| Record Information |
|---|
| Version | 2.0 |
|---|
| Creation Date | 2014-08-29 06:21:45 UTC |
|---|
| Update Date | 2014-12-24 20:26:46 UTC |
|---|
| Accession Number | T3D4319 |
|---|
| Identification |
|---|
| Common Name | L-Glutamine |
|---|
| Class | Small Molecule |
|---|
| Description | L-Glutamine (Gln) is one of the 20 amino acids encoded by the standard genetic code. Its side chain is an amide; it is formed by replacing a side-chain hydroxyl of glutamic acid with an amine functional group. glutamine is found in foods high in proteins, such as fish, red meat, beans, and dairy products. glutamine is a supplement that is used in weightlifting, bodybuilding, endurance and other sports, as well as by those who suffer from muscular cramps or pain particularly elderly people. The main use of glutamine within the diet of either group is as a means of replenishing the body's stores of amino acids that have been used during exercise or everyday activities. Studies which are looking into problems with excessive consumption of glutamine thus far have proved inconclusive. However, normal supplementation is healthy mainly because glutamine is supposed to be supplemented after prolonged periods of exercise (for example, a workout or exercise in which amino acids are required for use) and replenishes amino acid stores; this being the main reason glutamine is recommended during fasting or for people who suffer from physical trauma, immune deficiencies, or cancer. There is a significant body of evidence that links glutamine-enriched diets with intestinal effects; aiding maintenance of gut barrier function, intestinal cell proliferation and differentiation, as well as generally reducing septic morbidity and the symptoms of Irritable Bowel Syndrome. The reason for such cleansing properties is thought to stem from the fact that the intestinal extraction rate of glutamine is higher than that for other amino acids, and is therefore thought to be the most viable option when attempting to alleviate conditions relating to the gastrointestinal tract. These conditions were discovered after comparing plasma concentration within the gut between glutamine-enriched and non glutamine-enriched diets. However, even though glutamine is thought to have cleansing properties and effects, it is unknown to what extent glutamine has clinical benefits, due to the varied concentrations of glutamine in varieties of food. It is also known that glutamine has various effects in reducing healing time after operations. Hospital waiting times after abdominal surgery are reduced by providing parenteral nutrition regimens containing amounts of glutamine to patients. Clinical trials have revealed that patients on supplementation regimes containing glutamine have improved nitrogen balances, generation of cysteinyl-leukotrienes from polymorphonuclear neutrophil granulocytes and improved lymphocyte recovery and intestinal permeability (in postoperative patients) - in comparison to those who had no glutamine within their dietary regime; all without any side-effects. It is synthesized from glutamic acid and ammonia. It is the principal carrier of nitrogen in the body and is an important energy source for many cells. |
|---|
| Compound Type | - Amine
- Animal Toxin
- Dietary Supplement
- Drug
- Food Toxin
- Household Toxin
- Metabolite
- Micronutrient
- Natural Compound
