| Record Information |
|---|
| Version | 2.0 |
|---|
| Creation Date | 2014-08-30 21:04:00 UTC |
|---|
| Update Date | 2014-12-24 20:26:52 UTC |
|---|
| Accession Number | T3D4557 |
|---|
| Identification |
|---|
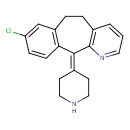
| Common Name | Desloratadine |
|---|
| Class | Small Molecule |
|---|
| Description | Desloratadine is a second generation, tricyclic antihistamine that which has a selective and peripheral H1-antagonist action. It is the active descarboethoxy metabolite of loratidine (a second generation histamine). Desloratidine has a long-lasting effect and does not cause drowsiness because it does not readily enter the central nervous system. |
|---|
| Compound Type | - Amine
- Cholinergic Antagonist
- Drug
- Histamine Antagonist
- Histamine H1 Antagonist, Non-Sedating
- Organic Compound
- Organochloride
- Synthetic Compound
|
|---|
| Chemical Structure | |
|---|
| Synonyms | | Synonym | | 8-Chloro-11-piperidin-4-ylidene-6,11-dihydro-5H-benzo[5,6]cyclohepta[1,2-b]pyridine | | 8-chloro-6,11-dihydro-11-(4-Piperidinylidene)-5H-benzo(5,6)cyclohepta(1,2-b)pyridine | | Aerius | | Claramax | | Clarinex | | Descarboethoxyloratadine | | DESLORATADINE | | NeoClarityn |
|
|---|
| Chemical Formula | C19H19ClN2 |
|---|
| Average Molecular Mass | 310.821 g/mol |
|---|
| Monoisotopic Mass | 310.124 g/mol |
|---|
| CAS Registry Number | 100643-71-8 |
|---|
| IUPAC Name | 13-chloro-2-(piperidin-4-ylidene)-4-azatricyclo[9.4.0.0³,⁸]pentadeca-1(11),3(8),4,6,12,14-hexaene |
|---|
| Traditional Name | clarinex |
|---|
| SMILES | ClC1=CC=C2C(CCC3=C(N=CC=C3)C2=C2CCNCC2)=C1 |
|---|
| InChI Identifier | InChI=1S/C19H19ClN2/c20-16-5-6-17-15(12-16)4-3-14-2-1-9-22-19(14)18(17)13-7-10-21-11-8-13/h1-2,5-6,9,12,21H,3-4,7-8,10-11H2 |
|---|
| InChI Key | InChIKey=JAUOIFJMECXRGI-UHFFFAOYSA-N |
|---|
| Chemical Taxonomy |
|---|
| Description | belongs to the class of organic compounds known as benzocycloheptapyridines. These are aromatic compounds containing a benzene ring and a pyridine ring fused to a seven membered carbocycle. |
|---|
| Kingdom | Organic compounds |
|---|
| Super Class | Organoheterocyclic compounds |
|---|
| Class | Benzocycloheptapyridines |
|---|
| Sub Class | Not Available |
|---|
| Direct Parent | Benzocycloheptapyridines |
|---|
| Alternative Parents | |
|---|
| Substituents | - Benzocycloheptapyridine
- Aryl chloride
- Aryl halide
- Piperidine
- Pyridine
- Benzenoid
- Heteroaromatic compound
- Secondary amine
- Secondary aliphatic amine
- Azacycle
- Amine
- Organonitrogen compound
- Organochloride
- Organohalogen compound
- Organic nitrogen compound
- Hydrocarbon derivative
- Organopnictogen compound
- Aromatic heteropolycyclic compound
|
|---|
| Molecular Framework | Aromatic heteropolycyclic compounds |
|---|
| External Descriptors | |
|---|
| Biological Properties |
|---|
| Status | Detected and Not Quantified |
|---|
| Origin | Exogenous |
|---|
| Cellular Locations | |
|---|
| Biofluid Locations | Not Available |
|---|
| Tissue Locations | Not Available |
|---|
| Pathways | Not Available |
|---|
| Applications | |
|---|
| Biological Roles | |
|---|
| Chemical Roles | Not Available |
|---|
| Physical Properties |
|---|
| State | Solid |
|---|
| Appearance | White powder. |
|---|
| Experimental Properties | | Property | Value |
|---|
| Melting Point | Not Available | | Boiling Point | Not Available | | Solubility | Not Available | | LogP | 3.2 |
|
|---|
| Predicted Properties | |
|---|
| Spectra |
|---|
| Spectra | | Spectrum Type | Description | Splash Key | Deposition Date | View |
|---|
| Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | splash10-00pm-1090000000-22109c83f6d7364ec27f | 2017-09-01 | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | Not Available | 2021-10-12 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-qTof , Positive | splash10-08fr-0095000000-2816aba50d0b94d568cf | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-qTof , Positive | splash10-08fr-0095000000-5b3f7555b72461661fa1 | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - , positive | splash10-0a4i-0940000000-d20368a648c20c37e1a0 | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - , positive | splash10-08fr-0095000000-2816aba50d0b94d568cf | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - , positive | splash10-0a4i-0940000000-9a0f5f4d81f4c42c85bb | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - , positive | splash10-08fr-0095000000-5b3f7555b72461661fa1 | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-QFT , positive | splash10-0bt9-0094000000-536b5a1d05f7479c2b04 | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - 10V, Positive | splash10-03di-0009000000-4cdd75cfb5b6a95978ee | 2021-09-20 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - 35V, Positive | splash10-0a4i-0091000000-702ea1b0afc5802588fe | 2021-09-20 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - 20V, Positive | splash10-08fr-0094000000-5d18c689b0d8db209b61 | 2021-09-20 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - 30V, Positive | splash10-0a4i-0090000000-0a46374188b55ae72bbb | 2021-09-20 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - 35V, Positive | splash10-0bt9-0094000000-c7895278fdf48ea14ae7 | 2021-09-20 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - 40V, Positive | splash10-0a4i-0090000000-8a8858c86845b74e96f3 | 2021-09-20 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - 50V, Positive | splash10-0a4l-0090000000-fe6da9796038e735965f | 2021-09-20 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-03di-0019000000-1fb53cf6a1a948b66616 | 2016-08-03 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-03di-5098000000-08255511bea5c886b910 | 2016-08-03 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-001i-9040000000-e8c991ebd37a0b0c2544 | 2016-08-03 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-0a4i-0009000000-3a9994f7ccd438482461 | 2016-08-03 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-0a4i-1029000000-5f479948c9e9c3620d41 | 2016-08-03 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-007p-5190000000-6fbaf13ffa17c703d90d | 2016-08-03 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-03di-0009000000-c678c7bde266b8b7c561 | 2021-10-11 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-03di-0009000000-40e78516a505b8e5b66e | 2021-10-11 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-0a6s-0190000000-d26af4a5ad57450875ac | 2021-10-11 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-0a4i-0009000000-3adde6a718d7e97498aa | 2021-10-11 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-053r-9005000000-07072a7bbfb51b942054 | 2021-10-11 | View Spectrum |
|
|---|
| Toxicity Profile |
|---|
| Route of Exposure | Not Available |
|---|
| Mechanism of Toxicity | Like other H1-blockers, Desloratadine competes with free histamine for binding at H1-receptors in the GI tract, uterus, large blood vessels, and bronchial smooth muscle. This blocks the action of endogenous histamine, which subsequently leads to temporary relief of the negative symptoms (eg. nasal congestion, watery eyes) brought on by histamine. |
|---|
| Metabolism |
Route of Elimination: Desloratadine (a major metabolite of loratadine) is extensively metabolized to 3-hydroxydesloratadine, an active metabolite, which is subsequently glucuronidated. Approximately 87% of a 14C-desloratadine dose was equally recovered in urine and feces.
