| Record Information |
|---|
| Version | 2.0 |
|---|
| Creation Date | 2013-04-25 07:56:55 UTC |
|---|
| Update Date | 2014-12-24 20:26:34 UTC |
|---|
| Accession Number | T3D3941 |
|---|
| Identification |
|---|
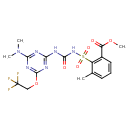
| Common Name | Triflusulfuron-methyl |
|---|
| Class | Small Molecule |
|---|
| Description | Triflusulfuron-methyl is a selective postemergence sulfonylurea herbicide for the control of annual and perennial broadleaf weeds and grasses in sugar beets (Beta vulgaris). Similar to other sulfonylureas, the site of action of triflusulfuron methyl is acetolactate synthase (ALS), an enzyme involved in branched-chain amino acid biosynthesis. |
|---|
| Compound Type | - Amide
- Amine
- Ester
- Ether
- Herbicide
- Organic Compound
- Organofluoride
- Pesticide
- Synthetic Compound
|
|---|
| Chemical Structure | |
|---|
| Synonyms | Not Available |
|---|
| Chemical Formula | C17H19F3N6O6S |
|---|
| Average Molecular Mass | 492.430 g/mol |
|---|
| Monoisotopic Mass | 492.104 g/mol |
|---|
| CAS Registry Number | 126535-15-7 |
|---|
| IUPAC Name | methyl 2-[({[4-(dimethylamino)-6-(2,2,2-trifluoroethoxy)-1,3,5-triazin-2-yl]carbamoyl}amino)sulfonyl]-3-methylbenzoate |
|---|
| Traditional Name | triflusulfuron-methyl |
|---|
| SMILES | COC(=O)C1=C(C(C)=CC=C1)S(=O)(=O)NC(=O)NC1=NC(=NC(OCC(F)(F)F)=N1)N(C)C |
|---|
| InChI Identifier | InChI=1S/C17H19F3N6O6S/c1-9-6-5-7-10(12(27)31-4)11(9)33(29,30)25-15(28)22-13-21-14(26(2)3)24-16(23-13)32-8-17(18,19)20/h5-7H,8H2,1-4H3,(H2,21,22,23,24,25,28) |
|---|
| InChI Key | InChIKey=IMEVJVISCHQJRM-UHFFFAOYSA-N |
|---|
| Chemical Taxonomy |
|---|
| Description | belongs to the class of organic compounds known as s-triazinyl-2-sulfonylureas. These are aromatic heterocyclic compounds containing a s-triazine ring which is substituted with a urea at the ring 2-position. |
|---|
| Kingdom | Organic compounds |
|---|
| Super Class | Organic nitrogen compounds |
|---|
| Class | Organonitrogen compounds |
|---|
| Sub Class | Sulfonylureas |
|---|
| Direct Parent | S-triazinyl-2-sulfonylureas |
|---|
| Alternative Parents | |
|---|
| Substituents | - S-triazinyl-2-sulfonylurea
- Benzenesulfonamide
- Benzoate ester
- Benzoic acid or derivatives
- Benzenesulfonyl group
- Alkoxy-s-triazine
- Benzoyl
- Dialkylarylamine
- Alkyl aryl ether
- Aminotriazine
- Amino-1,3,5-triazine
- Toluene
- N-aliphatic s-triazine
- 1,3,5-triazine
- Monocyclic benzene moiety
- Benzenoid
- Triazine
- Heteroaromatic compound
- Methyl ester
- Organic sulfonic acid or derivatives
- Organosulfonic acid or derivatives
- Sulfonyl
- Aminosulfonyl compound
- Carboxylic acid ester
- Carbonic acid derivative
- Carboxylic acid derivative
- Organoheterocyclic compound
- Ether
- Monocarboxylic acid or derivatives
- Azacycle
- Organic oxide
- Carbonyl group
- Alkyl fluoride
- Organopnictogen compound
- Organic oxygen compound
- Organohalogen compound
- Organofluoride
- Organooxygen compound
- Organosulfur compound
- Alkyl halide
- Hydrocarbon derivative
- Aromatic heteromonocyclic compound
|
|---|
| Molecular Framework | Aromatic heteromonocyclic compounds |
|---|
| External Descriptors | |
|---|
| Biological Properties |
|---|
| Status | Detected and Not Quantified |
|---|
| Origin | Exogenous |
|---|
| Cellular Locations | |
|---|
| Biofluid Locations | Not Available |
|---|
| Tissue Locations | Not Available |
|---|
| Pathways | Not Available |
|---|
| Applications | Not Available |
|---|
| Biological Roles | Not Available |
|---|
| Chemical Roles | Not Available |
|---|
| Physical Properties |
|---|
| State | Solid |
|---|
| Appearance | White powder. |
|---|
| Experimental Properties | | Property | Value |
