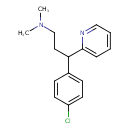
Chlorpheniramine (T3D2989)
| Record Information | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Version | 2.0 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Creation Date | 2009-07-21 20:28:24 UTC | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Update Date | 2014-12-24 20:25:54 UTC | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Accession Number | T3D2989 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Identification | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Common Name | Chlorpheniramine | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Class | Small Molecule | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Description | Chlorpheniramine is a histamine H1 antagonist used in allergic reactions, hay fever, rhinitis, urticaria, and asthma. It has also been used in veterinary applications. One of the most widely used of the classical antihistaminics, it generally causes less drowsiness and sedation than Promethazine. -- Pubchem; Chlorphenamine or chlorpheniramine, commonly marketed as its salt chlorphenamine maleate (Chlor- Trimeton, Piriton, Chlor- Tripolon), is a first generation antihistamine used in the prevention of the symptoms of allergic conditions such as rhinitis and urticaria.- wikipedia. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Compound Type |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Chemical Structure | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Synonyms |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Chemical Formula | C16H19ClN2 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Average Molecular Mass | 274.788 g/mol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Monoisotopic Mass | 274.124 g/mol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| CAS Registry Number | 132-22-9 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| IUPAC Name | [3-(4-chlorophenyl)-3-(pyridin-2-yl)propyl]dimethylamine | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Traditional Name | chlorpheniramine | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| SMILES | CN(C)CCC(C1=CC=C(Cl)C=C1)C1=CC=CC=N1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| InChI Identifier | InChI=1/C16H19ClN2/c1-19(2)12-10-15(16-5-3-4-11-18-16)13-6-8-14(17)9-7-13/h3-9,11,15H,10,12H2,1-2H3 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| InChI Key | InChIKey=SOYKEARSMXGVTM-UHFFFAOYNA-N | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Chemical Taxonomy | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Description | belongs to the class of organic compounds known as pheniramines. Pheniramines are compounds containing a pheniramine moiety, which is structurally characterized by the presence of a 2-benzylpyridine linked to an dimethyl(propyl)amine to form a dimethyl[3-phenyl-3-(pyridin-2-yl)propyl]amine skeleton. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Kingdom | Organic compounds | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Super Class | Organoheterocyclic compounds | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Class | Pyridines and derivatives | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Sub Class | Pheniramines | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Direct Parent | Pheniramines | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Alternative Parents | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Substituents |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Molecular Framework | Aromatic heteromonocyclic compounds | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| External Descriptors |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Biological Properties | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Status | Detected and Not Quantified | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Origin | Exogenous | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Cellular Locations |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Biofluid Locations | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Tissue Locations |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Pathways | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Applications | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Biological Roles | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Chemical Roles | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Physical Properties | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| State | Solid | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Appearance | White powder. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Experimental Properties |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Predicted Properties |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Spectra | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Spectra |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Toxicity Profile | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Route of Exposure | Well absorbed in the gastrointestinal tract. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Mechanism of Toxicity | Chlorpheniramine binds to the histamine H1 receptor. This blocks the action of endogenous histamine, which subsequently leads to temporary relief of the negative symptoms brought on by histamine. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Metabolism | Primarily hepatic via Cytochrome P450 (CYP450) enzymes. Half Life: 21-27 hours | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Toxicity Values | Oral LD50 (rat): 306 mg/kg Oral LD50 (mice): 130 mg/kg Oral LD50 (guinea pig): 198 mg/kg LD50: 306 mg/kg (Human) (1) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Lethal Dose | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Carcinogenicity (IARC Classification) | No indication of carcinogenicity to humans (not listed by IARC). | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Uses/Sources | Used used in the prevention of the symptoms of allergic conditions such as rhinitis, urticaria, allergy, common cold, asthma and hay fever. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Minimum Risk Level | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Health Effects | Chlorpheniramine posesses serotonergic and norepinephrinergic effects. It has been shown to work as a serotonin-norepinephrine reuptake inhibitor or SNRI. [Wikipedia] | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symptoms | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Treatment | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Normal Concentrations | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Not Available | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Abnormal Concentrations | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Not Available | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| External Links | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| DrugBank ID | DB01114 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| HMDB ID | HMDB01944 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PubChem Compound ID | 2725 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| ChEMBL ID | CHEMBL505 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| ChemSpider ID | 2624 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| KEGG ID | C06905 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| UniProt ID | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| OMIM ID | 146840 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| ChEBI ID | 52010 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| BioCyc ID | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| CTD ID | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Stitch ID | Chlorpheniramine | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PDB ID | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| ACToR ID | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Wikipedia Link | Chlorpheniramine | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| References | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Synthesis Reference | Anil M. Salpekar, John Johnson, “Acetaminophen compositions containing low doses of chlorpheniramine maleate, method for preparing same and tablets formed therefrom.” U.S. Patent US4631284, issued April, 1975. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| MSDS | Link | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| General References |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Gene Regulation | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Up-Regulated Genes | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Down-Regulated Genes | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Targets
- General Function:
- Histamine receptor activity
- Specific Function:
- In peripheral tissues, the H1 subclass of histamine receptors mediates the contraction of smooth muscles, increase in capillary permeability due to contraction of terminal venules, and catecholamine release from adrenal medulla, as well as mediating neurotransmission in the central nervous system.
