| Record Information |
|---|
| Version | 2.0 |
|---|
| Creation Date | 2014-09-11 05:14:58 UTC |
|---|
| Update Date | 2014-12-24 20:26:56 UTC |
|---|
| Accession Number | T3D4754 |
|---|
| Identification |
|---|
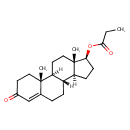
| Common Name | Testosterone Propionate |
|---|
| Class | Small Molecule |
|---|
| Description | An ester of testosterone with a propionate substitution at the 17-beta position. |
|---|
| Compound Type | - Drug
- Ester
- Ether
- Metabolite
- Organic Compound
- Synthetic Compound
|
|---|
| Chemical Structure | |
|---|
| Synonyms | | Synonym | | Testex | | Testosterone propionate | | Testosterone propionic acid |
|
|---|
| Chemical Formula | C22H32O3 |
|---|
| Average Molecular Mass | 344.488 g/mol |
|---|
| Monoisotopic Mass | 344.235 g/mol |
|---|
| CAS Registry Number | 57-85-2 |
|---|
| IUPAC Name | (1S,2R,10R,11S,14S,15S)-2,15-dimethyl-5-oxotetracyclo[8.7.0.0^{2,7}.0^{11,15}]heptadec-6-en-14-yl propanoate |
|---|
| Traditional Name | (1S,2R,10R,11S,14S,15S)-2,15-dimethyl-5-oxotetracyclo[8.7.0.0^{2,7}.0^{11,15}]heptadec-6-en-14-yl propanoate |
|---|
| SMILES | [H][C@@]1(CC[C@@]2([H])[C@]3([H])CCC4=CC(=O)CC[C@]4(C)[C@@]3([H])CC[C@]12C)OC(=O)CC |
|---|
| InChI Identifier | InChI=1S/C22H32O3/c1-4-20(24)25-19-8-7-17-16-6-5-14-13-15(23)9-11-21(14,2)18(16)10-12-22(17,19)3/h13,16-19H,4-12H2,1-3H3/t16-,17-,18-,19-,21-,22-/m0/s1 |
|---|
| InChI Key | InChIKey=PDMMFKSKQVNJMI-BLQWBTBKSA-N |
|---|
| Chemical Taxonomy |
|---|
| Description | belongs to the class of organic compounds known as steroid esters. Steroid esters are compounds containing a steroid moiety which bears a carboxylic acid ester group. |
|---|
| Kingdom | Organic compounds |
|---|
| Super Class | Lipids and lipid-like molecules |
|---|
| Class | Steroids and steroid derivatives |
|---|
| Sub Class | Steroid esters |
|---|
| Direct Parent | Steroid esters |
|---|
| Alternative Parents | |
|---|
| Substituents | - Steroid ester
- Androgen-skeleton
- Androstane-skeleton
- 3-oxosteroid
- 3-oxo-delta-4-steroid
- Oxosteroid
- Delta-4-steroid
- Cyclohexenone
- Ketone
- Carboxylic acid ester
- Cyclic ketone
- Monocarboxylic acid or derivatives
- Carboxylic acid derivative
- Organooxygen compound
- Carbonyl group
- Organic oxide
- Organic oxygen compound
- Hydrocarbon derivative
- Aliphatic homopolycyclic compound
|
|---|
| Molecular Framework | Aliphatic homopolycyclic compounds |
|---|
| External Descriptors | |
|---|
| Biological Properties |
|---|
| Status | Detected and Not Quantified |
|---|
| Origin | Exogenous |
|---|
| Cellular Locations | - Cytoplasm
- Extracellular
- Membrane
|
|---|
| Biofluid Locations | Not Available |
|---|
| Tissue Locations | Not Available |
|---|
| Pathways | Not Available |
|---|
| Applications | Not Available |
|---|
| Biological Roles | Not Available |
|---|
| Chemical Roles | Not Available |
|---|
| Physical Properties |
|---|
| State | Solid |
|---|
| Appearance | White powder. |
|---|
| Experimental Properties | | Property | Value |
|---|
| Melting Point | 120°C | | Boiling Point | Not Available | | Solubility | 1.48 mg/L (at 25°C) | | LogP | Not Available |
|
|---|
| Predicted Properties | |
|---|
| Spectra |
|---|
| Spectra | | Spectrum Type | Description | Splash Key | Deposition Date | View |
|---|
| Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | splash10-00or-4395000000-e888bc1b6c2a4c99d301 | 2017-09-01 | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | Not Available | 2021-10-12 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - 60V, Positive | splash10-0a4j-9700000000-764405bfcba7c5fe17d4 | 2021-09-20 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - 45V, Positive | splash10-052b-8900000000-4b2c3ab6d4ce48b53a23 | 2021-09-20 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - 15V, Positive | splash10-0002-0009000000-0b70d9f4dfe7d2b1882f | 2021-09-20 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - 30V, Positive | splash10-052b-8941000000-71e897f7bddbd3065fb6 | 2021-09-20 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - 90V, Positive | splash10-0a6s-9300000000-ae3c374d8e9e15f95613 | 2021-09-20 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - 75V, Positive | splash10-0a4j-9500000000-8e8b74c239d0be2293ab | 2021-09-20 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - 45V, Positive | splash10-052b-8900000000-3b051eb836dd8f828e7e | 2021-09-20 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-052b-4069000000-515cef716c1456ad0ff1 | 2016-08-03 