- Non-Essential Amino Acid
- Nutraceutical
- Organic Compound
- Supplement
|
|---|
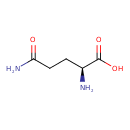
| Chemical Structure | |
|---|
| Synonyms | | Synonym | | (2S)-2,5-diamino-5-oxopentanoate | | (2S)-2,5-diamino-5-oxopentanoic acid | | (2S)-2-amino-4-carbamoylbutanoate | | (2S)-2-amino-4-carbamoylbutanoic acid | | (S)-2,5-Diamino-5-oxopentanoate | | (S)-2,5-Diamino-5-oxopentanoic acid | | 2-Aminoglutaramic acid | | Cebrogen | | Earthlink Science Glutamine Chews Chocolate | | gamma-Glutamine | | Glavamin | | Glumin | | Glutamic acid 5-amide | | Glutamic acid amide | | Glutamine | | Glutamine Express | | Glutamine Fuel Mega | | Glutamine fuel powder | | L-(+)-Glutamine | | L-2-Aminoglutaramic acid | | L-2-Aminoglutaramidic acid | | L-Glutamic acid 5-amide | | L-Glutamic acid gamma-amide | | L-glutamic acid γ-amide | | L-Glutamid | | L-Glutamide | | L-Glutamin | | L-Glutamine Power | | L-Glutaminsaeure-5-amid | | L-Glutaminsäure-5-amid | | Levoglutamid | | Levoglutamida | | Levoglutamide | | Levoglutamidum | | Levoglutamina | | NutreStore | | Polyglutamine | | Q | | Stimulina |
|
|---|
| Chemical Formula | C5H10N2O3 |
|---|
| Average Molecular Mass | 146.145 g/mol |
|---|
| Monoisotopic Mass | 146.069 g/mol |
|---|
| CAS Registry Number | 56-85-9 |
|---|
| IUPAC Name | (2S)-2-amino-4-carbamoylbutanoic acid |
|---|
| Traditional Name | L-glutamine |
|---|
| SMILES | [H][C@](N)(CCC(O)=N)C(O)=O |
|---|
| InChI Identifier | InChI=1S/C5H10N2O3/c6-3(5(9)10)1-2-4(7)8/h3H,1-2,6H2,(H2,7,8)(H,9,10)/t3-/m0/s1 |
|---|
| InChI Key | InChIKey=ZDXPYRJPNDTMRX-VKHMYHEASA-N |
|---|
| Chemical Taxonomy |
|---|
| Description | belongs to the class of organic compounds known as l-alpha-amino acids. These are alpha amino acids which have the L-configuration of the alpha-carbon atom. |
|---|
| Kingdom | Organic compounds |
|---|
| Super Class | Organic acids and derivatives |
|---|
| Class | Carboxylic acids and derivatives |
|---|
| Sub Class | Amino acids, peptides, and analogues |
|---|
| Direct Parent | L-alpha-amino acids |
|---|
| Alternative Parents | |
|---|
| Substituents | - L-alpha-amino acid
- Fatty acid
- Amino acid
- Carboximidic acid
- Carboximidic acid derivative
- Carboxylic acid
- Monocarboxylic acid or derivatives
- Amine
- Hydrocarbon derivative
- Primary amine
- Organooxygen compound
- Organonitrogen compound
- Organic oxide
- Primary aliphatic amine
- Organopnictogen compound
- Organic oxygen compound
- Organic nitrogen compound
- Carbonyl group
- Aliphatic acyclic compound
|
|---|
| Molecular Framework | Aliphatic acyclic compounds |
|---|
| External Descriptors | |
|---|
| Biological Properties |
|---|
| Status | Detected and Not Quantified |
|---|
| Origin | Endogenous |
|---|
| Cellular Locations | - Extracellular
- Membrane
- Mitochondria
|
|---|
| Biofluid Locations | Not Available |
|---|
| Tissue Locations | - Adipose Tissue
- Fibroblasts
- Gut
- Intestine
- Kidney
- Muscle
- Myelin
- Neuron
- Pancreas
- Placenta
- Prostate
- Skeletal Muscle
- Spleen
- Stratum Corneum
- Testes
|
|---|
| Pathways | |
|---|
| Applications | |
|---|
| Biological Roles | |
|---|
| Chemical Roles | |
|---|
| Physical Properties |
|---|
| State | Solid |
|---|
| Appearance | White powder. |
|---|
| Experimental Properties | | Property | Value |
|---|
| Melting Point | 185.5 dec°C | | Boiling Point | Not Available | | Solubility | 4.13E+004 mg/L (at 25°C) | | LogP | -3.64 |