- Gene Name:
- HRH1
- Uniprot ID:
- P35367
- Molecular Weight:
- 55783.61 Da
Binding/Activity Constants
| Type | Value | Assay Type | Assay Source |
|---|---|---|---|
| Inhibitory | 0.00251 uM | Not Available | BindingDB 35938 |
| Inhibitory | 0.00267 uM | Not Available | BindingDB 35938 |
| Inhibitory | 0.00481 uM | Not Available | BindingDB 35938 |
| Inhibitory | 0.007 uM | Not Available | BindingDB 50007468 |
| Inhibitory | 0.015 uM | Not Available | BindingDB 35938 |
| Inhibitory | 0.03 uM | Not Available | BindingDB 35938 |
| IC50 | 0.0088 uM | Not Available | BindingDB 50007468 |
| IC50 | 0.066 uM | Not Available | BindingDB 50007468 |
References
- Chen X, Ji ZL, Chen YZ: TTD: Therapeutic Target Database. Nucleic Acids Res. 2002 Jan 1;30(1):412-5. [11752352 ]
- Salata JJ, Jurkiewicz NK, Wallace AA, Stupienski RF 3rd, Guinosso PJ Jr, Lynch JJ Jr: Cardiac electrophysiological actions of the histamine H1-receptor antagonists astemizole and terfenadine compared with chlorpheniramine and pyrilamine. Circ Res. 1995 Jan;76(1):110-9. [8001268 ]
- Tagawa M, Kano M, Okamura N, Higuchi M, Matsuda M, Mizuki Y, Arai H, Iwata R, Fujii T, Komemushi S, Ido T, Itoh M, Sasaki H, Watanabe T, Yanai K: Neuroimaging of histamine H1-receptor occupancy in human brain by positron emission tomography (PET): a comparative study of ebastine, a second-generation antihistamine, and (+)-chlorpheniramine, a classical antihistamine. Br J Clin Pharmacol. 2001 Nov;52(5):501-9. [11736858 ]
- Hasenohrl RU, Kuhlen A, Frisch C, Galosi R, Brandao ML, Huston JP: Comparison of intra-accumbens injection of histamine with histamine H1-receptor antagonist chlorpheniramine in effects on reinforcement and memory parameters. Behav Brain Res. 2001 Oct 15;124(2):203-11. [11640974 ]
- Yasuda SU, Wellstein A, Likhari P, Barbey JT, Woosley RL: Chlorpheniramine plasma concentration and histamine H1-receptor occupancy. Clin Pharmacol Ther. 1995 Aug;58(2):210-20. [7648771 ]
- Nicholson AN, Pascoe PA, Turner C, Ganellin CR, Greengrass PM, Casy AF, Mercer AD: Sedation and histamine H1-receptor antagonism: studies in man with the enantiomers of chlorpheniramine and dimethindene. Br J Pharmacol. 1991 Sep;104(1):270-6. [1686208 ]
- Sleevi MC, Cale AD Jr, Gero TW, Jaques LW, Welstead WJ, Johnson AF, Kilpatrick BF, Demian I, Nolan JC, Jenkins H: Optical isomers of rocastine and close analogues: synthesis and H1 antihistaminic activity of its enantiomers and their structural relationship to the classical antihistamines. J Med Chem. 1991 Apr;34(4):1314-28. [1673158 ]
