Podofilox (T3D4617)
| Record Information | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Version | 2.0 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Creation Date | 2014-09-05 17:14:40 UTC | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Update Date | 2014-12-24 20:26:54 UTC | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Accession Number | T3D4617 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Identification | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Common Name | Podofilox | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Class | Small Molecule | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Description | A lignan (lignans) found in podophyllin resin from the roots of podophyllum plants. It is a potent spindle poison, toxic if taken internally, and has been used as a cathartic. It is very irritating to skin and mucous membranes, has keratolytic actions, has been used to treat warts and keratoses, and may have antineoplastic properties, as do some of its congeners and derivatives. | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Compound Type |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
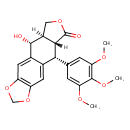
| Chemical Structure | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Synonyms |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Chemical Formula | C22H22O8 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Average Molecular Mass | 414.405 g/mol | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Monoisotopic Mass | 414.131 g/mol | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| CAS Registry Number | 477-47-4 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| IUPAC Name | (10R,11R,15R,16R)-16-hydroxy-10-(3,4,5-trimethoxyphenyl)-4,6,13-trioxatetracyclo[7.7.0.0³,⁷.0¹¹,¹⁵]hexadeca-1,3(7),8-trien-12-one | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Traditional Name | condylox | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| SMILES | [H][C@]12COC(=O)[C@]1([H])[C@]([H])(C1=CC(OC)=C(OC)C(OC)=C1)C1=CC3=C(OCO3)C=C1[C@]2([H])O | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| InChI Identifier | InChI=1S/C22H22O8/c1-25-16-4-10(5-17(26-2)21(16)27-3)18-11-6-14-15(30-9-29-14)7-12(11)20(23)13-8-28-22(24)19(13)18/h4-7,13,18-20,23H,8-9H2,1-3H3/t13-,18+,19-,20-/m0/s1 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| InChI Key | InChIKey=YJGVMLPVUAXIQN-XVVDYKMHSA-N | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Chemical Taxonomy | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Description | belongs to the class of organic compounds known as podophyllotoxins. These are tetralin lignans in which the benzene moiety of the tetralin skeleton is fused to a 1,3-dioxolane and the cyclohexane is fused to a butyrolactone (pyrrolidin-2-one). | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Kingdom | Organic compounds | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Super Class | Lignans, neolignans and related compounds | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Class | Lignan lactones | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Sub Class | Podophyllotoxins | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Direct Parent | Podophyllotoxins | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Alternative Parents |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Substituents |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Molecular Framework | Aromatic heteropolycyclic compounds | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| External Descriptors |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Biological Properties | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Status | Detected and Not Quantified | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Origin | Exogenous | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Cellular Locations |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Biofluid Locations | Not Available | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Tissue Locations | Not Available | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Pathways | Not Available | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Applications | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Biological Roles | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Chemical Roles | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Physical Properties | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| State | Solid | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Appearance | White powder. | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Experimental Properties |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Predicted Properties |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Spectra | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Spectra |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Toxicity Profile | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Route of Exposure | Topical application of 0.05 mL of 0.5% podofilox solution to external genitalia did not result in detectable serum levels. Applications of 0.1 to 1.5 mL resulted in peak serum levels of 1 to 17 ng/mL one to two hours after application. | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Mechanism of Toxicity | Etoposide, a semisynthetic derivative of podofilox, induces DNA breakage through its inhibition of topoisomerase II. The drug is most active in the late S and early G2 phases of the cell cycle. Teniposide is an analog with very similar pharmacologic characteristics. Podofilox derivatives display binding activity to the enzyme topoisomerase II during the late S and early G2 stage. For instance, etoposide binds and stabilizes the temporary break caused by the enzyme, disrupts the reparation of the break through which the double-stranded DNA passes, and consequently stops DNA unwinding and replication. Mutants resistant to either podofilox, or to its topoisomerase II inhibitory derivatives such as etoposide (VP-16), have been described in Chinese hamster cells. The mutually exclusive cross-resistance patterns of these mutants provide a highly specific mean to distinguish the two kinds of podofilox derivatives. Mutant Chinese hamster cells resistant to podofilox are affected in a protein P1 that was later identified as the mammalian HSP60 or chaperonin protein. (Wikipedia) | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Metabolism | Half Life: 1.0 to 4.5 hours. | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Toxicity Values | Not Available | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Lethal Dose | Not Available | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Carcinogenicity (IARC Classification) | No indication of carcinogenicity to humans (not listed by IARC). | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Uses/Sources | For treatment of external genital warts (Condyloma acuminatum). | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Minimum Risk Level | Not Available | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Health Effects | Not Available | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symptoms | Not Available | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Treatment | Not Available | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Normal Concentrations | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Abnormal Concentrations | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| External Links | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| DrugBank ID | DB01179 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| HMDB ID | HMDB15310 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PubChem Compound ID | 10607 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| ChEMBL ID | CHEMBL61 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| ChemSpider ID | 10162 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| KEGG ID | C10874 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| UniProt ID | Not Available | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| OMIM ID | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| ChEBI ID | 50305 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| BioCyc ID | Not Available | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| CTD ID | Not Available | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Stitch ID | Not Available | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PDB ID | POD | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| ACToR ID | Not Available | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Wikipedia Link | Podofilox | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| References | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Synthesis Reference | Not Available | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| MSDS | Link | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| General References | Not Available | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Gene Regulation | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Up-Regulated Genes | Not Available | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Down-Regulated Genes | Not Available | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Targets
- General Function:
- Ubiquitin binding
- Specific Function:
- Control of topological states of DNA by transient breakage and subsequent rejoining of DNA strands. Topoisomerase II makes double-strand breaks. Essential during mitosis and meiosis for proper segregation of daughter chromosomes. May play a role in regulating the period length of ARNTL/BMAL1 transcriptional oscillation (By similarity).