Half Life: 50 hours |
|---|
| Toxicity Values | Not Available |
|---|
| Lethal Dose | Not Available |
|---|
| Carcinogenicity (IARC Classification) | No indication of carcinogenicity to humans (not listed by IARC). |
|---|
| Uses/Sources | For the relief of symptoms of seasonal allergic rhinitis, perennial (non-seasonal) allergic rhinitis. Desloratidine is also used for the sympomatic treatment of pruritus and urticaria (hives) associated with chronic idiopathic urticaria. |
|---|
| Minimum Risk Level | Not Available |
|---|
| Health Effects | Not Available |
|---|
| Symptoms | Not Available |
|---|
| Treatment | Not Available |
|---|
| Normal Concentrations |
|---|
| Not Available |
|---|
| Abnormal Concentrations |
|---|
| Not Available |
|---|
| External Links |
|---|
| DrugBank ID | DB00967 |
|---|
| HMDB ID | Not Available |
|---|
| PubChem Compound ID | 124087 |
|---|
| ChEMBL ID | CHEMBL1172 |
|---|
| ChemSpider ID | 110575 |
|---|
| KEGG ID | Not Available |
|---|
| UniProt ID | Not Available |
|---|
| OMIM ID | |
|---|
| ChEBI ID | 291342 |
|---|
| BioCyc ID | Not Available |
|---|
| CTD ID | Not Available |
|---|
| Stitch ID | Not Available |
|---|
| PDB ID | Not Available |
|---|
| ACToR ID | Not Available |
|---|
| Wikipedia Link | Desloratadine |
|---|
| References |
|---|
| Synthesis Reference | Zoltan Toth, “Processes for preparation of polymorphic forms of desloratadine.” U.S. Patent US20040242619, issued December 02, 2004. |
|---|
| MSDS | T3D4557.pdf |
|---|
| General References | - Mann RD, Pearce GL, Dunn N, Shakir S: Sedation with "non-sedating" antihistamines: four prescription-event monitoring studies in general practice. BMJ. 2000 Apr 29;320(7243):1184-6. [10784544 ]
- Glass DJ, Harper AS: Assessing satisfaction with desloratadine and fexofenadine in allergy patients who report dissatisfaction with loratadine. BMC Fam Pract. 2003 Aug 13;4:10. Epub 2003 Aug 13. [12917016 ]
- See S: Desloratadine for allergic rhinitis. Am Fam Physician. 2003 Nov 15;68(10):2015-6. [14655812 ]
- Devillier P, Roche N, Faisy C: Clinical pharmacokinetics and pharmacodynamics of desloratadine, fexofenadine and levocetirizine : a comparative review. Clin Pharmacokinet. 2008;47(4):217-30. [18336052 ]
- Bachert C: A review of the efficacy of desloratadine, fexofenadine, and levocetirizine in the treatment of nasal congestion in patients with allergic rhinitis. Clin Ther. 2009 May;31(5):921-44. doi: 10.1016/j.clinthera.2009.05.017. [19539095 ]
- DuBuske L: Desloratadine for chronic idiopathic urticaria: a review of clinical efficacy. Am J Clin Dermatol. 2007;8(5):271-83. [17902729 ]
- Bachert C, Maurer M: Safety and efficacy of desloratadine in subjects with seasonal allergic rhinitis or chronic urticaria: results of four postmarketing surveillance studies. Clin Drug Investig. 2010;30(2):109-22. doi: 10.2165/11530930-000000000-00000. [20067329 ]
- Simons FE, Prenner BM, Finn A Jr: Efficacy and safety of desloratadine in the treatment of perennial allergic rhinitis. J Allergy Clin Immunol. 2003 Mar;111(3):617-22. [12642846 ]
|
|---|
| Gene Regulation |
|---|
| Up-Regulated Genes | Not Available |
|---|
| Down-Regulated Genes | Not Available |
|---|