- Kelly JX, Smilkstein MJ, Cooper RA, Lane KD, Johnson RA, Janowsky A, Dodean RA, Hinrichs DJ, Winter R, Riscoe M: Design, synthesis, and evaluation of 10-N-substituted acridones as novel chemosensitizers in Plasmodium falciparum. Antimicrob Agents Chemother. 2007 Nov;51(11):4133-40. Epub 2007 Sep 10. [17846138 ]
- Moguilevsky N, Varsalona F, Noyer M, Gillard M, Guillaume JP, Garcia L, Szpirer C, Szpirer J, Bollen A: Stable expression of human H1-histamine-receptor cDNA in Chinese hamster ovary cells. Pharmacological characterisation of the protein, tissue distribution of messenger RNA and chromosomal localisation of the gene. Eur J Biochem. 1994 Sep 1;224(2):489-95. [7925364 ]
- Moguilevsky N, Varsalona F, Guillaume JP, Noyer M, Gillard M, Daliers J, Henichart JP, Bollen A: Pharmacological and functional characterisation of the wild-type and site-directed mutants of the human H1 histamine receptor stably expressed in CHO cells. J Recept Signal Transduct Res. 1995 Jan-Mar;15(1-4):91-102. [8903934 ]
- Booth RG, Moniri NH, Bakker RA, Choksi NY, Nix WB, Timmerman H, Leurs R: A novel phenylaminotetralin radioligand reveals a subpopulation of histamine H(1) receptors. J Pharmacol Exp Ther. 2002 Jul;302(1):328-36. [12065734 ]
- Procopiou PA, Browning C, Buckley JM, Clark KL, Fechner L, Gore PM, Hancock AP, Hodgson ST, Holmes DS, Kranz M, Looker BE, Morriss KM, Parton DL, Russell LJ, Slack RJ, Sollis SL, Vile S, Watts CJ: The discovery of phthalazinone-based human H1 and H3 single-ligand antagonists suitable for intranasal administration for the treatment of allergic rhinitis. J Med Chem. 2011 Apr 14;54(7):2183-95. doi: 10.1021/jm1013874. Epub 2011 Mar 7. [21381763 ]
- Richelson E, Nelson A: Antagonism by neuroleptics of neurotransmitter receptors of normal human brain in vitro. Eur J Pharmacol. 1984 Aug 17;103(3-4):197-204. [6149136 ]
- Kanba S, Richelson E: Histamine H1 receptors in human brain labelled with [3H]doxepin. Brain Res. 1984 Jun 18;304(1):1-7. [6146381 ]
- Ruck LM, Rizzo CA, Anthes JC, Eckel S, Egan RW, Cuss FM, Hey JA: Synergistic antiallergic activity of combined histamine H1- and cysteinyl leukotriene1-receptor blockade in human bronchus. Life Sci. 2001 May 11;68(25):2825-34. [11432448 ]
- General Function:
- Voltage-gated potassium channel activity involved in ventricular cardiac muscle cell action potential repolarization
- Specific Function:
- Pore-forming (alpha) subunit of voltage-gated inwardly rectifying potassium channel. Channel properties are modulated by cAMP and subunit assembly. Mediates the rapidly activating component of the delayed rectifying potassium current in heart (IKr). Isoforms USO have no channel activity by themself, but modulates channel characteristics by forming heterotetramers with other isoforms which are retained intracellularly and undergo ubiquitin-dependent degradation.
