| Record Information |
|---|
| Version | 2.0 |
|---|
| Creation Date | 2009-07-30 17:58:54 UTC |
|---|
| Update Date | 2014-12-24 20:26:07 UTC |
|---|
| Accession Number | T3D3507 |
|---|
| Identification |
|---|
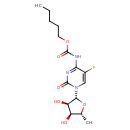
| Common Name | Capecitabine |
|---|
| Class | Small Molecule |
|---|
| Description | Capecitabine is an orally-administered chemotherapeutic agent used in the treatment of metastatic breast and colorectal cancers. Capecitabine is a prodrug, that is enzymatically converted to fluorouracil (antimetabolite) in the tumor, where it inhibits DNA synthesis and slows growth of tumor tissue. |
|---|
| Compound Type | - Amine
- Antimetabolite
- Antimetabolite, Antineoplastic
- Antineoplastic Agent
- Drug
- Ester
- Ether
- Metabolite
- Organic Compound
- Organofluoride
- Prodrug
- Synthetic Compound
|
|---|
| Chemical Structure | |
|---|
| Synonyms | | Synonym | | (1-(5-Deoxy-beta-D-ribofuranosyl)-5-fluoro-1,2-dihydro-2-oxo-4-pyrimidinyl)-carbamic acid pentyl ester | | Capecitabin | | Capecitabina | | Capecitabinum | | Pentyl 1-(5-deoxy-beta-D-ribofuranosyl)-5-fluoro-1,2-dihydro-2-oxo-4-pyrimidinecarbamate | | Pentyl [1-(5-deoxy-beta-D-ribofuranosyl)-5-fluoro-2-oxo-1,2-dihydropyrimidin-4-yl]carbamate | | R340 | | Xeloda |
|
|---|
| Chemical Formula | C15H22FN3O6 |
|---|
| Average Molecular Mass | 359.350 g/mol |
|---|
| Monoisotopic Mass | 359.149 g/mol |
|---|
| CAS Registry Number | 154361-50-9 |
|---|
| IUPAC Name | pentyl N-{1-[(2R,3R,4S,5R)-3,4-dihydroxy-5-methyloxolan-2-yl]-5-fluoro-2-oxo-1,2-dihydropyrimidin-4-yl}carbamate |
|---|
| Traditional Name | capecitabine |
|---|
| SMILES | [H][C@]1(C)O[C@@]([H])(N2C=C(F)C(N=C(O)OCCCCC)=NC2=O)[C@]([H])(O)[C@]1([H])O |
|---|
| InChI Identifier | InChI=1S/C15H22FN3O6/c1-3-4-5-6-24-15(23)18-12-9(16)7-19(14(22)17-12)13-11(21)10(20)8(2)25-13/h7-8,10-11,13,20-21H,3-6H2,1-2H3,(H,17,18,22,23)/t8-,10-,11-,13-/m1/s1 |
|---|
| InChI Key | InChIKey=GAGWJHPBXLXJQN-UORFTKCHSA-N |
|---|
| Chemical Taxonomy |
|---|
| Description | belongs to the class of organic compounds known as 5'-deoxyribonucleosides. These are nucleosides in which the oxygen atom at the 5'position of the ribose moiety has been replaced by another atom. The nucleobases here are limited to purine, pyrimidine, and pyridine derivatives. |
|---|
| Kingdom | Organic compounds |
|---|
| Super Class | Nucleosides, nucleotides, and analogues |
|---|
| Class | 5'-deoxyribonucleosides |
|---|
| Sub Class | Not Available |
|---|
| Direct Parent | 5'-deoxyribonucleosides |
|---|
| Alternative Parents | |
|---|
| Substituents | - 5'-deoxyribonucleoside
- Glycosyl compound
- N-glycosyl compound
- Halopyrimidine
- Pyrimidone
- Aryl fluoride
- Aryl halide
- Pyrimidine
- Hydropyrimidine
- Heteroaromatic compound
- Tetrahydrofuran
- Secondary alcohol
- 1,2-diol
- Oxacycle
- Azacycle
- Organoheterocyclic compound
- Organic 1,3-dipolar compound
- Propargyl-type 1,3-dipolar organic compound
- Carboximidic acid derivative
- Organopnictogen compound
- Organohalogen compound
- Organofluoride
- Organonitrogen compound
- Organic oxygen compound
- Organic nitrogen compound
- Organooxygen compound
- Alcohol
- Hydrocarbon derivative
- Organic oxide
- Aromatic heteromonocyclic compound
|
|---|
| Molecular Framework | Aromatic heteromonocyclic compounds |
|---|
| External Descriptors | |
|---|
| Biological Properties |
|---|
| Status | Detected and Not Quantified |
|---|
| Origin | Exogenous |
|---|
| Cellular Locations | - Cytoplasm
- Extracellular
- Membrane
|
|---|
| Biofluid Locations | Not Available |
|---|
| Tissue Locations | Not Available |
|---|
| Pathways | Not Available |
|---|
| Applications | |
|---|
| Biological Roles | |
|---|
| Chemical Roles | Not Available |
|---|
| Physical Properties |
|---|
| State | Solid |
|---|
| Appearance | White powder. |
|---|
| Experimental Properties | | Property | Value |
|---|
| Melting Point | 110-121°C | | Boiling Point | Not Available | | Solubility | 26 mg/mL | | LogP | 0.4 |
|
|---|
| Predicted Properties | |
|---|
| Spectra |
|---|
| Spectra | | Spectrum Type | Description | Splash Key | Deposition Date | View |
|---|
| Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | splash10-0ab9-9112000000-35a3e944454fc6eca103 | 2017-09-01 | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (2 TMS) - 70eV, Positive | splash10-0079-9502500000-bb4d3c6463fb0a87a997 | 2017-10-06 | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | Not Available | 2021-10-12 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-QFT , negative | splash10-0a4i-0109000000-c0c4de3c4233af3f34d4 | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-QFT , negative | splash10-0udi-0901000000-676f6925c6c255f4dd24 | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-QFT , negative | splash10-0udi-0900000000-236cb917e50b215b9bd1 | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-QFT , negative | splash10-0udi-1900000000-f06b7309e7a2ebfb3bde | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-QFT , negative | splash10-0ufr-3900000000-6bb1981f61b4205a9991 | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-QFT , negative | splash10-0kdi-6900000000-ac9a56401938b72ae4f1 | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-ITFT , negative | splash10-0a4i-0309000000-feac9f5163221adf2a7d | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-ITFT , negative | splash10-0udj-0912000000-258b837e7c98d16aafbf | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-ITFT , negative | splash10-0udi-0900000000-db3e2d6a5526611d6a6e | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-ITFT , negative | splash10-0udi-0900000000-b792e6750d1eaccca7aa | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-QFT , negative | splash10-0pb9-0609000000-cd05885e7f4f8d1853ec | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-QTOF , positive | splash10-0006-0090000000-8029f6bc09ef7062240e | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-QTOF , positive | splash10-006x-0590000000-8cea6a9f156f58ccb593 | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-QTOF , positive | splash10-00e9-0920000000-e942daf2b25f1829bec1 | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-QTOF , positive | splash10-0089-0900000000-2a5dcf8bb6ff7ae6d72f | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-QTOF , positive | splash10-001i-0900000000-460a3e3f1f8b25687c86 | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-QFT , positive | splash10-0006-0090000000-6105bf5c619fbe72100a | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-QFT , positive | splash10-001i-0900000000-af842efa254483370f53 | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-QFT , positive | splash10-001i-1900000000-61c84595e7254f9493c9 | 2017-09-14 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-0006-2090000000-4b8cf489e15586dfdae8 | 2016-08-01 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-006x-5390000000-679252cae00158da4c5a | 2016-08-01 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-0096-9250000000-13f9259c8bd82a744463 | 2016-08-01 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-006y-2792000000-3a1107d332a241bc2b9d | 2016-08-03 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-0f6x-3691000000-15dcc8cfd9d6e268fa7e | 2016-08-03 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-0076-7950000000-99f8f0926e3b4607c61b | 2016-08-03 | View Spectrum |
|
|---|
| Toxicity Profile |
|---|
| Route of Exposure | Readily absorbed through the GI tract (~70%) |
|---|
| Mechanism of Toxicity | Capecitabine is a prodrug that is selectively tumour-activated to its cytotoxic moiety, fluorouracil, by thymidine phosphorylase. Fluorouracil is further metabolized to two active metabolites, 5-fluoro-2-deoxyuridine monophosphate (FdUMP) and 5-fluorouridine triphosphate (FUTP), within normal and tumour cells. FdUMP inhibits DNA synthesis by reducing normal thymidine production, while FUTP inhibits RNA and protein synthesis by competing with uridine triphosphate.3 The active moiety of capecitabine, fluorouracil, is cell cycle phase-specific (Sphase). Both normal and tumor cells metabolize 5-FU to 5-fluoro-2-deoxyuridine monophosphate (FdUMP) and 5-fluorouridine triphosphate (FUTP). These metabolites cause cell injury by two different mechanisms. First, FdUMP and the folate cofactor, N5-10-methylenetetrahydrofolate, bind to thymidylate synthase (TS) to form a covalently bound ternary complex. This binding inhibits the formation of thymidylate from 2'-deaxyuridylate. Thymidylate is the necessary precursor of thymidine triphosphate, which is essential for the synthesis of DNA, so that a deficiency of this compound can inhibit cell division. Second nuclear transcriptional enzymes can mistakenly incorporate FUTP in place of uridine triphosphate (UTP) during the synthesis of RNA. This metabolic error can interfere with RNA processing and protein synthesis. |
|---|
| Metabolism | Metabolized by thymidine phosphorylase to fluoruracil.
