| Record Information |
|---|
| Version | 2.0 |
|---|
| Creation Date | 2009-07-30 17:59:10 UTC |
|---|
| Update Date | 2014-12-24 20:26:08 UTC |
|---|
| Accession Number | T3D3536 |
|---|
| Identification |
|---|
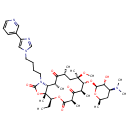
| Common Name | Telithromycin |
|---|
| Class | Small Molecule |
|---|
| Description | Telithromycin, a semi-synthetic erythromycin derivative, belongs to a new chemical class of antibiotics called ketolides. Ketolides have been recently added to the macrolide-lincosamide-streptogramin class of antibiotics. Similar to the macrolide antibiotics, telithromycin prevents bacterial growth by interfering with bacterial protein synthesis. Telithromycin binds to the 50S subunit of the 70S bacterial ribosome and blocks further peptide elongation. Binding occurs simultaneously at to two domains of 23S RNA of the 50S ribosomal subunit, domain II and V, where older macrolides bind only to one. It is used to treat mild to moderate respiratory infections. |
|---|
| Compound Type | - Amine
- Anti-Bacterial Agent
- Drug
- Ester
- Ether
- Ketolide
- Metabolite
- Organic Compound
- Synthetic Compound
|
|---|
| Chemical Structure | |
|---|
| Synonyms | |
|---|
| Chemical Formula | C43H65N5O10 |
|---|
| Average Molecular Mass | 812.004 g/mol |
|---|
| Monoisotopic Mass | 811.473 g/mol |
|---|
| CAS Registry Number | 191114-48-4 |
|---|
| IUPAC Name | (3aR,4S,7R,9R,10R,11R,13R,15R,15aR)-10-{[(2S,3R,4S,6R)-4-(dimethylamino)-3-hydroxy-6-methyloxan-2-yl]oxy}-4-ethyl-11-methoxy-3a,7,9,11,13,15-hexamethyl-1-{4-[4-(pyridin-3-yl)-1H-imidazol-1-yl]butyl}-tetradecahydro-1H-oxacyclotetradeca[4,3-d][1,3]oxazole-2,6,8,14-tetrone |
|---|
| Traditional Name | telithromycin |
|---|
| SMILES | [H][C@]12N(CCCCN3C=NC(=C3)C3=CN=CC=C3)C(=O)O[C@@]1(C)[C@]([H])(CC)OC(=O)[C@]([H])(C)C(=O)[C@]([H])(C)[C@@]([H])(O[C@]1([H])O[C@]([H])(C)C[C@]([H])(N(C)C)[C@@]1([H])O)[C@@](C)(C[C@@]([H])(C)C(=O)[C@]2([H])C)OC |
|---|
| InChI Identifier | InChI=1S/C43H65N5O10/c1-12-33-43(8)37(48(41(53)58-43)19-14-13-18-47-23-31(45-24-47)30-16-15-17-44-22-30)27(4)34(49)25(2)21-42(7,54-11)38(28(5)35(50)29(6)39(52)56-33)57-40-36(51)32(46(9)10)20-26(3)55-40/h15-17,22-29,32-33,36-38,40,51H,12-14,18-21H2,1-11H3/t25-,26-,27+,28+,29-,32+,33+,36-,37-,38-,40+,42-,43+/m1/s1 |
|---|
| InChI Key | InChIKey=LJVAJPDWBABPEJ-RMNISARHSA-N |
|---|
| Chemical Taxonomy |
|---|
| Description | belongs to the class of organic compounds known as aminoglycosides. These are molecules or a portion of a molecule composed of amino-modified sugars. |
|---|
| Kingdom | Organic compounds |
|---|
| Super Class | Organic oxygen compounds |
|---|
| Class | Organooxygen compounds |
|---|
| Sub Class | Carbohydrates and carbohydrate conjugates |
|---|
| Direct Parent | Aminoglycosides |
|---|
| Alternative Parents | |
|---|
| Substituents | - Aminoglycoside core
- N-substituted imidazole
- Oxane
- Oxazolidinone
- Pyridine
- 1,3-dicarbonyl compound
- Oxazolidine
- Azole
- Heteroaromatic compound
- Carbamic acid ester
- Imidazole
- Tertiary aliphatic amine
- Tertiary amine
- Secondary alcohol
- Lactone
- Carbonic acid derivative
- Amino acid or derivatives
- Carboxylic acid ester
- Cyclic ketone
- Ketone
- 1,2-aminoalcohol
- Acetal
- Carboxylic acid derivative
- Oxacycle
- Dialkyl ether
- Ether
- Azacycle
- Organoheterocyclic compound
- Monocarboxylic acid or derivatives
- Carbonyl group
- Organic nitrogen compound
- Alcohol
- Amine
- Organopnictogen compound
- Organic oxide
- Organonitrogen compound
- Hydrocarbon derivative
- Aromatic heteropolycyclic compound
|
|---|
| Molecular Framework | Aromatic heteropolycyclic compounds |
|---|
| External Descriptors | Not Available |
|---|
| Biological Properties |
|---|
| Status | Detected and Not Quantified |
|---|
| Origin | Exogenous |
|---|
| Cellular Locations | |
|---|
| Biofluid Locations | Not Available |
|---|
| Tissue Locations | Not Available |
|---|