- Gene Name:
- KCNH2
- Uniprot ID:
- Q12809
- Molecular Weight:
- 126653.52 Da
Binding/Activity Constants
| Type | Value | Assay Type | Assay Source |
|---|---|---|---|
| IC50 | 1.1 uM | Not Available | BindingDB 50007468 |
| IC50 | 20.89296 uM | Not Available | BindingDB 50007468 |
| IC50 | 20.893 uM | Not Available | BindingDB 50007468 |
References
- Du LP, Tsai KC, Li MY, You QD, Xia L: The pharmacophore hypotheses of I(Kr) potassium channel blockers: novel class III antiarrhythmic agents. Bioorg Med Chem Lett. 2004 Sep 20;14(18):4771-7. [15324906 ]
- Tobita M, Nishikawa T, Nagashima R: A discriminant model constructed by the support vector machine method for HERG potassium channel inhibitors. Bioorg Med Chem Lett. 2005 Jun 2;15(11):2886-90. [15911273 ]
- Jia L, Sun H: Support vector machines classification of hERG liabilities based on atom types. Bioorg Med Chem. 2008 Jun 1;16(11):6252-60. doi: 10.1016/j.bmc.2008.04.028. Epub 2008 Apr 16. [18448342 ]
- Keseru GM: Prediction of hERG potassium channel affinity by traditional and hologram qSAR methods. Bioorg Med Chem Lett. 2003 Aug 18;13(16):2773-5. [12873512 ]
- General Function:
- Serotonin receptor activity
- Specific Function:
- G-protein coupled receptor for 5-hydroxytryptamine (serotonin). Also functions as a receptor for various ergot alkaloid derivatives and psychoactive substances. Ligand binding causes a conformation change that triggers signaling via guanine nucleotide-binding proteins (G proteins) and modulates the activity of down-stream effectors. Beta-arrestin family members inhibit signaling via G proteins and mediate activation of alternative signaling pathways. Signaling activates a phosphatidylinositol-calcium second messenger system that modulates the activity of phosphatidylinositol 3-kinase and down-stream signaling cascades and promotes the release of Ca(2+) ions from intracellular stores. Plays a role in the regulation of dopamine and 5-hydroxytryptamine release, 5-hydroxytryptamine uptake and in the regulation of extracellular dopamine and 5-hydroxytryptamine levels, and thereby affects neural activity. May play a role in the perception of pain. Plays a role in the regulation of behavior, including impulsive behavior. Required for normal proliferation of embryonic cardiac myocytes and normal heart development. Protects cardiomyocytes against apoptosis. Plays a role in the adaptation of pulmonary arteries to chronic hypoxia. Plays a role in vasoconstriction. Required for normal osteoblast function and proliferation, and for maintaining normal bone density. Required for normal proliferation of the interstitial cells of Cajal in the intestine.
- Gene Name:
- HTR2B
- Uniprot ID:
- P41595
- Molecular Weight:
- 54297.41 Da
Binding/Activity Constants
| Type | Value | Assay Type | Assay Source |
|---|---|---|---|
| Inhibitory | 7.22 uM | Not Available | BindingDB 35938 |
References
- Wainscott DB, Lucaites VL, Kursar JD, Baez M, Nelson DL: Pharmacologic characterization of the human 5-hydroxytryptamine2B receptor: evidence for species differences. J Pharmacol Exp Ther. 1996 Feb;276(2):720-7. [8632342 ]
- General Function:
- Serotonin receptor activity
- Specific Function:
- G-protein coupled receptor for 5-hydroxytryptamine (serotonin). Also functions as a receptor for various drugs and psychoactive substances, including ergot alkaloid derivatives, 1-2,5,-dimethoxy-4-iodophenyl-2-aminopropane (DOI) and lysergic acid diethylamide (LSD). Ligand binding causes a conformation change that triggers signaling via guanine nucleotide-binding proteins (G proteins) and modulates the activity of down-stream effectors. Beta-arrestin family members inhibit signaling via G proteins and mediate activation of alternative signaling pathways. Signaling activates a phosphatidylinositol-calcium second messenger system that modulates the activity of phosphatidylinositol 3-kinase and down-stream signaling cascades and promotes the release of Ca(2+) ions from intracellular stores. Regulates neuronal activity via the activation of short transient receptor potential calcium channels in the brain, and thereby modulates the activation of pro-opiomelacortin neurons and the release of CRH that then regulates the release of corticosterone. Plays a role in the regulation of appetite and eating behavior, responses to anxiogenic stimuli and stress. Plays a role in insulin sensitivity and glucose homeostasis.
- Gene Name:
- HTR2C
- Uniprot ID:
- P28335
- Molecular Weight:
- 51820.705 Da
Binding/Activity Constants
| Type | Value | Assay Type | Assay Source |
|---|---|---|---|
| Inhibitory | >10 uM | Not Available | BindingDB 35938 |
References
- Wainscott DB, Lucaites VL, Kursar JD, Baez M, Nelson DL: Pharmacologic characterization of the human 5-hydroxytryptamine2B receptor: evidence for species differences. J Pharmacol Exp Ther. 1996 Feb;276(2):720-7. [8632342 ]
- General Function:
- Histamine receptor activity
- Specific Function:
- The H4 subclass of histamine receptors could mediate the histamine signals in peripheral tissues. Displays a significant level of constitutive activity (spontaneous activity in the absence of agonist).
