| Record Information |
|---|
| Version | 2.0 |
|---|
| Creation Date | 2009-07-21 20:28:36 UTC |
|---|
| Update Date | 2014-12-24 20:25:55 UTC |
|---|
| Accession Number | T3D3015 |
|---|
| Identification |
|---|
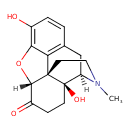
| Common Name | Oxymorphone |
|---|
| Class | Small Molecule |
|---|
| Description | An opioid analgesic with actions and uses similar to those of morphine, apart from an absence of cough suppressant activity. It is used in the treatment of moderate to severe pain, including pain in obstetrics. It may also be used as an adjunct to anesthesia. (From Martindale, The Extra Pharmacopoeia, 30th ed, p1092) |
|---|
| Compound Type | - Adjuvant
- Adjuvant, Anesthesia
- Amine
- Analgesic
- Analgesic, Opioid
- Drug
- Ether
- Metabolite
- Narcotic
- Opiate Agonist
- Organic Compound
- Synthetic Compound
|
|---|
| Chemical Structure | |
|---|
| Synonyms | | Synonym | | 14-Hydroxydihydromorphinone | | Dihydrohydroxymorphinone | | Dihydroxymorphinone | | EN3202 | | Numorphan | | Opana | | OPANA ER | | Oximorphonum | | Oxymorphine |
|
|---|
| Chemical Formula | C17H19NO4 |
|---|
| Average Molecular Mass | 301.337 g/mol |
|---|
| Monoisotopic Mass | 301.131 g/mol |
|---|
| CAS Registry Number | 76-41-5 |
|---|
| IUPAC Name | (1S,5R,13R,17S)-10,17-dihydroxy-4-methyl-12-oxa-4-azapentacyclo[9.6.1.0¹,¹³.0⁵,¹⁷.0⁷,¹⁸]octadeca-7(18),8,10-trien-14-one |
|---|
| Traditional Name | oxymorphone |
|---|
| SMILES | [H][C@@]12OC3=C(O)C=CC4=C3[C@@]11CCN(C)[C@]([H])(C4)[C@]1(O)CCC2=O |
|---|
| InChI Identifier | InChI=1S/C17H19NO4/c1-18-7-6-16-13-9-2-3-10(19)14(13)22-15(16)11(20)4-5-17(16,21)12(18)8-9/h2-3,12,15,19,21H,4-8H2,1H3/t12-,15+,16+,17-/m1/s1 |
|---|
| InChI Key | InChIKey=UQCNKQCJZOAFTQ-ISWURRPUSA-N |
|---|
| Chemical Taxonomy |
|---|
| Description | belongs to the class of organic compounds known as phenanthrenes and derivatives. These are polycyclic compounds containing a phenanthrene moiety, which is a tricyclic aromatic compound with three non-linearly fused benzene. |
|---|
| Kingdom | Organic compounds |
|---|
| Super Class | Benzenoids |
|---|
| Class | Phenanthrenes and derivatives |
|---|
| Sub Class | Not Available |
|---|
| Direct Parent | Phenanthrenes and derivatives |
|---|
| Alternative Parents | |
|---|
| Substituents | - Phenanthrene
- Isoquinolone
- Tetralin
- Coumaran
- 1-hydroxy-2-unsubstituted benzenoid
- Alkyl aryl ether
- Aralkylamine
- Piperidine
- Cyclic alcohol
- Tertiary alcohol
- 1,2-aminoalcohol
- Ketone
- Tertiary aliphatic amine
- Tertiary amine
- Ether
- Oxacycle
- Azacycle
- Organoheterocyclic compound
- Organonitrogen compound
- Hydrocarbon derivative
- Organic oxide
- Organic nitrogen compound
- Organopnictogen compound
- Carbonyl group
- Organooxygen compound
- Alcohol
- Organic oxygen compound
- Amine
- Aromatic heteropolycyclic compound
|
|---|
| Molecular Framework | Aromatic heteropolycyclic compounds |
|---|
| External Descriptors | |
|---|
| Biological Properties |
|---|
| Status | Detected and Not Quantified |
|---|
| Origin | Exogenous |
|---|
| Cellular Locations | - Cytoplasm
- Extracellular
- Membrane
|
|---|
| Biofluid Locations | Not Available |
|---|
| Tissue Locations | Not Available |
|---|
| Pathways | Not Available |
|---|
| Applications | Not Available |
|---|
| Biological Roles | Not Available |
|---|
| Chemical Roles | Not Available |
|---|
| Physical Properties |
|---|
| State | Solid |
|---|
| Appearance | White powder. |
|---|
| Experimental Properties | | Property | Value |
|---|
| Melting Point | 248-249°C | | Boiling Point | Not Available | | Solubility | 2.4E+004 mg/L | | LogP | 0.83 |
|
|---|
| Predicted Properties | |
|---|
| Spectra |
|---|
| Spectra | | Spectrum Type | Description | Splash Key | Deposition Date | View |
|---|
| Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | splash10-0a4l-9050000000-d955cbd739d3bad18756 | 2017-09-01 | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (2 TMS) - 70eV, Positive | splash10-00c0-6907600000-2e98fd21eb80cd8204b3 | 2017-10-06 | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | Not Available | 2021-10-12 | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | Not Available | 2021-10-12 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - 35V, Positive | splash10-0f89-0093000000-d20c48538ad6696fd83b | 2021-09-20 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-0f89-0095000000-3fa32e9f5773ea0cc992 | 2016-08-03 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-001i-0091000000-4c5b42673e83312d1305 | 2016-08-03 