Route of Elimination: Capecitabine and its metabolites are predominantly excreted in urine; 95.5% of administered capecitabine dose is recovered in urine. Fecal excretion is minimal (2.6%). The major metabolite excreted in urine is FBAL which represents 57% of the administered dose.About 3% of the administered dose is excreted in urine as unchanged drug.
Half Life: 45-60 minutes for capecitabine and its metabolites. |
|---|
| Toxicity Values | Not Available |
|---|
| Lethal Dose | Not Available |
|---|
| Carcinogenicity (IARC Classification) | No indication of carcinogenicity to humans (not listed by IARC). |
|---|
| Uses/Sources | For the treatment of patients with metastatic breast cancer resistant to both paclitaxel and an anthracycline-containing chemotherapy regimen. May also be used in combination with docetaxel for the treatment of metastatic breast cancer in patients who have failed to respond to, or recurred or relasped during or following anthracycline-containing chemotherapy. Capecitabine is used alone as an adjuvant therapy following the complete resection of primary tumor in patients with stage III colon cancer when monotherapy with fluroprymidine is preferred. The use or capecitabine in combination regimens for advanced gastric cancer is currently being investigated. |
|---|
| Minimum Risk Level | Not Available |
|---|
| Health Effects | Not Available |
|---|
| Symptoms | Not Available |
|---|
| Treatment | Not Available |
|---|
| Normal Concentrations |
|---|
| Not Available |
|---|
| Abnormal Concentrations |
|---|
| Not Available |
|---|
| External Links |
|---|
| DrugBank ID | DB01101 |
|---|
| HMDB ID | HMDB15233 |
|---|
| PubChem Compound ID | 60953 |
|---|
| ChEMBL ID | CHEMBL1773 |
|---|
| ChemSpider ID | 54916 |
|---|
| KEGG ID | C12650 |
|---|
| UniProt ID | Not Available |
|---|
| OMIM ID | |
|---|
| ChEBI ID | 31348 |
|---|
| BioCyc ID | Not Available |
|---|
| CTD ID | Not Available |
|---|
| Stitch ID | Capecitabine |
|---|
| PDB ID | Not Available |
|---|
| ACToR ID | Not Available |
|---|
| Wikipedia Link | Capecitabine |
|---|
| References |
|---|
| Synthesis Reference | DrugSyn.org |
|---|
| MSDS | T3D3507.pdf |
|---|
| General References | - Walko CM, Lindley C: Capecitabine: a review. Clin Ther. 2005 Jan;27(1):23-44. [15763604 ]
- Wagstaff AJ, Ibbotson T, Goa KL: Capecitabine: a review of its pharmacology and therapeutic efficacy in the management of advanced breast cancer. Drugs. 2003;63(2):217-36. [12515569 ]
- Koukourakis GV, Kouloulias V, Koukourakis MJ, Zacharias GA, Zabatis H, Kouvaris J: Efficacy of the oral fluorouracil pro-drug capecitabine in cancer treatment: a review. Molecules. 2008 Aug 27;13(8):1897-922. [18794792 ]
- Twelves C: Vision of the future: capecitabine. Oncologist. 2001;6 Suppl 4:35-9. [11585973 ]
- Milano G, Ferrero JM, Francois E: Comparative pharmacology of oral fluoropyrimidines: a focus on pharmacokinetics, pharmacodynamics and pharmacomodulation. Br J Cancer. 2004 Aug 16;91(4):613-7. [15280932 ]
- de Bono JS, Twelves CJ: The oral fluorinated pyrimidines. Invest New Drugs. 2001;19(1):41-59. [11291832 ]
- Drugs.com [Link]
|
|---|
| Gene Regulation |
|---|
| Up-Regulated Genes | | Gene | Gene Symbol | Gene ID | Interaction | Chromosome | Details |
|---|
|
|---|
| Down-Regulated Genes | Not Available |
|---|