- Gene Name:
- HRH4
- Uniprot ID:
- Q9H3N8
- Molecular Weight:
- 44495.375 Da
Binding/Activity Constants
| Type | Value | Assay Type | Assay Source |
|---|---|---|---|
| Inhibitory | 2.91 uM | Not Available | BindingDB 35938 |
References
- Nguyen T, Shapiro DA, George SR, Setola V, Lee DK, Cheng R, Rauser L, Lee SP, Lynch KR, Roth BL, O'Dowd BF: Discovery of a novel member of the histamine receptor family. Mol Pharmacol. 2001 Mar;59(3):427-33. [11179435 ]
- General Function:
- Zinc ion binding
- Specific Function:
- Polycomb group (PcG) protein that specifically recognizes and binds mono- and dimethyllysine residues on target proteins, therey acting as a 'reader' of a network of post-translational modifications. PcG proteins maintain the transcriptionally repressive state of genes: acts as a chromatin compaction factor by recognizing and binding mono- and dimethylated histone H1b/HIST1H1E at 'Lys-26' (H1bK26me1 and H1bK26me2) and histone H4 at 'Lys-20' (H4K20me1 and H4K20me2), leading to condense chromatin and repress transcription. Recognizes and binds p53/TP53 monomethylated at 'Lys-382', leading to repress p53/TP53-target genes. Also recognizes and binds RB1/RB monomethylated at 'Lys-860'. Participates in the ETV6-mediated repression. Probably plays a role in cell proliferation. Overexpression induces multinucleated cells, suggesting that it is required to accomplish normal mitosis.
- Gene Name:
- L3MBTL1
- Uniprot ID:
- Q9Y468
- Molecular Weight:
- 83882.985 Da
Binding/Activity Constants
| Type | Value | Assay Type | Assay Source |
|---|---|---|---|
| IC50 | 41 uM | Not Available | BindingDB 50007468 |
References
- Kireev D, Wigle TJ, Norris-Drouin J, Herold JM, Janzen WP, Frye SV: Identification of non-peptide malignant brain tumor (MBT) repeat antagonists by virtual screening of commercially available compounds. J Med Chem. 2010 Nov 11;53(21):7625-31. doi: 10.1021/jm1007374. [20931980 ]
- General Function:
- Not Available
- Specific Function:
- Putative Polycomb group (PcG) protein. PcG proteins maintain the transcriptionally repressive state of genes, probably via a modification of chromatin, rendering it heritably changed in its expressibility. Required for normal maturation of myeloid progenitor cells (By similarity).
- Gene Name:
- L3MBTL3
- Uniprot ID:
- Q96JM7
- Molecular Weight:
- 88336.035 Da
Binding/Activity Constants
| Type | Value | Assay Type | Assay Source |
|---|---|---|---|
| IC50 | 42 uM | Not Available | BindingDB 50007468 |
References
- Kireev D, Wigle TJ, Norris-Drouin J, Herold JM, Janzen WP, Frye SV: Identification of non-peptide malignant brain tumor (MBT) repeat antagonists by virtual screening of commercially available compounds. J Med Chem. 2010 Nov 11;53(21):7625-31. doi: 10.1021/jm1007374. [20931980 ]
- General Function:
- Zinc ion binding
- Specific Function:
- Putative Polycomb group (PcG) protein. PcG proteins maintain the transcriptionally repressive state of genes, probably via a modification of chromatin, rendering it heritably changed in its expressibility (By similarity).