|
|---|
| Predicted Properties | |
|---|
| Spectra |
|---|
| Spectra | | Spectrum Type | Description | Splash Key | Deposition Date | View |
|---|
| GC-MS | GC-MS Spectrum - GC-EI-TOF (Pegasus III TOF-MS system, Leco; GC 6890, Agilent Technologies) (3 TMS) | splash10-0a4i-0910000000-adb283bb40327f705680 | 2014-06-16 | View Spectrum | | GC-MS | GC-MS Spectrum - GC-EI-TOF (Pegasus III TOF-MS system, Leco; GC 6890, Agilent Technologies) (3 TMS) | splash10-0a4i-0910000000-134e80840320dadf6ad1 | 2014-06-16 | View Spectrum | | GC-MS | GC-MS Spectrum - GC-EI-TOF (Pegasus III TOF-MS system, Leco; GC 6890, Agilent Technologies) (3 TMS) | splash10-05fr-7910000000-89f87d4acc18299244f5 | 2014-06-16 | View Spectrum | | GC-MS | GC-MS Spectrum - GC-MS (2 TMS) | splash10-0a4i-0900000000-952a471cad5e5ead0a7e | 2014-06-16 | View Spectrum | | GC-MS | GC-MS Spectrum - GC-MS (4 TMS) | splash10-004i-1961000000-94183211889cfe72d4ff | 2014-06-16 | View Spectrum | | GC-MS | GC-MS Spectrum - GC-MS (3 TMS) | splash10-0a4i-1920000000-6505cd814f3707a0feba | 2014-06-16 | View Spectrum | | GC-MS | GC-MS Spectrum - GC-MS (4 TMS) | splash10-004i-0692000000-9788416fdefb051b9586 | 2014-06-16 | View Spectrum | | GC-MS | GC-MS Spectrum - GC-EI-TOF (Non-derivatized) | splash10-0a4i-0910000000-adb283bb40327f705680 | 2017-09-12 | View Spectrum | | GC-MS | GC-MS Spectrum - GC-EI-TOF (Non-derivatized) | splash10-0a4i-0910000000-134e80840320dadf6ad1 | 2017-09-12 | View Spectrum | | GC-MS | GC-MS Spectrum - GC-EI-QQ (Non-derivatized) | splash10-00dj-4921200000-5446333d9aa592da8a07 | 2017-09-12 | View Spectrum | | GC-MS | GC-MS Spectrum - GC-EI-TOF (Non-derivatized) | splash10-05fr-7910000000-89f87d4acc18299244f5 | 2017-09-12 | View Spectrum | | GC-MS | GC-MS Spectrum - GC-MS (Non-derivatized) | splash10-0a4i-0900000000-952a471cad5e5ead0a7e | 2017-09-12 | View Spectrum | | GC-MS | GC-MS Spectrum - GC-MS (Non-derivatized) | splash10-004i-1961000000-94183211889cfe72d4ff | 2017-09-12 | View Spectrum | | GC-MS | GC-MS Spectrum - GC-MS (Non-derivatized) | splash10-0a4i-1920000000-6505cd814f3707a0feba | 2017-09-12 | View Spectrum | | GC-MS | GC-MS Spectrum - GC-MS (Non-derivatized) | splash10-004i-0692000000-9788416fdefb051b9586 | 2017-09-12 | View Spectrum | | GC-MS | GC-MS Spectrum - GC-EI-TOF (Non-derivatized) | splash10-0a4i-0910000000-ce2b15fe45c57a30c6bb | 2017-09-12 | View Spectrum | | GC-MS | GC-MS Spectrum - GC-EI-TOF (Non-derivatized) | splash10-0ab9-1900000000-7be5eab1056d2a249ab5 | 2017-09-12 | View Spectrum | | GC-MS | GC-MS Spectrum - GC-EI-TOF (Non-derivatized) | splash10-0a4i-0900000000-5dc2d147bea250c7b80e | 2017-09-12 | View Spectrum | | GC-MS | GC-MS Spectrum - GC-EI-TOF (Non-derivatized) | splash10-0fb9-0921000000-5dcbab01982f543f7925 | 2017-09-12 | View Spectrum | | GC-MS | GC-MS Spectrum - GC-EI-TOF (Non-derivatized) | splash10-004j-0940000000-102c9a43b5153f06927f | 2017-09-12 | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | splash10-0006-9300000000-716b1947d48f42862cdf | 2016-09-22 | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (1 TMS) - 70eV, Positive | splash10-00dl-9610000000-bfd7aa529a6e7f62de43 | 2017-10-06 | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | Not Available | 2021-10-12 | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | Not Available | 2021-10-12 | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_1_2) - 70eV, Positive | Not Available | 2021-11-05 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - Quattro_QQQ 10V, Positive (Annotated) | splash10-0059-3900000000-c8f487d1a561e49327c0 | 2012-07-24 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - Quattro_QQQ 25V, Positive (Annotated) | splash10-001i-9000000000-eee04836e7577e11d0d0 | 2012-07-24 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - Quattro_QQQ 40V, Positive (Annotated) | splash10-0a59-9000000000-54ffbe5be8d7dbede389 | 2012-07-24 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-ITFT (LTQ Orbitrap XL, Thermo Scientfic) , Positive | splash10-0002-0900000000-79283391c985f8286677 | 2012-08-31 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-ITFT (LTQ Orbitrap XL, Thermo Scientfic) , Positive | splash10-001i-9000000000-6afbd5929b6e2371a371 | 2012-08-31 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-ITFT (LTQ Orbitrap XL, Thermo Scientfic) , Positive | splash10-001i-0900000000-d5bd83614703b048228e | 2012-08-31 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-ITFT (LTQ Orbitrap XL, Thermo Scientfic) , Positive | splash10-001i-0900000000-dc8def340a2655f9a399 | 2012-08-31 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-ITFT (LTQ Orbitrap XL, Thermo Scientfic) , Positive | splash10-0002-0900000000-47e3e94a4387c53e0bd6 | 2012-08-31 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-ITFT (LTQ Orbitrap XL, Thermo Scientfic) , Positive | splash10-001i-9000000000-c1acfa3750a90d81ce15 | 2012-08-31 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-ITFT (LTQ Orbitrap XL, Thermo Scientfic) , Positive | splash10-004i-0900000000-93f01bccbc98f18d4b07 | 2012-08-31 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-ITFT (LTQ Orbitrap XL, Thermo Scientfic) , Positive | splash10-001i-0900000000-ce03708ec9ce55cd8952 | 2012-08-31 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-ITFT (LTQ Orbitrap XL, Thermo Scientfic) , Negative | splash10-0002-0933201000-9339494cebedd9c39135 | 2012-08-31 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-ITFT (LTQ Orbitrap XL, Thermo Scientfic) , Negative | splash10-004i-0900000000-64561a4c3590d5994e37 | 2012-08-31 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-ITFT (LTQ Orbitrap XL, Thermo Scientfic) , Negative | splash10-0002-0900000000-fdcdd7591fb182d804e2 | 2012-08-31 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-ITFT (LTQ Orbitrap XL, Thermo Scientfic) , Negative | splash10-03dm-0030900000-66314ae704783630947a | 2012-08-31 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-ITFT (LTQ Orbitrap XL, Thermo Scientfic) , Negative | splash10-0002-0933100000-8798d2e46a415bb1029a | 2012-08-31 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-ITFT (LTQ Orbitrap XL, Thermo Scientfic) , Negative | splash10-004i-0900000000-dddea137e500a9b364cc | 2012-08-31 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-ITFT (LTQ Orbitrap XL, Thermo Scientfic) , Negative | splash10-0002-0900000000-a78f8e4b2d6b37917b21 | 2012-08-31 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-ITFT (LTQ Orbitrap XL, Thermo Scientfic) , Negative | splash10-0002-0920000000-8c6e1eb7de6f70911080 | 2012-08-31 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-QQ (API3000, Applied Biosystems) 10V, Negative | splash10-0002-0900000000-fa42fc4bcba608992804 | 2012-08-31 