|---|
| Melting Point | Not Available | | Boiling Point | Not Available | | Solubility | Not Available | | LogP | Not Available |
|
|---|
| Predicted Properties | |
|---|
| Spectra |
|---|
| Spectra | | Spectrum Type | Description | Splash Key | Deposition Date | View |
|---|
| Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | splash10-01p9-3290400000-c08e4438f00882703eaa | 2021-09-24 | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | Not Available | 2021-10-12 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - 15V, Negative | splash10-000i-0190000000-88fa8ce5acf0f5607b6c | 2021-09-20 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - 30V, Negative | splash10-000i-0490000000-6787ed88169b889b4c24 | 2021-09-20 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - 30V, Positive | splash10-03di-0090000000-9b4f94f85638316b6016 | 2021-09-20 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - 60V, Negative | splash10-03fs-0900000000-2621452d6a9f50bdd17f | 2021-09-20 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - 45V, Negative | splash10-0002-0910000000-6207aa8acf303fecd873 | 2021-09-20 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - 15V, Positive | splash10-03di-0090000000-0fd577deb3d0d3deaafd | 2021-09-20 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - 45V, Positive | splash10-03di-1090000000-6f0fcc20545568c6f124 | 2021-09-20 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - 60V, Positive | splash10-01ot-9140000000-8441dc974f092c459306 | 2021-09-20 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-000x-0060900000-f6c80327ef12225a78b6 | 2016-06-03 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-02ai-3090000000-cbe665bffa4aee6b2b80 | 2016-06-03 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-0g4i-7590000000-b86a77d31894a3d6d89e | 2016-06-03 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-000f-0091800000-5086fb002e0f72402381 | 2016-08-03 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-004i-0590200000-c2d4aa19224d8c6533ad | 2016-08-03 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-0fi0-5940000000-69df3e1037ac1e0f7fee | 2016-08-03 | View Spectrum |
|
|---|
| Toxicity Profile |
|---|
| Route of Exposure | Not Available |
|---|
| Mechanism of Toxicity | Not Available |
|---|
| Metabolism | Not Available |
|---|
| Toxicity Values | Not Available |
|---|
| Lethal Dose | Not Available |
|---|
| Carcinogenicity (IARC Classification) | No indication of carcinogenicity to humans (not listed by IARC). |
|---|
| Uses/Sources | This is a man-made compound that is used as a pesticide. |
|---|
| Minimum Risk Level | Not Available |
|---|
| Health Effects | Not Available |
|---|
| Symptoms | Not Available |
|---|
| Treatment | Not Available |
|---|
| Normal Concentrations |
|---|
| Not Available |
|---|
| Abnormal Concentrations |
|---|
| Not Available |
|---|
| External Links |
|---|
| DrugBank ID | Not Available |
|---|
| HMDB ID | Not Available |
|---|
| PubChem Compound ID | 92434 |
|---|
| ChEMBL ID | CHEMBL2143050 |
|---|
| ChemSpider ID | 83452 |
|---|
| KEGG ID | C18901 |
|---|
| UniProt ID | Not Available |
|---|
| OMIM ID | |
|---|
| ChEBI ID | Not Available |
|---|
| BioCyc ID | Not Available |
|---|
| CTD ID | Not Available |
|---|
| Stitch ID | Not Available |
|---|
| PDB ID | Not Available |
|---|
| ACToR ID | Not Available |
|---|
| Wikipedia Link | Not Available |
|---|
| References |
|---|
| Synthesis Reference | Not Available |
|---|
| MSDS | T3D3941.pdf |
|---|
| General References | Not Available |
|---|
| Gene Regulation |
|---|
| Up-Regulated Genes | Not Available |
|---|
| Down-Regulated Genes | Not Available |
|---|