C=C1 |
|---|
| InChI Identifier | InChI=1/C21H25ClN2O3/c22-19-8-6-18(7-9-19)21(17-4-2-1-3-5-17)24-12-10-23(11-13-24)14-15-27-16-20(25)26/h1-9,21H,10-16H2,(H,25,26) |
|---|
| InChI Key | InChIKey=ZKLPARSLTMPFCP-UHFFFAOYNA-N |
|---|
| Chemical Taxonomy |
|---|
| Description | belongs to the class of organic compounds known as diphenylmethanes. Diphenylmethanes are compounds containing a diphenylmethane moiety, which consists of a methane wherein two hydrogen atoms are replaced by two phenyl groups. |
|---|
| Kingdom | Organic compounds |
|---|
| Super Class | Benzenoids |
|---|
| Class | Benzene and substituted derivatives |
|---|
| Sub Class | Diphenylmethanes |
|---|
| Direct Parent | Diphenylmethanes |
|---|
| Alternative Parents | |
|---|
| Substituents | - Diphenylmethane
- Chlorobenzene
- Halobenzene
- N-alkylpiperazine
- Aralkylamine
- Aryl halide
- 1,4-diazinane
- Aryl chloride
- Piperazine
- Amino acid
- Amino acid or derivatives
- Tertiary amine
- Tertiary aliphatic amine
- Organoheterocyclic compound
- Carboxylic acid derivative
- Carboxylic acid
- Dialkyl ether
- Monocarboxylic acid or derivatives
- Azacycle
- Ether
- Organic oxygen compound
- Organic nitrogen compound
- Organopnictogen compound
- Organohalogen compound
- Amine
- Organochloride
- Organonitrogen compound
- Carbonyl group
- Hydrocarbon derivative
- Organooxygen compound
- Organic oxide
- Aromatic heteromonocyclic compound
|
|---|
| Molecular Framework | Aromatic heteromonocyclic compounds |
|---|
| External Descriptors | |
|---|
| Biological Properties |
|---|
| Status | Detected and Not Quantified |
|---|
| Origin | Exogenous |
|---|
| Cellular Locations | |
|---|
| Biofluid Locations | Not Available |
|---|
| Tissue Locations | |
|---|
| Pathways | Not Available |
|---|
| Applications | |
|---|
| Biological Roles | |
|---|
| Chemical Roles | Not Available |
|---|
| Physical Properties |
|---|
| State | Solid |
|---|
| Appearance | White powder. |
|---|
| Experimental Properties | | Property | Value |
|---|
| Melting Point | 112.5°C | | Boiling Point | Not Available | | Solubility | 101 mg/L | | LogP | 2.8 |
|
|---|
| Predicted Properties | |
|---|
| Spectra |
|---|
| Spectra | | Spectrum Type | Description | Splash Key | Deposition Date | View |
|---|
| Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | splash10-0udr-3292000000-e20abaab20740ce81a18 | 2017-09-01 | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (1 TMS) - 70eV, Positive | splash10-0uds-7498000000-d6daecf118171395d6b7 | 2017-10-06 | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | Not Available | 2021-10-12 | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | Not Available | 2021-10-12 | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TBDMS_1_1) - 70eV, Positive | Not Available | 2021-11-03 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-ITFT , negative | splash10-000j-0900000000-9801bb859e79fa9922aa | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-ITFT , negative | splash10-000i-0009000000-014b559d56b0f32984ff | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-ITFT , negative | splash10-000i-0009000000-213bd47e17d5ab69ad4b | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-ITFT , negative | splash10-004r-9006000000-f496e22c8e0e293fe98c | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-ITFT , negative | splash10-000i-0009000000-d8a55ab1a7aff0580a91 | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-ITFT , negative | splash10-000i-0009000000-d8a55ab1a7aff0580a91 | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-ITFT , negative | splash10-000i-0009000000-dc957bb6f947575ae4fa | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-ITFT , negative | splash10-000b-0900000000-48fc0581ff9fff15e16d | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-ITFT , negative | splash10-000i-0009000000-04ca95fadd952d0aaeb8 | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-ITFT , negative | splash10-000i-0009000000-014b559d56b0f32984ff | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-ITFT , negative | splash10-0006-0009000000-26a7769df28c2325ec53 | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-ITFT , negative | splash10-0006-0009000000-284c2e7b25733d7c9875 | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-QFT , negative | splash10-000i-0009000000-04befed283d691217ba2 | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-ITFT , positive | splash10-0udi-0190000000-14715ff4d57580fbb166 | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-ITFT , positive | splash10-0f79-0069000000-431df9c3533b77cf08bc | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-ITFT , positive | splash10-0udi-0090000000-e042b67e2048318e7dbd | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-ITFT , positive | splash10-0udi-0190000000-b0dfb67717c01fd0c0b9 | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-ITFT , positive | splash10-0gb9-0970000000-e2810b242fefb4a419e7 | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-ITFT , positive | splash10-014i-0900000000-72f7cddc4ff6d45f98c0 | 2017-09-14 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-000i-0019000000-e2cc239f762011eb0c4d | 2016-08-03 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-0w2i-1169000000-185e6899ae017256a767 | 2016-08-03 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-0udi-3391000000-bfd70499959ae71093eb | 2016-08-03 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-000i-0009000000-caf1604c9603d45a6712 | 2016-08-03 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-000i-1119000000-61c40cd9a0e9b76b5e91 | 2016-08-03 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-056r-9162000000-08731719ee47b215e4c7 | 2016-08-03 | View Spectrum |
|
|---|
| Toxicity Profile |
|---|
| Route of Exposure | Oral; mean peak plasma concentration (Cmax) of 114 ng/mL at a time (Tmax) of 2.2 hours postdose was observed for cetirizine. |
|---|
| Mechanism of Toxicity | Cetirizine competes with histamine for binding at H1-receptor sites on the effector cell surface, resulting in suppression of histaminic edema, flare, and pruritus. The low incidence of sedation can be attributed to reduced penetration of cetirizine into the CNS as a result of the less lipophilic carboxyl group on the ethylamine side chain. |
|---|
| Metabolism |
Half Life: 8.3 hours |
|---|
| Toxicity Values | Not Available |
|---|
| Lethal Dose | Not Available |
|---|
| Carcinogenicity (IARC Classification) | No indication of carcinogenicity to humans (not listed by IARC). |
|---|
| Uses/Sources | For the relief of symptoms associated with seasonal allergic rhinitis, perennial allergic rhinitis and the treatment of the uncomplicated skin manifestations of chronic idiopathic urticaria |
|---|
| Minimum Risk Level | Not Available |
|---|
| Health Effects | Overdosage has been reported with ZYRTEC (cetirizine) . In one adult patient who took 150 mg of ZYRTEC (cetirizine) , the patient was somnolent but did not display any other clinical signs or abnormal blood chemistry or hematology results. In an 18 month old pediatric patient who took an overdose of ZYRTEC (cetirizine) (approximately 180 mg), restlessness and irritability were observed initially; this was followed by drowsiness. (13) |
|---|
| Symptoms | Somnolence (sleepiness or unusual drowsiness), restlessness, irritability. |
|---|