| Pathways | | Name | SMPDB Link | KEGG Link |
|---|
| Telithromycin Pathway | Not Available | Not Available |
|
|---|
| Applications | Not Available |
|---|
| Biological Roles | Not Available |
|---|
| Chemical Roles | Not Available |
|---|
| Physical Properties |
|---|
| State | Solid |
|---|
| Appearance | White powder. |
|---|
| Experimental Properties | | Property | Value |
|---|
| Melting Point | 176-188°C | | Boiling Point | Not Available | | Solubility | 300 mg/L | | LogP | 3 |
|
|---|
| Predicted Properties | |
|---|
| Spectra |
|---|
| Spectra | | Spectrum Type | Description | Splash Key | Deposition Date | View |
|---|
| Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_1_1) - 70eV, Positive | Not Available | 2021-10-19 | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_1_2) - 70eV, Positive | Not Available | 2021-10-19 | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_1_3) - 70eV, Positive | Not Available | 2021-10-19 | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_1_4) - 70eV, Positive | Not Available | 2021-10-19 | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_1_5) - 70eV, Positive | Not Available | 2021-10-19 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-0bti-0110009330-a597d6786f2ec155d709 | 2016-08-03 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-0a4r-2311119000-a3fc63423a63d99a5fb7 | 2016-08-03 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-0udj-5692335100-c9dcfe66d206e059a42a | 2016-08-03 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-0ik9-0912217680-c23ce8bf8a7dc6585c65 | 2016-08-03 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-0f6x-0913128500-dfbbb5029daf5212939d | 2016-08-03 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-0ktf-7812694000-d210db4913a3e8fedd3d | 2016-08-03 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-03di-0100001190-101d9f97452eadd98caa | 2021-10-11 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-074i-2400009320-dd1ca952f0b45c335762 | 2021-10-11 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-002f-2900002100-7a252e17a6b4beea42fb | 2021-10-11 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-03di-0110001390-4bf88a3178c537fe09a1 | 2021-10-11 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-08gi-1900002750-173fb451507d24f93587 | 2021-10-11 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-06r2-3910001000-6b208b50ef647efcc9fc | 2021-10-11 | View Spectrum |
|
|---|
| Toxicity Profile |
|---|
| Route of Exposure | Absolute bioavailability is approximately 57%. Maximal concentrations are reached 0.5 - 4 hours following oral administration. Food intake does not affected absorption. |
|---|
| Mechanism of Toxicity | Telithromycin acts by binding to domains II and V of 23S rRNA of the 50S ribosomal subunit. By binding at domain II, telithromycin retains activity against gram-positive cocci (e.g., Streptococcus pneumoniae) in the presence of resistance mediated by methylases (erm genes) that alter the domain V binding site of telithromycin. Telithromycin may also inhibit the assembly of nascent ribosomal units. Compared to erythromycin A, telithromycin binds to the 23S rRNA with 10 times greater affinity in erythromycin-susceptible organisms and 25 times greater affinity in macrolide-resistant strains. This increased binding affinity may be conferred by the C11-12 carbamate side chain of telithromycin. The side chain appears to maintain binding at domain II in the presence of resistance mediated by alterations in domain V. |
|---|
| Metabolism | Hepatic - estimated 50% metabolized by CYP3A4 and 50% metabolized independent of cytochrome P450
Route of Elimination: The systemically available telithromycin is eliminated by multiple pathways as follows: 7% of the dose is excreted unchanged in feces by biliary and/or intestinal secretion; 13% of the dose is excreted unchanged in urine by renal excretion; and 37% of the dose is metabolized by the liver.