- Gene Name:
- L3MBTL4
- Uniprot ID:
- Q8NA19
- Molecular Weight:
- 71121.56 Da
Binding/Activity Constants
| Type | Value | Assay Type | Assay Source |
|---|---|---|---|
| IC50 | >100 uM | Not Available | BindingDB 50007468 |
References
- Kireev D, Wigle TJ, Norris-Drouin J, Herold JM, Janzen WP, Frye SV: Identification of non-peptide malignant brain tumor (MBT) repeat antagonists by virtual screening of commercially available compounds. J Med Chem. 2010 Nov 11;53(21):7625-31. doi: 10.1021/jm1007374. [20931980 ]
- General Function:
- Zinc ion binding
- Specific Function:
- Putative Polycomb group (PcG) protein. PcG proteins maintain the transcriptionally repressive state of genes, probably via a modification of chromatin, rendering it heritably changed in its expressibility (By similarity). Specifically binds to monomethylated and dimethylated 'Lys-20' on histone H4.
- Gene Name:
- MBTD1
- Uniprot ID:
- Q05BQ5
- Molecular Weight:
- 70546.805 Da
Binding/Activity Constants
| Type | Value | Assay Type | Assay Source |
|---|---|---|---|
| IC50 | 34 uM | Not Available | BindingDB 50007468 |
References
- Kireev D, Wigle TJ, Norris-Drouin J, Herold JM, Janzen WP, Frye SV: Identification of non-peptide malignant brain tumor (MBT) repeat antagonists by virtual screening of commercially available compounds. J Med Chem. 2010 Nov 11;53(21):7625-31. doi: 10.1021/jm1007374. [20931980 ]
- General Function:
- Platelet activating factor receptor activity
- Specific Function:
- Receptor for platelet activating factor, a chemotactic phospholipid mediator that possesses potent inflammatory, smooth-muscle contractile and hypotensive activity. Seems to mediate its action via a G protein that activates a phosphatidylinositol-calcium second messenger system.
- Gene Name:
- PTAFR
- Uniprot ID:
- P25105
- Molecular Weight:
- 39203.075 Da
Binding/Activity Constants
| Type | Value | Assay Type | Assay Source |
|---|---|---|---|
| IC50 | >50 uM | Not Available | BindingDB 50007468 |
References
- Piwinski JJ, Wong JK, Green MJ, Ganguly AK, Billah MM, West RE Jr, Kreutner W: Dual antagonists of platelet activating factor and histamine. Identification of structural requirements for dual activity of N-Acyl-4-(5,6-dihydro-11H-benzo [5,6]cyclohepta-[1,2-b]pyridin-11-ylidene)piperidines. J Med Chem. 1991 Jan;34(1):457-61. [1671420 ]
- General Function:
- Monoamine transmembrane transporter activity
- Specific Function:
- Amine transporter. Terminates the action of dopamine by its high affinity sodium-dependent reuptake into presynaptic terminals.
- Gene Name:
- SLC6A3
- Uniprot ID:
- Q01959
- Molecular Weight:
- 68494.255 Da
References
- Tatsumi M, Groshan K, Blakely RD, Richelson E: Pharmacological profile of antidepressants and related compounds at human monoamine transporters. Eur J Pharmacol. 1997 Dec 11;340(2-3):249-58. [9537821 ]
- General Function:
- Norepinephrine:sodium symporter activity
- Specific Function:
- Amine transporter. Terminates the action of noradrenaline by its high affinity sodium-dependent reuptake into presynaptic terminals.
- Gene Name:
- SLC6A2
- Uniprot ID:
- P23975
- Molecular Weight:
- 69331.42 Da
References
- Tatsumi M, Groshan K, Blakely RD, Richelson E: Pharmacological profile of antidepressants and related compounds at human monoamine transporters. Eur J Pharmacol. 1997 Dec 11;340(2-3):249-58. [9537821 ]
- General Function:
- Serotonin:sodium symporter activity
- Specific Function:
- Serotonin transporter whose primary function in the central nervous system involves the regulation of serotonergic signaling via transport of serotonin molecules from the synaptic cleft back into the pre-synaptic terminal for re-utilization. Plays a key role in mediating regulation of the availability of serotonin to other receptors of serotonergic systems. Terminates the action of serotonin and recycles it in a sodium-dependent manner.
- Gene Name:
- SLC6A4
- Uniprot ID:
- P31645
- Molecular Weight:
- 70324.165 Da
References
- Tatsumi M, Groshan K, Blakely RD, Richelson E: Pharmacological profile of antidepressants and related compounds at human monoamine transporters. Eur J Pharmacol. 1997 Dec 11;340(2-3):249-58. [9537821 ]