- Gene Name:
- TOP2A
- Uniprot ID:
- P11388
- Molecular Weight:
- 174383.88 Da
References
- Iida A, Kano M, Kubota Y, Koga K, Tomioka K: Podophyllotoxin aza-analogue, a novel DNA topoisomerase II inhibitor. Chem Pharm Bull (Tokyo). 2000 Apr;48(4):486-9. [10783066 ]
- Zhang YL, Shen YC, Wang ZQ, Chen HX, Guo X, Cheng YC, Lee KH: Antitumor agents, 130, Novel 4 beta-arylamino derivatives of 3',4'-didemethoxy-3',4'-dioxo-4-deoxypodophyllotoxin as potent inhibitors of human DNA topoisomerase II. J Nat Prod. 1992 Aug;55(8):1100-11. [1331331 ]
- Terada T, Fujimoto K, Nomura M, Yamashita J, Kobunai T, Takeda S, Wierzba K, Yamada Y, Yamaguchi H: Antitumor agents. I. DNA topoisomerase II inhibitory activity and the structural relationship of podophyllotoxin derivatives as antitumor agents. Chem Pharm Bull (Tokyo). 1992 Oct;40(10):2720-7. [1334447 ]
- Kamal A, Gayatri NL, Reddy DR, Mohan Reddy PS, Arifuddin M, Dastidar SG, Kondapi AK, Rajkumar M: Synthesis and biological evaluation of new 4beta-anilino- and 4beta-imido-substituted podophyllotoxin congeners. Bioorg Med Chem. 2005 Nov 15;13(22):6218-25. Epub 2005 Aug 2. [16061385 ]
- Ruckdeschel JC: Etoposide in the management of non-small cell lung cancer. Cancer. 1991 Jan 1;67(1 Suppl):250-3. [1845848 ]
- Chen X, Ji ZL, Chen YZ: TTD: Therapeutic Target Database. Nucleic Acids Res. 2002 Jan 1;30(1):412-5. [11752352 ]
- General Function:
- Structural constituent of cytoskeleton
- Specific Function:
- Tubulin is the major constituent of microtubules. It binds two moles of GTP, one at an exchangeable site on the beta chain and one at a non-exchangeable site on the alpha chain.