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-0a4i-6392000000-9fad427c5fd42708c35b | 2016-08-03 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-0a4r-3590000000-8e5391da57f43fc2f8d0 | 2016-08-03 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-0006-1049000000-96980216f4e665c10566 | 2016-08-03 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-000i-4094000000-3757f1e13b7d134d9d32 | 2016-08-03 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-0ab9-4090000000-94160172dab6424eaf2d | 2016-08-03 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-00ds-0094000000-8c969c8df525e9804ec3 | 2021-10-11 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-044s-0962000000-2061e148bffa88e3c96e | 2021-10-11 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-0a4i-2910000000-d364cf854d78cc7d4e9f | 2021-10-11 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-00dl-9008000000-fde8ba8c4ed57693204c | 2021-10-11 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-00di-9001000000-02ee41375158c3c608c9 | 2021-10-11 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-0ab9-9061000000-82ba060dba23b610bd79 | 2021-10-11 | View Spectrum | | MS | Mass Spectrum (Electron Ionization) | splash10-0a4i-9800000000-8aff9c2cbe0b5c20d85a | 2014-09-20 | View Spectrum | | 1D NMR | 13C NMR Spectrum (1D, 50.18 MHz, CDCl3, experimental) | Not Available | 2014-09-23 | View Spectrum |
|
|---|
| Toxicity Profile |
|---|
| Route of Exposure | Not Available |
|---|
| Mechanism of Toxicity | The effects of testosterone in humans and other vertebrates occur by way of two main mechanisms: by activation of the androgen receptor (directly or as DHT), and by conversion to estradiol and activation of certain estrogen receptors. Free testosterone (T) is transported into the cytoplasm of target tissue cells, where it can bind to the androgen receptor, or can be reduced to 5α-dihydrotestosterone (DHT) by the cytoplasmic enzyme 5α-reductase. DHT binds to the same androgen receptor even more strongly than T, so that its androgenic potency is about 2.5 times that of T. The T-receptor or DHT-receptor complex undergoes a structural change that allows it to move into the cell nucleus and bind directly to specific nucleotide sequences of the chromosomal DNA. The areas of binding are called hormone response elements (HREs), and influence transcriptional activity of certain genes, producing the androgen effects. |
|---|
| Metabolism | Testosterone propionate is rapidly hydrolysed into testosterone. Testosterone is metabolized to 17-keto steroids through two different pathways. The major active metabolites are estradiol and dihydrotestosterone (DHT).
Route of Elimination: About 90% of a dose of testosterone given intramuscularly is excreted in the urine as glucuronic and sulfuric acid conjugates of testosterone and its metabolites; about 6% of a dose is excreted in the feces, mostly in the unconjugated form. |
|---|
| Toxicity Values | Not Available |
|---|
| Lethal Dose | Not Available |
|---|
| Carcinogenicity (IARC Classification) | No indication of carcinogenicity to humans (not listed by IARC). |
|---|
| Uses/Sources | Testosterone propionate is an anabolic steroid and a short ester form of testosterone that becomes active in the body. It is often used for muscle mass building. |
|---|
| Minimum Risk Level | Not Available |
|---|
| Health Effects | Not Available |
|---|
| Symptoms | Side effects include amnesia, anxiety, discolored hair, dizziness, dry skin, hirsutism, hostility, impaired urination, paresthesia, penis disorder, peripheral edema, sweating, and vasodilation. |
|---|
| Treatment | Not Available |
|---|
| Normal Concentrations |
|---|
| Not Available |
|---|
| Abnormal Concentrations |
|---|
| Not Available |
|---|
| External Links |
|---|
| DrugBank ID | DB01420 |
|---|
| HMDB ID | HMDB15489 |
|---|
| PubChem Compound ID | 5995 |
|---|
| ChEMBL ID | CHEMBL1170 |
|---|
| ChemSpider ID | 5774 |
|---|
| KEGG ID | C08158 |
|---|
| UniProt ID | Not Available |
|---|
| OMIM ID | |
|---|
| ChEBI ID | 290629 |
|---|
| BioCyc ID | Not Available |
|---|
| CTD ID | Not Available |
|---|
| Stitch ID | Not Available |
|---|
| PDB ID | Not Available |
|---|
| ACToR ID | Not Available |
|---|
| Wikipedia Link | Not Available |
|---|
| References |
|---|
| Synthesis Reference | Not Available |
|---|
| MSDS | T3D4754.pdf |
|---|
| General References | Not Available |
|---|
| Gene Regulation |
|---|
| Up-Regulated Genes | Not Available |
|---|
| Down-Regulated Genes | Not Available |
|---|