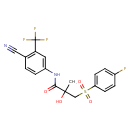| Record Information |
|---|
| Version | 2.0 |
|---|
| Creation Date | 2014-09-11 05:17:56 UTC |
|---|
| Update Date | 2014-12-24 20:26:57 UTC |
|---|
| Accession Number | T3D4818 |
|---|
| Identification |
|---|
| Common Name | Bicalutamide |
|---|
| Class | Small Molecule |
|---|
| Description | Bicalutamide is an oral non-steroidal anti-androgen for prostate cancer. It binds to the androgen receptor. |
|---|
| Compound Type | - Drug
- Ether
- Metabolite
- Nitrile
- Organic Compound
- Organofluoride
- Synthetic Compound
|
|---|
| Chemical Structure | |
|---|
| Synonyms | |
|---|
| Chemical Formula | C18H14F4N2O4S |
|---|
| Average Molecular Mass | 430.373 g/mol |
|---|
| Monoisotopic Mass | 430.061 g/mol |
|---|
| CAS Registry Number | 90357-06-5 |
|---|
| IUPAC Name | N-[4-cyano-3-(trifluoromethyl)phenyl]-3-(4-fluorobenzenesulfonyl)-2-hydroxy-2-methylpropanamide |
|---|
| Traditional Name | bicalutamide |
|---|
| SMILES | CC(O)(CS(=O)(=O)C1=CC=C(F)C=C1)C(O)=NC1=CC(=C(C=C1)C#N)C(F)(F)F |
|---|
| InChI Identifier | InChI=1/C18H14F4N2O4S/c1-17(26,10-29(27,28)14-6-3-12(19)4-7-14)16(25)24-13-5-2-11(9-23)15(8-13)18(20,21)22/h2-8,26H,10H2,1H3,(H,24,25) |
|---|
| InChI Key | InChIKey=LKJPYSCBVHEWIU-UHFFFAOYNA-N |
|---|
| Chemical Taxonomy |
|---|
| Description | belongs to the class of organic compounds known as trifluoromethylbenzenes. These are organofluorine compounds that contain a benzene ring substituted with one or more trifluoromethyl groups. |
|---|
| Kingdom | Organic compounds |
|---|
| Super Class | Benzenoids |
|---|
| Class | Benzene and substituted derivatives |
|---|
| Sub Class | Trifluoromethylbenzenes |
|---|
| Direct Parent | Trifluoromethylbenzenes |
|---|
| Alternative Parents | |
|---|
| Substituents | - Trifluoromethylbenzene
- Benzenesulfonyl group
- Anilide
- Benzonitrile
- N-arylamide
- Fluorobenzene
- Halobenzene
- Aryl fluoride
- Aryl halide
- Tertiary alcohol
- Sulfonyl
- Sulfone
- Carboxamide group
- Secondary carboxylic acid amide
- Carboxylic acid derivative
- Carbonitrile
- Nitrile
- Organohalogen compound
- Alkyl halide
- Organosulfur compound
- Hydrocarbon derivative
- Organic oxide
- Alkyl fluoride
- Alcohol
- Carbonyl group
- Organopnictogen compound
- Organic oxygen compound
- Organic nitrogen compound
- Organofluoride
- Organonitrogen compound
- Organooxygen compound
- Aromatic homomonocyclic compound
|
|---|
| Molecular Framework | Aromatic homomonocyclic compounds |
|---|
| External Descriptors | Not Available |
|---|
| Biological Properties |
|---|
| Status | Detected and Not Quantified |
|---|
| Origin | Exogenous |
|---|
| Cellular Locations | |
|---|
| Biofluid Locations | Not Available |
|---|
| Tissue Locations | Not Available |
|---|
| Pathways | Not Available |
|---|
| Applications | Not Available |
|---|
| Biological Roles | Not Available |
|---|
| Chemical Roles | Not Available |
|---|
| Physical Properties |
|---|
| State | Solid |
|---|
| Appearance | White powder. |
|---|
| Experimental Properties | | Property | Value |
|---|
| Melting Point | 191-193°C | | Boiling Point | Not Available | | Solubility | 5 mg/L | | LogP | 2.5 |
|
|---|
| Predicted Properties | |
|---|
| Spectra |
|---|
| Spectra | | Spectrum Type | Description | Splash Key | Deposition Date | View |
|---|
| Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | splash10-08g3-8980200000-64d743e9fc999bfbcba9 | 2017-09-01 | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (1 TMS) - 70eV, Positive | splash10-01p9-7953200000-a9b28d0415e04d7763bb | 2017-10-06 | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | Not Available | 2021-10-12 | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | Not Available | 2021-10-12 | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | Not Available | 2021-10-12 | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_1_2) - 70eV, Positive | Not Available | 2021-11-03 | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TBDMS_1_1) - 70eV, Positive | Not Available | 2021-11-03 | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TBDMS_1_2) - 70eV, Positive | Not Available | 2021-11-03 | View Spectrum |