- Gene Name:
- TUBA4A
- Uniprot ID:
- P68366
- Molecular Weight:
- 49923.995 Da
References
- Labruere R, Gautier B, Testud M, Seguin J, Lenoir C, Desbene-Finck S, Helissey P, Garbay C, Chabot GG, Vidal M, Giorgi-Renault S: Design, synthesis, and biological evaluation of the first podophyllotoxin analogues as potential vascular-disrupting agents. ChemMedChem. 2010 Dec 3;5(12):2016-25. doi: 10.1002/cmdc.201000305. [20979080 ]
- Li CM, Lu Y, Ahn S, Narayanan R, Miller DD, Dalton JT: Competitive mass spectrometry binding assay for characterization of three binding sites of tubulin. J Mass Spectrom. 2010 Oct;45(10):1160-6. doi: 10.1002/jms.1804. [20814887 ]
- Kim ND, Park ES, Kim YH, Moon SK, Lee SS, Ahn SK, Yu DY, No KT, Kim KH: Structure-based virtual screening of novel tubulin inhibitors and their characterization as anti-mitotic agents. Bioorg Med Chem. 2010 Oct 1;18(19):7092-100. doi: 10.1016/j.bmc.2010.07.072. Epub 2010 Aug 6. [20810285 ]
- Screpanti E, Santaguida S, Nguyen T, Silvestri R, Gussio R, Musacchio A, Hamel E, De Wulf P: A screen for kinetochore-microtubule interaction inhibitors identifies novel antitubulin compounds. PLoS One. 2010 Jul 15;5(7):e11603. doi: 10.1371/journal.pone.0011603. [20657644 ]
- Alam MA, Naik PK: Applying linear interaction energy method for binding affinity calculations of podophyllotoxin analogues with tubulin using continuum solvent model and prediction of cytotoxic activity. J Mol Graph Model. 2009 Jun-Jul;27(8):930-43. doi: 10.1016/j.jmgm.2009.02.003. Epub 2009 Feb 20. [19286405 ]
- Clark PI: Clinical pharmacology and schedule dependency of the podophyllotoxin derivatives. Semin Oncol. 1992 Apr;19(2 Suppl 6):20-7. [1411635 ]
- General Function:
- Zinc ion binding
- Specific Function:
- Receptor for glucocorticoids (GC). Has a dual mode of action: as a transcription factor that binds to glucocorticoid response elements (GRE), both for nuclear and mitochondrial DNA, and as a modulator of other transcription factors. Affects inflammatory responses, cellular proliferation and differentiation in target tissues. Could act as a coactivator for STAT5-dependent transcription upon growth hormone (GH) stimulation and could reveal an essential role of hepatic GR in the control of body growth. Involved in chromatin remodeling. May play a negative role in adipogenesis through the regulation of lipolytic and antilipogenic genes expression.
- Gene Name:
- NR3C1
- Uniprot ID:
- P04150
- Molecular Weight:
- 85658.57 Da
Binding/Activity Constants
| Type | Value | Assay Type | Assay Source |
|---|---|---|---|
| IC50 | 0.014 uM | Not Available | BindingDB 50035218 |
References
- Holloway GA, Charman WN, Fairlamb AH, Brun R, Kaiser M, Kostewicz E, Novello PM, Parisot JP, Richardson J, Street IP, Watson KG, Baell JB: Trypanothione reductase high-throughput screening campaign identifies novel classes of inhibitors with antiparasitic activity. Antimicrob Agents Chemother. 2009 Jul;53(7):2824-33. doi: 10.1128/AAC.01568-08. Epub 2009 Apr 13. [19364854 ]
- General Function:
- Zinc ion binding
- Specific Function:
- Nuclear receptor that binds DNA as a monomer to ROR response elements (RORE) containing a single core motif half-site 5'-AGGTCA-3' preceded by a short A-T-rich sequence. Key regulator of embryonic development, cellular differentiation, immunity, circadian rhythm as well as lipid, steroid, xenobiotics and glucose metabolism. Considered to have intrinsic transcriptional activity, have some natural ligands like oxysterols that act as agonists (25-hydroxycholesterol) or inverse agonists (7-oxygenated sterols), enhancing or repressing the transcriptional activity, respectively. Recruits distinct combinations of cofactors to target genes regulatory regions to modulate their transcriptional expression, depending on the tissue, time and promoter contexts. Regulates genes involved in photoreceptor development including OPN1SW, OPN1SM and ARR3 and skeletal muscle development with MYOD1. Required for proper cerebellum development, regulates SHH gene expression, among others, to induce granule cells proliferation as well as expression of genes involved in calcium-mediated signal transduction. Regulates the circadian expression of several clock genes, including CLOCK, ARNTL/BMAL1, NPAS2 and CRY1. Competes with NR1D1 for binding to their shared DNA response element on some clock genes such as ARNTL/BMAL1, CRY1 and NR1D1 itself, resulting in NR1D1-mediated repression or RORA-mediated activation of clock genes expression, leading to the circadian pattern of clock genes expression. Therefore influences the period length and stability of the clock. Regulates genes involved in lipid metabolism such as apolipoproteins APOA1, APOA5, APOC3 and PPARG. In liver, has specific and redundant functions with RORC as positive or negative modulator of expression of genes encoding phase I and phase II proteins involved in the metabolism of lipids, steroids and xenobiotics, such as CYP7B1 and SULT2A1. Induces a rhythmic expression of some of these genes. In addition, interplays functionally with NR1H2 and NR1H3 for the regulation of genes involved in cholesterol metabolism. Also involved in the regulation of hepatic glucose metabolism through the modulation of G6PC and PCK1. In adipose tissue, plays a role as negative regulator of adipocyte differentiation, probably acting through dual mechanisms. May suppress CEBPB-dependent adipogenesis through direct interaction and PPARG-dependent adipogenesis through competition for DNA-binding. Downstream of IL6 and TGFB and synergistically with RORC isoform 2, is implicated in the lineage specification of uncommitted CD4(+) T-helper (T(H)) cells into T(H)17 cells, antagonizing the T(H)1 program. Probably regulates IL17 and IL17F expression on T(H) by binding to the essential enhancer conserved non-coding sequence 2 (CNS2) in the IL17-IL17F locus. Involved in hypoxia signaling by interacting with and activating the transcriptional activity of HIF1A. May inhibit cell growth in response to cellular stress. May exert an anti-inflammatory role by inducing CHUK expression and inhibiting NF-kappa-B signaling.