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-QQ (API3000, Applied Biosystems) 20V, Negative | splash10-004i-2900000000-074610880ea19a417f60 | 2012-08-31 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-QQ (API3000, Applied Biosystems) 30V, Negative | splash10-059x-9200000000-8b7dae54eb8325667bd7 | 2012-08-31 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-QQ (API3000, Applied Biosystems) 40V, Negative | splash10-0006-9000000000-e115d9215eea3f0c3629 | 2012-08-31 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-QQ (API3000, Applied Biosystems) 50V, Negative | splash10-0006-9000000000-55b1ac62eea66274b984 | 2012-08-31 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-QQ (API3000, Applied Biosystems) 10V, Positive | splash10-001j-0900000000-c9c71895033d66cb2bec | 2012-08-31 | View Spectrum | | MS | Mass Spectrum (Electron Ionization) | splash10-001i-9000000000-bc4e294b8a3fd8c5abce | 2014-09-20 | View Spectrum | | 1D NMR | 13C NMR Spectrum (1D, 125 MHz, H2O, experimental) | Not Available | 2012-12-04 | View Spectrum | | 1D NMR | 1H NMR Spectrum (1D, 500 MHz, H2O, experimental) | Not Available | 2012-12-04 | View Spectrum | | 1D NMR | 1H NMR Spectrum (1D, 400 MHz, D2O, experimental) | Not Available | 2014-09-20 | View Spectrum | | 1D NMR | 13C NMR Spectrum (1D, 22.5 MHz, D2O, experimental) | Not Available | 2014-09-23 | View Spectrum | | 1D NMR | 1H NMR Spectrum (1D, 100 MHz, D2O, predicted) | Not Available | 2021-09-29 | View Spectrum | | 1D NMR | 13C NMR Spectrum (1D, 100 MHz, D2O, predicted) | Not Available | 2021-09-29 | View Spectrum | | 1D NMR | 1H NMR Spectrum (1D, 1000 MHz, D2O, predicted) | Not Available | 2021-09-29 | View Spectrum | | 1D NMR | 13C NMR Spectrum (1D, 1000 MHz, D2O, predicted) | Not Available | 2021-09-29 | View Spectrum | | 1D NMR | 1H NMR Spectrum (1D, 200 MHz, D2O, predicted) | Not Available | 2021-09-29 | View Spectrum | | 1D NMR | 13C NMR Spectrum (1D, 200 MHz, D2O, predicted) | Not Available | 2021-09-29 | View Spectrum | | 1D NMR | 1H NMR Spectrum (1D, 300 MHz, D2O, predicted) | Not Available | 2021-09-29 | View Spectrum | | 1D NMR | 13C NMR Spectrum (1D, 300 MHz, D2O, predicted) | Not Available | 2021-09-29 | View Spectrum | | 1D NMR | 1H NMR Spectrum (1D, 400 MHz, D2O, predicted) | Not Available | 2021-09-29 | View Spectrum | | 1D NMR | 13C NMR Spectrum (1D, 400 MHz, D2O, predicted) | Not Available | 2021-09-29 | View Spectrum | | 1D NMR | 1H NMR Spectrum (1D, 500 MHz, D2O, predicted) | Not Available | 2021-09-29 | View Spectrum | | 1D NMR | 13C NMR Spectrum (1D, 500 MHz, D2O, predicted) | Not Available | 2021-09-29 | View Spectrum | | 1D NMR | 1H NMR Spectrum (1D, 600 MHz, D2O, predicted) | Not Available | 2021-09-29 | View Spectrum | | 1D NMR | 13C NMR Spectrum (1D, 600 MHz, D2O, predicted) | Not Available | 2021-09-29 | View Spectrum | | 1D NMR | 1H NMR Spectrum (1D, 700 MHz, D2O, predicted) | Not Available | 2021-09-29 | View Spectrum | | 1D NMR | 13C NMR Spectrum (1D, 700 MHz, D2O, predicted) | Not Available | 2021-09-29 | View Spectrum | | 1D NMR | 1H NMR Spectrum (1D, 800 MHz, D2O, predicted) | Not Available | 2021-09-29 | View Spectrum | | 1D NMR | 13C NMR Spectrum (1D, 800 MHz, D2O, predicted) | Not Available | 2021-09-29 | View Spectrum | | 1D NMR | 1H NMR Spectrum (1D, 900 MHz, D2O, predicted) | Not Available | 2021-09-29 | View Spectrum | | 1D NMR | 13C NMR Spectrum (1D, 900 MHz, D2O, predicted) | Not Available | 2021-09-29 | View Spectrum | | 1D NMR | 13C NMR Spectrum (1D, 400 MHz, H2O, experimental) | Not Available | 2021-10-10 | View Spectrum | | 2D NMR | [1H, 13C]-HSQC NMR Spectrum (2D, 600 MHz, H2O, experimental) | Not Available | 2012-12-05 | View Spectrum | | 2D NMR | [1H, 1H]-TOCSY 2D NMR Spectrum (experimental) | Not Available | 2021-10-14 | View Spectrum |