|
|---|
| Toxicity Profile |
|---|
| Route of Exposure | Bicalutamide is well-absorbed following oral administration, although the absolute bioavailability is unknown. |
|---|
| Mechanism of Toxicity | Bicalutamide competes with androgen for the binding of androgen receptors, consequently blocking the action of androgens of adrenal and testicular origin which stimulate the growth of normal and malignant prostatic tissue. Organic nitriles decompose into cyanide ions both in vivo and in vitro. Consequently the primary mechanism of toxicity for organic nitriles is their production of toxic cyanide ions or hydrogen cyanide. Cyanide is an inhibitor of cytochrome c oxidase in the fourth complex of the electron transport chain (found in the membrane of the mitochondria of eukaryotic cells). It complexes with the ferric iron atom in this enzyme. The binding of cyanide to this cytochrome prevents transport of electrons from cytochrome c oxidase to oxygen. As a result, the electron transport chain is disrupted and the cell can no longer aerobically produce ATP for energy. Tissues that mainly depend on aerobic respiration, such as the central nervous system and the heart, are particularly affected. Cyanide is also known produce some of its toxic effects by binding to catalase, glutathione peroxidase, methemoglobin, hydroxocobalamin, phosphatase, tyrosinase, ascorbic acid oxidase, xanthine oxidase, succinic dehydrogenase, and Cu/Zn superoxide dismutase. Cyanide binds to the ferric ion of methemoglobin to form inactive cyanmethemoglobin. (1) |
|---|
| Metabolism | Bicalutamide undergoes stereo specific metabolism. The S (inactive) isomer is metabolized primarily by glucuronidation. The R (active) isomer also undergoes glucuronidation but is predominantly oxidized to an inactive metabolite followed by glucuronidation.
Half Life: 5.9 days |
|---|
| Toxicity Values | Not Available |
|---|
| Lethal Dose | Not Available |
|---|
| Carcinogenicity (IARC Classification) | No indication of carcinogenicity to humans (not listed by IARC). |
|---|
| Uses/Sources | For treatment (together with surgery or LHRH analogue) of advanced prostatic cancer. |
|---|
| Minimum Risk Level | Not Available |
|---|
| Health Effects | Not Available |
|---|
| Symptoms | Not Available |
|---|
| Treatment | Not Available |
|---|
| Normal Concentrations |
|---|
| Not Available |
|---|
| Abnormal Concentrations |
|---|
| Not Available |
|---|
| External Links |
|---|
| DrugBank ID | DB01128 |
|---|
| HMDB ID | HMDB15260 |
|---|
| PubChem Compound ID | 2375 |
|---|
| ChEMBL ID | CHEMBL409 |
|---|
| ChemSpider ID | 50614 |
|---|
| KEGG ID | C08160 |
|---|
| UniProt ID | Not Available |
|---|
| OMIM ID | |
|---|
| ChEBI ID | 100717 |
|---|
| BioCyc ID | Not Available |
|---|
| CTD ID | C053541 |
|---|
| Stitch ID | Not Available |
|---|
| PDB ID | Not Available |
|---|
| ACToR ID | Not Available |
|---|
| Wikipedia Link | Bicalutamide |
|---|
| References |
|---|
| Synthesis Reference | Nnochiri Ekwuribe, “METHODS OF SYNTHESIZING ACYLANILIDES INCLUDING BICALUTAMIDE AND DERIVATIVES THEREOF.” U.S. Patent US20020165406, issued November 07, 2002. |
|---|
| MSDS | Link |
|---|
| General References | - Wikipedia. Cyanide poisoning. Last Updated 30 March 2009. [Link]
|
|---|
| Gene Regulation |
|---|
| Up-Regulated Genes | Not Available |
|---|
| Down-Regulated Genes | Not Available |
|---|