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-069u-3090000000-b1a88d98c02d968d1332 | 2016-08-03 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-0udi-0029000000-0f9dac6d2c5bb246b565 | 2016-08-03 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-0ue9-0097000000-2bab2cfd5aabe92e0abe | 2016-08-03 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-000x-2090000000-7d02b1358307dc0ce4d2 | 2016-08-03 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-0udi-0009000000-ab6f6c1d45a17ba966cd | 2021-10-11 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-0ue9-0049000000-68376f76c73135dac7e3 | 2021-10-11 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-0uk9-0093000000-3e6504da4a5c1b25ff70 | 2021-10-11 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-0udi-0009000000-867b26b8e9b8bd5f9273 | 2021-10-12 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-0udi-0009000000-867b26b8e9b8bd5f9273 | 2021-10-12 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-0udi-0097000000-9087b236d7dbdab76ba5 | 2021-10-12 | View Spectrum | | MS | Mass Spectrum (Electron Ionization) | splash10-0udl-7932000000-89141254d526ca629dac | 2014-09-20 | View Spectrum | | 1D NMR | 1H NMR Spectrum (1D, 100 MHz, D2O, predicted) | Not Available | 2021-09-25 | View Spectrum | | 1D NMR | 13C NMR Spectrum (1D, 100 MHz, D2O, predicted) | Not Available | 2021-09-25 | View Spectrum | | 1D NMR | 1H NMR Spectrum (1D, 1000 MHz, D2O, predicted) | Not Available | 2021-09-25 | View Spectrum | | 1D NMR | 13C NMR Spectrum (1D, 1000 MHz, D2O, predicted) | Not Available | 2021-09-25 | View Spectrum | | 1D NMR | 1H NMR Spectrum (1D, 200 MHz, D2O, predicted) | Not Available | 2021-09-25 | View Spectrum | | 1D NMR | 13C NMR Spectrum (1D, 200 MHz, D2O, predicted) | Not Available | 2021-09-25 | View Spectrum | | 1D NMR | 1H NMR Spectrum (1D, 300 MHz, D2O, predicted) | Not Available | 2021-09-25 | View Spectrum | | 1D NMR | 13C NMR Spectrum (1D, 300 MHz, D2O, predicted) | Not Available | 2021-09-25 | View Spectrum | | 1D NMR | 1H NMR Spectrum (1D, 400 MHz, D2O, predicted) | Not Available | 2021-09-25 | View Spectrum | | 1D NMR | 13C NMR Spectrum (1D, 400 MHz, D2O, predicted) | Not Available | 2021-09-25 | View Spectrum | | 1D NMR | 1H NMR Spectrum (1D, 500 MHz, D2O, predicted) | Not Available | 2021-09-25 | View Spectrum | | 1D NMR | 13C NMR Spectrum (1D, 500 MHz, D2O, predicted) | Not Available | 2021-09-25 | View Spectrum | | 1D NMR | 1H NMR Spectrum (1D, 600 MHz, D2O, predicted) | Not Available | 2021-09-25 | View Spectrum | | 1D NMR | 13C NMR Spectrum (1D, 600 MHz, D2O, predicted) | Not Available | 2021-09-25 | View Spectrum | | 1D NMR | 1H NMR Spectrum (1D, 700 MHz, D2O, predicted) | Not Available | 2021-09-25 | View Spectrum | | 1D NMR | 13C NMR Spectrum (1D, 700 MHz, D2O, predicted) | Not Available | 2021-09-25 | View Spectrum | | 1D NMR | 1H NMR Spectrum (1D, 800 MHz, D2O, predicted) | Not Available | 2021-09-25 | View Spectrum | | 1D NMR | 13C NMR Spectrum (1D, 800 MHz, D2O, predicted) | Not Available | 2021-09-25 | View Spectrum | | 1D NMR | 1H NMR Spectrum (1D, 900 MHz, D2O, predicted) | Not Available | 2021-09-25 | View Spectrum | | 1D NMR | 13C NMR Spectrum (1D, 900 MHz, D2O, predicted) | Not Available | 2021-09-25 | View Spectrum |
|
|---|
| Toxicity Profile |
|---|
| Route of Exposure | Enteral(rectal). |
|---|
| Mechanism of Toxicity | Oxymorphone interacts predominantly with the opioid mu-receptor. These mu-binding sites are discretely distributed in the human brain, with high densities in the posterior amygdala, hypothalamus, thalamus, nucleus caudatus, putamen, and certain cortical areas. They are also found on the terminal axons of primary afferents within laminae I and II (substantia gelatinosa) of the spinal cord and in the spinal nucleus of the trigeminal nerve. Also, it has been shown that oxymorphone binds to and inhibits GABA inhibitory interneurons via mu-receptors. These interneurons normally inhibit the descending pain inhibition pathway. So, without the inhibitory signals, pain modulation can proceed downstream. |
|---|
| Metabolism | Oxymorphone undergoes extensive hepatic metabolism in humans. After a 10 mg oral dose, 49% was excreted over a five-day period in the urine. Of this, 82% was excreted in the first 24 hours after administration. The recovered drug-related products contained the oxymorphone (1.9%), the conjugate of oxymorphone (44.1%), the 6(beta)-carbinol produced by 6-keto reduction of oxymorphone (0.3%), and the conjugates of 6(beta)-carbinol (2.6%) and 6(alpha)-carbinol (0.1%).