|
|---|
| Toxicity Profile |
|---|
| Route of Exposure | Absorption is efficient and occurs by an active transport mechanism |
|---|
| Mechanism of Toxicity | Supplemental L-glutamine's possible immunomodulatory role may be accounted for in a number of ways. L-glutamine appears to play a major role in protecting the integrity of the gastrointestinal tract and, in particular, the large intestine. During catabolic states, the integrity of the intestinal mucosa may be compromised with consequent increased intestinal permeability and translocation of Gram-negative bacteria from the large intestine into the body. The demand for L-glutamine by the intestine, as well as by cells such as lymphocytes, appears to be much greater than that supplied by skeletal muscle, the major storage tissue for L-glutamine. L-glutamine is the preferred respiratory fuel for enterocytes, colonocytes and lymphocytes. Therefore, supplying supplemental L-glutamine under these conditions may do a number of things. For one, it may reverse the catabolic state by sparing skeletal muscle L-glutamine. It also may inhibit translocation of Gram-negative bacteria from the large intestine. L-glutamine helps maintain secretory IgA, which functions primarily by preventing the attachment of bacteria to mucosal cells. L-glutamine appears to be required to support the proliferation of mitogen-stimulated lymphocytes, as well as the production of interleukin-2 (IL-2) and interferon-gamma (IFN-gamma). It is also required for the maintenance of lymphokine-activated killer cells (LAK). L-glutamine can enhance phagocytosis by neutrophils and monocytes. It can lead to an increased synthesis of glutathione in the intestine, which may also play a role in maintaining the integrity of the intestinal mucosa by ameliorating oxidative stress. The exact mechanism of the possible immunomodulatory action of supplemental L-glutamine, however, remains unclear. It is conceivable that the major effect of L-glutamine occurs at the level of the intestine. Perhaps enteral L-glutamine acts directly on intestine-associated lymphoid tissue and stimulates overall immune function by that mechanism, without passing beyond the splanchnic bed. |
|---|
| Metabolism | Enterocytes, Hepatic |
|---|
| Toxicity Values | Not Available |
|---|
| Lethal Dose | Not Available |
|---|
| Carcinogenicity (IARC Classification) | No indication of carcinogenicity to humans (not listed by IARC). |
|---|
| Uses/Sources | Used for nutritional supplementation, also for treating dietary shortage or imbalance. |
|---|
| Minimum Risk Level | Not Available |
|---|
| Health Effects | Not Available |
|---|
| Symptoms | Not Available |
|---|
| Treatment | Not Available |
|---|
| Normal Concentrations |
|---|
| Not Available |
|---|
| Abnormal Concentrations |
|---|
| Not Available |
|---|
| External Links |
|---|
| DrugBank ID | DB00130 |
|---|
| HMDB ID | HMDB00641 |
|---|
| PubChem Compound ID | 5961 |
|---|
| ChEMBL ID | CHEMBL930 |