| Treatment | Should overdose occur, treatment should be symptomatic or supportive, taking into account any concomitantly ingested medications. There is no known specific antidote to Cetirizine. (13) |
|---|
| Normal Concentrations |
|---|
| Not Available |
|---|
| Abnormal Concentrations |
|---|
| Not Available |
|---|
| External Links |
|---|
| DrugBank ID | DB00341 |
|---|
| HMDB ID | HMDB05032 |
|---|
| PubChem Compound ID | 2678 |
|---|
| ChEMBL ID | CHEMBL1000 |
|---|
| ChemSpider ID | 2577 |
|---|
| KEGG ID | C07778 |
|---|
| UniProt ID | Not Available |
|---|
| OMIM ID | |
|---|
| ChEBI ID | 3561 |
|---|
| BioCyc ID | Not Available |
|---|
| CTD ID | Not Available |
|---|
| Stitch ID | Cetirizine |
|---|
| PDB ID | Not Available |
|---|
| ACToR ID | Not Available |
|---|
| Wikipedia Link | Cetirizine |
|---|
| References |
|---|
| Synthesis Reference | Manne Reddy, “Polymorphic forms of dihydrochloride salts of cetirizine and processes for preparation thereof.” U.S. Patent US20040186112, issued September 23, 2004. |
|---|
| MSDS | Link |
|---|
| General References | - Purohit A, Melac M, Pauli G, Frossard N: Comparative activity of cetirizine and desloratadine on histamine-induced wheal-and-flare responses during 24 hours. Ann Allergy Asthma Immunol. 2004 Jun;92(6):635-40. [15237765 ]
- Ramboer I, Bumtbacea R, Lazarescu D, Radu JR: Cetirizine and loratadine: a comparison using the ED50 in skin reactions. J Int Med Res. 2000 Mar-Apr;28(2):69-77. [10898119 ]
- Townley RG, Okada C: Use of cetirizine to investigate non-H1 effects of second-generation antihistamines. Ann Allergy. 1992 Feb;68(2):190-6. [1346737 ]
- Brik A, Tashkin DP, Gong H Jr, Dauphinee B, Lee E: Effect of cetirizine, a new histamine H1 antagonist, on airway dynamics and responsiveness to inhaled histamine in mild asthma. J Allergy Clin Immunol. 1987 Jul;80(1):51-6. [2885355 ]
- Fadel R, Herpin-Richard N, Rihoux JP, Henocq E: Inhibitory effect of cetirizine 2HCl on eosinophil migration in vivo. Clin Allergy. 1987 Jul;17(4):373-9. [2887304 ]
- Tashiro M, Sakurada Y, Iwabuchi K, Mochizuki H, Kato M, Aoki M, Funaki Y, Itoh M, Iwata R, Wong DF, Yanai K: Central effects of fexofenadine and cetirizine: measurement of psychomotor performance, subjective sleepiness, and brain histamine H1-receptor occupancy using 11C-doxepin positron emission tomography. J Clin Pharmacol. 2004 Aug;44(8):890-900. [15286093 ]
- Petersen LJ, Church MK, Rihoux JP, Skov PS: Measurement of interstitial cetirizine concentrations in human skin: correlation of drug levels with inhibition of histamine-induced skin responses. Allergy. 1999 Jun;54(6):607-11. [10435475 ]
- Siergiejko Z, Michalska I, Rogalewska A, Chyrek-Borowska S: [Effect of cetirizine, selective H1 antagonist of histamine on skin and bronchial reactivity and cellular histamine release in allergic diseases]. Pneumonol Alergol Pol. 1992;60(11-12):22-7. [1303774 ]
- Purohit A, Duvernelle C, Melac M, Pauli G, Frossard N: Twenty-four hours of activity of cetirizine and fexofenadine in the skin. Ann Allergy Asthma Immunol. 2001 Apr;86(4):387-92. [11345280 ]
- Atkins PC, Zweiman B, Moskovitz A, von Allmen C, Ciliberti M: Cellular inflammatory responses and mediator release during early developing late-phase allergic cutaneous inflammatory responses: effects of cetirizine. J Allergy Clin Immunol. 1997 Jun;99(6 Pt 1):806-11. [9215249 ]
- Simons FE, Murray HE, Simons KJ: Quantitation of H1-receptor antagonists in skin and serum. J Allergy Clin Immunol. 1995 Mar;95(3):759-64. [7897161 ]
- Drugs.com [Link]
- RxList: The Internet Drug Index (2009). [Link]
|
|---|
| Gene Regulation |
|---|
| Up-Regulated Genes | Not Available |
|---|
| Down-Regulated Genes | Not Available |
|---|