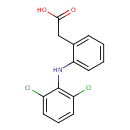
Diclofenac (T3D4729)
| Record Information | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Version | 2.0 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Creation Date | 2014-09-11 05:13:44 UTC | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Update Date | 2014-12-24 20:26:56 UTC | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Accession Number | T3D4729 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Identification | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Common Name | Diclofenac | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Class | Small Molecule | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Description | A non-steroidal anti-inflammatory agent (NSAID) with antipyretic and analgesic actions. It is primarily available as the sodium salt. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Compound Type |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Chemical Structure | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Synonyms |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Chemical Formula | C14H11Cl2NO2 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Average Molecular Mass | 296.149 g/mol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Monoisotopic Mass | 295.017 g/mol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| CAS Registry Number | 15307-86-5 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| IUPAC Name | 2-{2-[(2,6-dichlorophenyl)amino]phenyl}acetic acid | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Traditional Name | diclofenac | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| SMILES | OC(=O)CC1=CC=CC=C1NC1=C(Cl)C=CC=C1Cl | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| InChI Identifier | InChI=1S/C14H11Cl2NO2/c15-10-5-3-6-11(16)14(10)17-12-7-2-1-4-9(12)8-13(18)19/h1-7,17H,8H2,(H,18,19) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| InChI Key | InChIKey=DCOPUUMXTXDBNB-UHFFFAOYSA-N | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Chemical Taxonomy | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Description | belongs to the class of organic compounds known as dichlorobenzenes. Dichlorobenzenes are compounds containing a benzene with exactly two chlorine atoms attached to it. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Kingdom | Organic compounds | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Super Class | Benzenoids | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Class | Benzene and substituted derivatives | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Sub Class | Halobenzenes | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Direct Parent | Dichlorobenzenes | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Alternative Parents | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Substituents |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Molecular Framework | Aromatic homomonocyclic compounds | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| External Descriptors |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Biological Properties | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Status | Detected and Not Quantified | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Origin | Exogenous | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Cellular Locations |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Biofluid Locations | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Tissue Locations | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Pathways | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Applications | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Biological Roles | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Chemical Roles | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Physical Properties | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| State | Solid | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Appearance | White powder. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Experimental Properties |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Predicted Properties |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Spectra | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Spectra |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Toxicity Profile | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Route of Exposure | Completely absorbed from the gastrointestinal tract. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Mechanism of Toxicity | The antiinflammatory effects of diclofenac are believed to be due to inhibition of both leukocyte migration and the enzyme cylooxygenase (COX-1 and COX-2), leading to the peripheral inhibition of prostaglandin synthesis. As prostaglandins sensitize pain receptors, inhibition of their synthesis is responsible for the analgesic effects of diclofenac. Antipyretic effects may be due to action on the hypothalamus, resulting in peripheral dilation, increased cutaneous blood flow, and subsequent heat dissipation. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Metabolism | Hepatic. Route of Elimination: Diclofenac is eliminated through metabolism and subsequent urinary and biliary excretion of the glucuronide and the sulfate conjugates of the metabolites. Little or no free unchanged diclofenac is excreted in the urine. Approximately 65% of the dose is excreted in the urine and approximately 35% in the bile as conjugates of unchanged diclofenac plus metabolites. Half Life: 2 hours | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Toxicity Values | LD50=390mg/kg (orally in mice) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Lethal Dose | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Carcinogenicity (IARC Classification) | No indication of carcinogenicity to humans (not listed by IARC). | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Uses/Sources | For the acute and chronic treatment of signs and symptoms of osteoarthritis and rheumatoid arthritis. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Minimum Risk Level | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Health Effects | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symptoms | Symptoms of overdose include loss of consciousness, increased intracranial pressure, and aspiration pneumonitis. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Treatment | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Normal Concentrations | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Not Available | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Abnormal Concentrations | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Not Available | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| External Links | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| DrugBank ID | DB00586 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| HMDB ID | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PubChem Compound ID | 3033 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| ChEMBL ID | CHEMBL139 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| ChemSpider ID | 2925 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| KEGG ID | C01690 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| UniProt ID | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| OMIM ID | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| ChEBI ID | 4507 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| BioCyc ID | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| CTD ID | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Stitch ID | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PDB ID | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| ACToR ID | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Wikipedia Link | Diclofenac | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| References | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Synthesis Reference | Takuzo Kamishita, “Gel preparations for topical application of diclofenac sodium.” U.S. Patent US4670254, issued October, 1983. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| MSDS | T3D4729.pdf | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| General References |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Gene Regulation | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Up-Regulated Genes | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Down-Regulated Genes | Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Targets
- General Function:
- Prostaglandin-endoperoxide synthase activity
- Specific Function:
- Converts arachidonate to prostaglandin H2 (PGH2), a committed step in prostanoid synthesis. Involved in the constitutive production of prostanoids in particular in the stomach and platelets. In gastric epithelial cells, it is a key step in the generation of prostaglandins, such as prostaglandin E2 (PGE2), which plays an important role in cytoprotection. In platelets, it is involved in the generation of thromboxane A2 (TXA2), which promotes platelet activation and aggregation, vasoconstriction and proliferation of vascular smooth muscle cells.