|---|
| ChemSpider ID | 5746 |
|---|
| KEGG ID | C00064 |
|---|
| UniProt ID | Not Available |
|---|
| OMIM ID | |
|---|
| ChEBI ID | 18050 |
|---|
| BioCyc ID | GLN |
|---|
| CTD ID | Not Available |
|---|
| Stitch ID | Not Available |
|---|
| PDB ID | GLN |
|---|
| ACToR ID | Not Available |
|---|
| Wikipedia Link | L-Glutamine |
|---|
| References |
|---|
| Synthesis Reference | Stephen Paul, “Novel preparation of fiber, L-glutamine and a soy derivative for the purpose of enhancement of isoflavone bioavailability.” U.S. Patent US20020076455, issued June 20, 2002. |
|---|
| MSDS | Link |
|---|
| General References | - Boza JJ, Dangin M, Moennoz D, Montigon F, Vuichoud J, Jarret A, Pouteau E, Gremaud G, Oguey-Araymon S, Courtois D, Woupeyi A, Finot PA, Ballevre O: Free and protein-bound glutamine have identical splanchnic extraction in healthy human volunteers. Am J Physiol Gastrointest Liver Physiol. 2001 Jul;281(1):G267-74. [11408280 ]
- McAnena OJ, Moore FA, Moore EE, Jones TN, Parsons P: Selective uptake of glutamine in the gastrointestinal tract: confirmation in a human study. Br J Surg. 1991 Apr;78(4):480-2. [1903318 ]
- Morlion BJ, Stehle P, Wachtler P, Siedhoff HP, Koller M, Konig W, Furst P, Puchstein C: Total parenteral nutrition with glutamine dipeptide after major abdominal surgery: a randomized, double-blind, controlled study. Ann Surg. 1998 Feb;227(2):302-8. [9488531 ]
- Jian ZM, Cao JD, Zhu XG, Zhao WX, Yu JC, Ma EL, Wang XR, Zhu MW, Shu H, Liu YW: The impact of alanyl-glutamine on clinical safety, nitrogen balance, intestinal permeability, and clinical outcome in postoperative patients: a randomized, double-blind, controlled study of 120 patients. JPEN J Parenter Enteral Nutr. 1999 Sep-Oct;23(5 Suppl):S62-6. [10483898 ]
- Szebenyi G, Morfini GA, Babcock A, Gould M, Selkoe K, Stenoien DL, Young M, Faber PW, MacDonald ME, McPhaul MJ, Brady ST: Neuropathogenic forms of huntingtin and androgen receptor inhibit fast axonal transport. Neuron. 2003 Sep 25;40(1):41-52. [14527432 ]
- Wada A, Yoshida R, Oda K, Fukuba E, Uchida N, Kitagaki H: Acute encephalopathy associated with intravenous immunoglobulin therapy. AJNR Am J Neuroradiol. 2005 Oct;26(9):2311-5. [16219838 ]
- Pennisi P, Gavrilova O, Setser-Portas J, Jou W, Santopietro S, Clemmons D, Yakar S, LeRoith D: Recombinant human insulin-like growth factor-I treatment inhibits gluconeogenesis in a transgenic mouse model of type 2 diabetes mellitus. Endocrinology. 2006 Jun;147(6):2619-30. Epub 2006 Mar 2. [16513827 ]
- Peng CT, Wu KH, Lan SJ, Tsai JJ, Tsai FJ, Tsai CH: Amino acid concentrations in cerebrospinal fluid in children with acute lymphoblastic leukemia undergoing chemotherapy. Eur J Cancer. 2005 May;41(8):1158-63. Epub 2005 Apr 14. [15911239 ]
- Cynober LA: Plasma amino acid levels with a note on membrane transport: characteristics, regulation, and metabolic significance. Nutrition. 2002 Sep;18(9):761-6. [12297216 ]
- Rainesalo S, Keranen T, Palmio J, Peltola J, Oja SS, Saransaari P: Plasma and cerebrospinal fluid amino acids in epileptic patients. Neurochem Res. 2004 Jan;29(1):319-24. [14992292 ]
- Subramanian A, Gupta A, Saxena S, Gupta A, Kumar R, Nigam A, Kumar R, Mandal SK, Roy R: Proton MR CSF analysis and a new software as predictors for the differentiation of meningitis in children. NMR Biomed. 2005 Jun;18(4):213-25. [15627241 ]