Route of Elimination: Oxymorphone is highly metabolized, principally in the liver, and undergoes reduction or conjugation with glucuronic acid to form both active and inactive products. Because oxymorphone is extensively metabolized, <1% of the administered dose is excreted unchanged in the urine.
Half Life: 1.3 (+/-0.7) hours |
|---|
| Toxicity Values | Intravenous mouse LD50 is 172 mg/kg.
|
|---|
| Lethal Dose | Not Available |
|---|
| Carcinogenicity (IARC Classification) | No indication of carcinogenicity to humans (not listed by IARC). |
|---|
| Uses/Sources | For the treatment of moderate-to-severe pain. |
|---|
| Minimum Risk Level | Not Available |
|---|
| Health Effects | Medical problems can include congested lungs, liver disease, tetanus, infection of the heart valves, skin abscesses, anemia and pneumonia. Death can occur from overdose. |
|---|
| Symptoms | Oxymorphone overdosage is characterized by respiratory depression, extreme somnolence progressing to stupor or coma, skeletal muscle flaccidity, cold and clammy skin, and sometimes bradycardia and hypotension. In a severe case of overdose, apnea, circulatory collapse, cardiac arrest, and death may occur. |
|---|
| Treatment | Primary attention should be given to the reestablishment of adequate respiratory exchange through provision of a patent airway and the institution of assisted or controlled ventilation. The opioid antagonist naloxone hydrochloride is a specific antidote against respiratory depression which may result from overdosage or unusual sensitivity to opioids including oxymorphone. Therefore, an appropriate dose of naloxone hydrochloride should be administered (usual initial adult dose 0.4 mg-2 mg) preferably by the intravenous route and simultaneously with efforts at respiratory resuscitation. Since the duration of action of oxymorphone may exceed that of the antagonist, the patient should be kept under continued surveillance and repeated doses of the antagonist should be administered as needed to maintain adequate respiration. Naloxone hydrochloride should not be administered in the absence of clinically significant respiratory or cardiovascular depression. In addition, it should be considered that the use of an opioid antagonist in patients physically dependent on opioids may precipitate an acute withdrawal syndrome that cannot be readily suppressed while the action of the antagonist persists. If respiratory depression is associated with muscular rigidity, administration of a neuromuscular blocking agent may be necessary to facilitate assisted or controlled ventilation. Muscular rigidity may also respond to opioid antagonist therapy. Oxygen, intravenous fluids, vasopressors and other supportive measures should be employed as indicated. (4) |
|---|
| Normal Concentrations |
|---|
| Not Available |
|---|
| Abnormal Concentrations |
|---|
| Not Available |
|---|
| External Links |
|---|
| DrugBank ID | DB01192 |
|---|
| HMDB ID | HMDB15323 |
|---|
| PubChem Compound ID | 5284604 |
|---|
| ChEMBL ID | CHEMBL963 |
|---|
| ChemSpider ID | 4447650 |
|---|
| KEGG ID | C08019 |
|---|
| UniProt ID | Not Available |
|---|
| OMIM ID | |
|---|
| ChEBI ID | 194484 |
|---|
| BioCyc ID | Not Available |
|---|
| CTD ID | Not Available |
|---|
| Stitch ID | Oxymorphone |
|---|
| PDB ID | Not Available |
|---|
| ACToR ID | Not Available |
|---|
| Wikipedia Link | Oxymorphone |
|---|
| References |
|---|
| Synthesis Reference | Bao-Shan Huang, Yansong Lu, Ben-Yi Ji, Aris P Christodoulou, “Preparation of oxymorphone from morphine.” U.S. Patent US5922876, issued May, 1992. |
|---|
| MSDS | Link |
|---|
| General References | - Martindale. The Extra Pharmacopoeia, 30th ed.
- Drugs.com [Link]
- Drugs.com [Link]
- RxList: The Internet Drug Index (2009). [Link]
|
|---|
| Gene Regulation |
|---|
| Up-Regulated Genes | Not Available |
|---|
| Down-Regulated Genes | Not Available |
|---|