Half Life: Main elimination half-life is 2-3 hours; terminal elimination half-life is 10 hours |
|---|
| Toxicity Values | LD50>2000 mg/kg (PO in rats). |
|---|
| Lethal Dose | Not Available |
|---|
| Carcinogenicity (IARC Classification) | No indication of carcinogenicity to humans (not listed by IARC). |
|---|
| Uses/Sources | For the treatment of Pneumococcal infection, acute sinusitis, acute bacterial tonsillitis, acute bronchitis and bronchiolitis, lower respiratory tract infection and lobar (pneumococcal) pneumonia. |
|---|
| Minimum Risk Level | Not Available |
|---|
| Health Effects | Not Available |
|---|
| Symptoms | Most common side-effects are gastrointestinal, including diarrhea, nausea, abdominal pain and vomiting. Headache and disturbances in taste also occur. Less common side-effects include palpitations, blurred vision and rashes. [Wikipedia] |
|---|
| Treatment | Not Available |
|---|
| Normal Concentrations |
|---|
| Not Available |
|---|
| Abnormal Concentrations |
|---|
| Not Available |
|---|
| External Links |
|---|
| DrugBank ID | DB00976 |
|---|
| HMDB ID | HMDB15111 |
|---|
| PubChem Compound ID | 5462516 |
|---|
| ChEMBL ID | CHEMBL1136 |
|---|
| ChemSpider ID | 26329513 |
|---|
| KEGG ID | C12009 |
|---|
| UniProt ID | Not Available |
|---|
| OMIM ID | |
|---|
| ChEBI ID | Not Available |
|---|
| BioCyc ID | Not Available |
|---|
| CTD ID | Not Available |
|---|
| Stitch ID | Telithromycin |
|---|
| PDB ID | Not Available |
|---|
| ACToR ID | Not Available |
|---|
| Wikipedia Link | Telithromycin |
|---|
| References |
|---|
| Synthesis Reference | Suhas Sohani, Mandar Deodhar, Nishant Patel, Manish Patel, Mahesh Davadra, Vinodhamar Kansal, “Process for the Preparation of Telithromycin.” U.S. Patent US20070260066, issued November 08, 2007. |
|---|
| MSDS | Link |
|---|
| General References | - Clay KD, Hanson JS, Pope SD, Rissmiller RW, Purdum PP 3rd, Banks PM: Brief communication: severe hepatotoxicity of telithromycin: three case reports and literature review. Ann Intern Med. 2006 Mar 21;144(6):415-20. Epub 2006 Feb 15. [16481451 ]
- Drugs.com [Link]
|
|---|
| Gene Regulation |
|---|
| Up-Regulated Genes | Not Available |
|---|
| Down-Regulated Genes | Not Available |
|---|