- Redjems-Bennani N, Jeandel C, Lefebvre E, Blain H, Vidailhet M, Gueant JL: Abnormal substrate levels that depend upon mitochondrial function in cerebrospinal fluid from Alzheimer patients. Gerontology. 1998;44(5):300-4. [9693263 ]
- Avila J, Barbaro B, Gangemi A, Romagnoli T, Kuechle J, Hansen M, Shapiro J, Testa G, Sankary H, Benedetti E, Lakey J, Oberholzer J: Intra-ductal glutamine administration reduces oxidative injury during human pancreatic islet isolation. Am J Transplant. 2005 Dec;5(12):2830-7. [16302995 ]
- Cooper AJ: Ammonia metabolism in normal and portacaval-shunted rats. Adv Exp Med Biol. 1990;272:23-46. [2103690 ]
- Melis GC, Boelens PG, van der Sijp JR, Popovici T, De Bandt JP, Cynober L, van Leeuwen PA: The feeding route (enteral or parenteral) affects the plasma response of the dipetide Ala-Gln and the amino acids glutamine, citrulline and arginine, with the administration of Ala-Gln in preoperative patients. Br J Nutr. 2005 Jul;94(1):19-26. [16115328 ]
- Choudry HA, Pan M, Karinch AM, Souba WW: Branched-chain amino acid-enriched nutritional support in surgical and cancer patients. J Nutr. 2006 Jan;136(1 Suppl):314S-8S. [16365105 ]
- Commodari F, Arnold DL, Sanctuary BC, Shoubridge EA: 1H NMR characterization of normal human cerebrospinal fluid and the detection of methylmalonic acid in a vitamin B12 deficient patient. NMR Biomed. 1991 Aug;4(4):192-200. [1931558 ]
- Frayn KN, Khan K, Coppack SW, Elia M: Amino acid metabolism in human subcutaneous adipose tissue in vivo. Clin Sci (Lond). 1991 May;80(5):471-4. [1851687 ]
- Silwood CJ, Lynch E, Claxson AW, Grootveld MC: 1H and (13)C NMR spectroscopic analysis of human saliva. J Dent Res. 2002 Jun;81(6):422-7. [12097436 ]
- Coeffier M, Miralles-Barrachina O, Le Pessot F, Lalaude O, Daveau M, Lavoinne A, Lerebours E, Dechelotte P: Influence of glutamine on cytokine production by human gut in vitro. Cytokine. 2001 Feb 7;13(3):148-54. [11161457 ]
- Nicholson JK, O'Flynn MP, Sadler PJ, Macleod AF, Juul SM, Sonksen PH: Proton-nuclear-magnetic-resonance studies of serum, plasma and urine from fasting normal and diabetic subjects. Biochem J. 1984 Jan 15;217(2):365-75. [6696735 ]
- Rutten EP, Engelen MP, Wouters EF, Schols AM, Deutz NE: Metabolic effects of glutamine and glutamate ingestion in healthy subjects and in persons with chronic obstructive pulmonary disease. Am J Clin Nutr. 2006 Jan;83(1):115-23. [16400059 ]
- van der Hulst RR, von Meyenfeldt MF, Deutz NE, Soeters PB: Glutamine extraction by the gut is reduced in depleted [corrected] patients with gastrointestinal cancer. Ann Surg. 1997 Jan;225(1):112-21. [8998127 ]
- Hagenfeldt L, Bjerkenstedt L, Edman G, Sedvall G, Wiesel FA: Amino acids in plasma and CSF and monoamine metabolites in CSF: interrelationship in healthy subjects. J Neurochem. 1984 Mar;42(3):833-7. [6198473 ]
- Sreekumar A, Poisson LM, Rajendiran TM, Khan AP, Cao Q, Yu J, Laxman B, Mehra R, Lonigro RJ, Li Y, Nyati MK, Ahsan A, Kalyana-Sundaram S, Han B, Cao X, Byun J, Omenn GS, Ghosh D, Pennathur S, Alexander DC, Berger A, Shuster JR, Wei JT, Varambally S, Beecher C, Chinnaiyan AM: Metabolomic profiles delineate potential role for sarcosine in prostate cancer progression. Nature. 2009 Feb 12;457(7231):910-4. doi: 10.1038/nature07762. [19212411 ]
|
|---|
| Gene Regulation |
|---|
| Up-Regulated Genes | Not Available |
|---|
| Down-Regulated Genes | Not Available |
|---|