- Gene Name:
- PTGS1
- Uniprot ID:
- P23219
- Molecular Weight:
- 68685.82 Da
References
- Calkin AC, Sudhir K, Honisett S, Williams MR, Dawood T, Komesaroff PA: Rapid potentiation of endothelium-dependent vasodilation by estradiol in postmenopausal women is mediated via cyclooxygenase 2. J Clin Endocrinol Metab. 2002 Nov;87(11):5072-5. [12414874 ]
- Kirchheiner J, Meineke I, Steinbach N, Meisel C, Roots I, Brockmoller J: Pharmacokinetics of diclofenac and inhibition of cyclooxygenases 1 and 2: no relationship to the CYP2C9 genetic polymorphism in humans. Br J Clin Pharmacol. 2003 Jan;55(1):51-61. [12534640 ]
- Kampfer H, Brautigam L, Geisslinger G, Pfeilschifter J, Frank S: Cyclooxygenase-1-coupled prostaglandin biosynthesis constitutes an essential prerequisite for skin repair. J Invest Dermatol. 2003 May;120(5):880-90. [12713596 ]
- Chavez ML, DeKorte CJ: Valdecoxib: a review. Clin Ther. 2003 Mar;25(3):817-51. [12852704 ]
- Hinz B, Rau T, Auge D, Werner U, Ramer R, Rietbrock S, Brune K: Aceclofenac spares cyclooxygenase 1 as a result of limited but sustained biotransformation to diclofenac. Clin Pharmacol Ther. 2003 Sep;74(3):222-35. [12966366 ]
- Chen X, Ji ZL, Chen YZ: TTD: Therapeutic Target Database. Nucleic Acids Res. 2002 Jan 1;30(1):412-5. [11752352 ]
- General Function:
- Prostaglandin-endoperoxide synthase activity
- Specific Function:
- Converts arachidonate to prostaglandin H2 (PGH2), a committed step in prostanoid synthesis. Constitutively expressed in some tissues in physiological conditions, such as the endothelium, kidney and brain, and in pathological conditions, such as in cancer. PTGS2 is responsible for production of inflammatory prostaglandins. Up-regulation of PTGS2 is also associated with increased cell adhesion, phenotypic changes, resistance to apoptosis and tumor angiogenesis. In cancer cells, PTGS2 is a key step in the production of prostaglandin E2 (PGE2), which plays important roles in modulating motility, proliferation and resistance to apoptosis.
- Gene Name:
- PTGS2
- Uniprot ID:
- P35354
- Molecular Weight:
- 68995.625 Da
References
- Kirchheiner J, Meineke I, Steinbach N, Meisel C, Roots I, Brockmoller J: Pharmacokinetics of diclofenac and inhibition of cyclooxygenases 1 and 2: no relationship to the CYP2C9 genetic polymorphism in humans. Br J Clin Pharmacol. 2003 Jan;55(1):51-61. [12534640 ]
- Blomme EA, Chinn KS, Hardy MM, Casler JJ, Kim SH, Opsahl AC, Hall WA, Trajkovic D, Khan KN, Tripp CS: Selective cyclooxygenase-2 inhibition does not affect the healing of cutaneous full-thickness incisional wounds in SKH-1 mice. Br J Dermatol. 2003 Feb;148(2):211-23. [12588370 ]
- Beubler E: [Pharmacology of cyclooxygenase 2 inhibition]. Wien Med Wochenschr. 2003;153(5-6):95-9. [12705061 ]
- Chavez ML, DeKorte CJ: Valdecoxib: a review. Clin Ther. 2003 Mar;25(3):817-51. [12852704 ]
- Rowlinson SW, Kiefer JR, Prusakiewicz JJ, Pawlitz JL, Kozak KR, Kalgutkar AS, Stallings WC, Kurumbail RG, Marnett LJ: A novel mechanism of cyclooxygenase-2 inhibition involving interactions with Ser-530 and Tyr-385. J Biol Chem. 2003 Nov 14;278(46):45763-9. Epub 2003 Aug 18. [12925531 ]
- General Function:
- Iron ion binding
- Specific Function:
- Catalyzes the first step in leukotriene biosynthesis, and thereby plays a role in inflammatory processes.