- Gene Name:
- RORA
- Uniprot ID:
- P35398
- Molecular Weight:
- 58974.35 Da
Binding/Activity Constants
| Type | Value | Assay Type | Assay Source |
|---|---|---|---|
| IC50 | 0.081 uM | Not Available | BindingDB 50035218 |
References
- Liu T, Lin Y, Wen X, Jorissen RN, Gilson MK: BindingDB: a web-accessible database of experimentally determined protein-ligand binding affinities. Nucleic Acids Res. 2007 Jan;35(Database issue):D198-201. Epub 2006 Dec 1. [17145705 ]
- General Function:
- Transcription factor binding
- Specific Function:
- Orphan nuclear receptor. Component of a cascade required for the development of the hypothalamic-pituitary-adrenal-gonadal axis. Acts as a coregulatory protein that inhibits the transcriptional activity of other nuclear receptors through heterodimeric interactions. May also have a role in the development of the embryo and in the maintenance of embryonic stem cell pluripotency (By similarity).
- Gene Name:
- NR0B1
- Uniprot ID:
- P51843
- Molecular Weight:
- 51717.185 Da
Binding/Activity Constants
| Type | Value | Assay Type | Assay Source |
|---|---|---|---|
| IC50 | 2.145 uM | Not Available | BindingDB 50035218 |
| IC50 | >67.55 uM | Not Available | BindingDB 50035218 |
References
- Liu T, Lin Y, Wen X, Jorissen RN, Gilson MK: BindingDB: a web-accessible database of experimentally determined protein-ligand binding affinities. Nucleic Acids Res. 2007 Jan;35(Database issue):D198-201. Epub 2006 Dec 1. [17145705 ]
- General Function:
- Zinc ion binding
- Specific Function:
- Transcriptional activator. Seems to be essential for sexual differentiation and formation of the primary steroidogenic tissues. Binds to the Ad4 site found in the promoter region of steroidogenic P450 genes such as CYP11A, CYP11B and CYP21B. Also regulates the AMH/Muellerian inhibiting substance gene as well as the AHCH and STAR genes. 5'-YCAAGGYC-3' and 5'-RRAGGTCA-3' are the consensus sequences for the recognition by NR5A1. The SFPQ-NONO-NR5A1 complex binds to the CYP17 promoter and regulates basal and cAMP-dependent transcriptional avtivity. Binds phosphatidylcholine (By similarity). Binds phospholipids with a phosphatidylinositol (PI) headgroup, in particular PI(3,4)P2 and PI(3,4,5)P3. Activated by the phosphorylation of NR5A1 by HIPK3 leading to increased steroidogenic gene expression upon cAMP signaling pathway stimulation.
- Gene Name:
- NR5A1
- Uniprot ID:
- Q13285
- Molecular Weight:
- 51635.47 Da
Binding/Activity Constants
| Type | Value | Assay Type | Assay Source |
|---|---|---|---|
| IC50 | 0.04535 uM | Not Available | BindingDB 50035218 |
| IC50 | >67.54 uM | Not Available | BindingDB 50035218 |
| IC50 | >67.55 uM | Not Available | BindingDB 50035218 |
References
- Liu T, Lin Y, Wen X, Jorissen RN, Gilson MK: BindingDB: a web-accessible database of experimentally determined protein-ligand binding affinities. Nucleic Acids Res. 2007 Jan;35(Database issue):D198-201. Epub 2006 Dec 1. [17145705 ]
- General Function:
- Ubiquitin protein ligase binding
- Specific Function:
- Tubulin is the major constituent of microtubules. It binds two moles of GTP, one at an exchangeable site on the beta chain and one at a non-exchangeable site on the alpha chain.
- Gene Name:
- TUBB
- Uniprot ID:
- P07437
- Molecular Weight:
- 49670.515 Da
References
- Wolff J, Knipling L, Cahnmann HJ, Palumbo G: Direct photoaffinity labeling of tubulin with colchicine. Proc Natl Acad Sci U S A. 1991 Apr 1;88(7):2820-4. [2011590 ]