- Gene Name:
- ALOX5
- Uniprot ID:
- P09917
- Molecular Weight:
- 77982.595 Da
References
- Charlier C, Michaux C: Dual inhibition of cyclooxygenase-2 (COX-2) and 5-lipoxygenase (5-LOX) as a new strategy to provide safer non-steroidal anti-inflammatory drugs. Eur J Med Chem. 2003 Jul-Aug;38(7-8):645-59. [12932896 ]
- Kudo C, Kori M, Matsuzaki K, Yamai K, Nakajima A, Shibuya A, Niwa H, Kamisaki Y, Wada K: Diclofenac inhibits proliferation and differentiation of neural stem cells. Biochem Pharmacol. 2003 Jul 15;66(2):289-95. [12826271 ]
- Whittle BJ: Cyclooxygenase and nitric oxide systems in the gut as therapeutic targets for safer anti-inflammatory drugs. Curr Opin Pharmacol. 2004 Dec;4(6):538-45. [15525540 ]
- General Function:
- Phospholipid binding
- Specific Function:
- Thought to participate in the regulation of the phospholipid metabolism in biomembranes including eicosanoid biosynthesis. Catalyzes the calcium-dependent hydrolysis of the 2-acyl groups in 3-sn-phosphoglycerides.
- Gene Name:
- PLA2G2A
- Uniprot ID:
- P14555
- Molecular Weight:
- 16082.525 Da
References
- Madanick RD, O'Loughlin CJ, Barkin JS: Diclofenac reduces the incidence of acute pancreatitis after endoscopic retrograde cholangiopancreatography. Dig Dis Sci. 2005 May;50(5):879-81. [15906762 ]
- Singh N, Jabeen T, Sharma S, Somvanshi RK, Dey S, Srinivasan A, Singh TP: Specific binding of non-steroidal anti-inflammatory drugs (NSAIDs) to phospholipase A2: structure of the complex formed between phospholipase A2 and diclofenac at 2.7 A resolution. Acta Crystallogr D Biol Crystallogr. 2006 Apr;62(Pt 4):410-6. Epub 2006 Mar 18. [16552142 ]
- Makela A, Kuusi T, Schroder T: Inhibition of serum phospholipase-A2 in acute pancreatitis by pharmacological agents in vitro. Scand J Clin Lab Invest. 1997 Aug;57(5):401-7. [9279965 ]
- General Function:
- Voltage-gated sodium channel activity
- Specific Function:
- This protein mediates the voltage-dependent sodium ion permeability of excitable membranes. Assuming opened or closed conformations in response to the voltage difference across the membrane, the protein forms a sodium-selective channel through which Na(+) ions may pass in accordance with their electrochemical gradient. This sodium channel may be present in both denervated and innervated skeletal muscle.
- Gene Name:
- SCN4A
- Uniprot ID:
- P35499
- Molecular Weight:
- 208059.175 Da
References
- Yang YC, Kuo CC: An inactivation stabilizer of the Na+ channel acts as an opportunistic pore blocker modulated by external Na+. J Gen Physiol. 2005 May;125(5):465-81. Epub 2005 Apr 11. [15824190 ]
- Voilley N: Acid-sensing ion channels (ASICs): new targets for the analgesic effects of non-steroid anti-inflammatory drugs (NSAIDs). Curr Drug Targets Inflamm Allergy. 2004 Mar;3(1):71-9. [15032643 ]
- Jones NG, Slater R, Cadiou H, McNaughton P, McMahon SB: Acid-induced pain and its modulation in humans. J Neurosci. 2004 Dec 1;24(48):10974-9. [15574747 ]
- General Function:
- Voltage-gated potassium channel activity
- Specific Function:
- Probably important in the regulation of neuronal excitability. Associates with KCNQ3 to form a potassium channel with essentially identical properties to the channel underlying the native M-current, a slowly activating and deactivating potassium conductance which plays a critical role in determining the subthreshold electrical excitability of neurons as well as the responsiveness to synaptic inputs. KCNQ2/KCNQ3 current is blocked by linopirdine and XE991, and activated by the anticonvulsant retigabine. Muscarinic agonist oxotremorine-M strongly suppress KCNQ2/KCNQ3 current in cells in which cloned KCNQ2/KCNQ3 channels were coexpressed with M1 muscarinic receptors.
- Gene Name:
- KCNQ2
- Uniprot ID:
- O43526
- Molecular Weight:
- 95846.575 Da
References
- Peretz A, Degani N, Nachman R, Uziyel Y, Gibor G, Shabat D, Attali B: Meclofenamic acid and diclofenac, novel templates of KCNQ2/Q3 potassium channel openers, depress cortical neuron activity and exhibit anticonvulsant properties. Mol Pharmacol. 2005 Apr;67(4):1053-66. Epub 2004 Dec 14. [15598972 ]
- Xiong Q, Gao Z, Wang W, Li M: Activation of Kv7 (KCNQ) voltage-gated potassium channels by synthetic compounds. Trends Pharmacol Sci. 2008 Feb;29(2):99-107. doi: 10.1016/j.tips.2007.11.010. Epub 2008 Jan 18. [18206251 ]
- General Function:
- Voltage-gated potassium channel activity
- Specific Function:
- Probably important in the regulation of neuronal excitability. Associates with KCNQ2 or KCNQ5 to form a potassium channel with essentially identical properties to the channel underlying the native M-current, a slowly activating and deactivating potassium conductance which plays a critical role in determining the subthreshold electrical excitability of neurons as well as the responsiveness to synaptic inputs.
- Gene Name:
- KCNQ3
- Uniprot ID:
- O43525
- Molecular Weight:
- 96741.515 Da
References
- Xiong Q, Gao Z, Wang W, Li M: Activation of Kv7 (KCNQ) voltage-gated potassium channels by synthetic compounds. Trends Pharmacol Sci. 2008 Feb;29(2):99-107. doi: 10.1016/j.tips.2007.11.010. Epub 2008 Jan 18. [18206251 ]
- Peretz A, Degani N, Nachman R, Uziyel Y, Gibor G, Shabat D, Attali B: Meclofenamic acid and diclofenac, novel templates of KCNQ2/Q3 potassium channel openers, depress cortical neuron activity and exhibit anticonvulsant properties. Mol Pharmacol. 2005 Apr;67(4):1053-66. Epub 2004 Dec 14. [15598972 ]
- General Function:
- Ligand-gated sodium channel activity
- Specific Function:
- Isoform 2 and isoform 3 function as proton-gated sodium channels; they are activated by a drop of the extracellular pH and then become rapidly desensitized. The channel generates a biphasic current with a fast inactivating and a slow sustained phase. Has high selectivity for sodium ions and can also transport lithium ions with high efficiency. Isoform 2 can also transport potassium, but with lower efficiency. It is nearly impermeable to the larger rubidium and cesium ions. Isoform 3 can also transport calcium ions. Mediates glutamate-independent Ca(2+) entry into neurons upon acidosis. This Ca(2+) overloading is toxic for cortical neurons and may be in part responsible for ischemic brain injury. Heteromeric channel assembly seems to modulate channel properties. Functions as a postsynaptic proton receptor that influences intracellular Ca(2+) concentration and calmodulin-dependent protein kinase II phosphorylation and thereby the density of dendritic spines. Modulates activity in the circuits underlying innate fear.Isoform 1 does not display proton-gated cation channel activity.
- Gene Name:
- ASIC1
- Uniprot ID:
- P78348
- Molecular Weight:
- 59908.915 Da
References
- Voilley N, de Weille J, Mamet J, Lazdunski M: Nonsteroid anti-inflammatory drugs inhibit both the activity and the inflammation-induced expression of acid-sensing ion channels in nociceptors. J Neurosci. 2001 Oct 15;21(20):8026-33. [11588175 ]