| Record Information |
|---|
| Version | 2.0 |
|---|
| Creation Date | 2014-09-11 05:16:04 UTC |
|---|
| Update Date | 2014-12-24 20:26:57 UTC |
|---|
| Accession Number | T3D4775 |
|---|
| Identification |
|---|
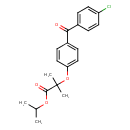
| Common Name | Fenofibrate |
|---|
| Class | Small Molecule |
|---|
| Description | An antilipemic agent which reduces both cholesterol and triglycerides in the blood. |
|---|
| Compound Type | - Drug
- Ester
- Ether
- Hypolipidemic Agent
- Metabolite
- Organic Compound
- Organochloride
- Synthetic Compound
|
|---|
| Chemical Structure | |
|---|
| Synonyms | | Synonym | | 2-(4-(4-Chlorobenzoyl)phenoxy)-2-methylpropanoic acid 1-methylethyl ester | | Antara | | Fenofibrato | | Fenofibratum | | Fenofibric acid | | Fenogal | | Fenoglide | | FIBRICOR | | Finofibrate | | FNF | | Isopropyl (4'-(P-chlorobenzoyl)-2-phenoxy-2-methyl)propionate | | Isopropyl 2-(4-(4-chlorobenzoyl)phenoxy)-2-methylpropionate | | Lipanthyl | | Lipantil | | Lipidil | | Lipofen | | Lofibra | | Procetofen | | Tricor | | TriCor | | Triglide | | Trilipix |
|
|---|
| Chemical Formula | C20H21ClO4 |
|---|
| Average Molecular Mass | 360.831 g/mol |
|---|
| Monoisotopic Mass | 360.113 g/mol |
|---|
| CAS Registry Number | 49562-28-9 |
|---|
| IUPAC Name | propan-2-yl 2-[4-(4-chlorobenzoyl)phenoxy]-2-methylpropanoate |
|---|
| Traditional Name | antara |
|---|
| SMILES | CC(C)OC(=O)C(C)(C)OC1=CC=C(C=C1)C(=O)C1=CC=C(Cl)C=C1 |
|---|
| InChI Identifier | InChI=1S/C20H21ClO4/c1-13(2)24-19(23)20(3,4)25-17-11-7-15(8-12-17)18(22)14-5-9-16(21)10-6-14/h5-13H,1-4H3 |
|---|
| InChI Key | InChIKey=YMTINGFKWWXKFG-UHFFFAOYSA-N |
|---|
| Chemical Taxonomy |
|---|
| Description | belongs to the class of organic compounds known as benzophenones. These are organic compounds containing a ketone attached to two phenyl groups. |
|---|
| Kingdom | Organic compounds |
|---|
| Super Class | Benzenoids |
|---|
| Class | Benzene and substituted derivatives |
|---|
| Sub Class | Benzophenones |
|---|
| Direct Parent | Benzophenones |
|---|
| Alternative Parents | |
|---|
| Substituents | - Benzophenone
- Aryl-phenylketone
- Diphenylmethane
- Phenoxyacetate
- Phenoxy compound
- Aryl ketone
- Phenol ether
- Benzoyl
- Alkyl aryl ether
- Chlorobenzene
- Halobenzene
- Aryl chloride
- Aryl halide
- Carboxylic acid ester
- Ketone
- Monocarboxylic acid or derivatives
- Carboxylic acid derivative
- Ether
- Organohalogen compound
- Organic oxygen compound
- Carbonyl group
- Organochloride
- Organooxygen compound
- Hydrocarbon derivative
- Organic oxide
- Aromatic homomonocyclic compound
|
|---|
| Molecular Framework | Aromatic homomonocyclic compounds |
|---|
| External Descriptors | |
|---|
| Biological Properties |
|---|
| Status | Detected and Not Quantified |
|---|
| Origin | Exogenous |
|---|
| Cellular Locations | |
|---|
| Biofluid Locations | Not Available |
|---|
| Tissue Locations | Not Available |
|---|
| Pathways | Not Available |
|---|
| Applications | Not Available |
|---|
| Biological Roles | Not Available |
|---|
| Chemical Roles | Not Available |
|---|
| Physical Properties |
|---|
| State | Solid |
|---|
| Appearance | White powder. |
|---|
| Experimental Properties | | Property | Value |
|---|
| Melting Point | 80.5°C | | Boiling Point | Not Available | | Solubility | 0.25mg/ml at 25°C | | LogP | 5.3 |
|
|---|
| Predicted Properties | |
|---|
| Spectra |
|---|
| Spectra | | Spectrum Type | Description | Splash Key | Deposition Date | View |
|---|
| Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | splash10-00dl-6390000000-5464dc4b809f70f9340d | 2017-09-01 | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | Not Available | 2021-10-12 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-qTof , Positive | splash10-000i-0122900000-4e3cc64221a0b278b61e | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-ITFT , positive | splash10-001i-0090000000-f4e7816cd30c8701cce3 | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-ITFT , positive | splash10-03di-0029000000-60bbd2de375b9b56ec55 | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-ITFT , positive | splash10-001i-0190000000-6e83f7c906eaeb363caf | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-ITFT , positive | splash10-0019-0930000000-fccf28abf1a426a6949b | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-ITFT , positive | splash10-000i-0900000000-07739caadc1c26a7baaa | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-ITFT , positive | splash10-000i-0900000000-5045a0039b18e3d7dd9a | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-ITFT , positive | splash10-000i-1900000000-25a145c2138d34aa4b52 | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-ITFT , positive | splash10-03di-0029000000-a98245ce05ccb6a6707e | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-ITFT , positive | splash10-001i-0190000000-4e9b3e71fc7960db389e | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-ITFT , positive | splash10-0019-0930000000-9d3908b5ee8b5f59218d | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-ITFT , positive | splash10-000i-0900000000-1090af5e239a872654d9 | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-ITFT , positive | splash10-000i-0900000000-5dcab26837d06f9756f8 | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-ITFT , positive | splash10-000i-1900000000-2663fed5f3ba78d3c7a5 | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-ITFT , positive | splash10-001i-0090000000-4933bea6ce44715c6172 | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - , positive | splash10-01qi-0595000000-d3c4c1cce3b5e1fbb33f | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - , positive | splash10-01qi-1594000000-9756f301ee5d44577654 | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - , positive | splash10-000i-3920000000-91103d7fd9d1063bae9a | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-QFT , positive | splash10-0019-0941000000-27e0cd2c4e952b499a9a | 2017-09-14 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-03di-0039000000-944af46f336da55a8475 | 2016-06-03 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-001i-2293000000-4801131855d2da97d5ee | 2016-06-03 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-00ec-5970000000-1378e4f4b3a07ef9697d | 2016-06-03 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-0a4i-0119000000-2b2235f5a6097ea83d1e | 2016-08-03 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-053r-1297000000-e56c9f467ebb0930e34a | 2016-08-03 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-00lr-0970000000-abe9ec8d570cd8856680 | 2016-08-03 | View Spectrum |
|
|---|
| Toxicity Profile |
|---|
| Route of Exposure | Fenofibrate is well absorbed from the gastrointestinal tract. After absorption, fenofibrate is mainly excreted in the urine in the form of metabolites, primarily fenofibric acid and fenofibric acid glucuronide |
|---|
| Mechanism of Toxicity | Fenofibrate exerts its therapeutic effects through activation of peroxisome proliferator activated receptor a (PPARa). This increases lipolysis and elimination of triglyceride-rich particles from plasma by activating lipoprotein lipase and reducing production of apoprotein C-III. The resulting fall in triglycerides produces an alteration in the size and composition of LDL from small, dense particles, to large buoyant particles. These larger particles have a greater affinity for cholesterol receptors and are catabolized rapidly. |
|---|
| Metabolism | Route of Elimination: Fenofibric acid is primarily conjugated with glucuronic acid and then excreted in urine. Following oral administration in healthy volunteers, approximately 60% of a single dose of radiolabelled fenofibrate appeared in urine, primarily as fenofibric acid and its glucuronate conjugate and 25% was excreted in the feces.
Half Life: 20 hours |
|---|
| Toxicity Values | LD50=1600 mg/kg (Oral, in mice) |
|---|
| Lethal Dose | Not Available |
|---|
| Carcinogenicity (IARC Classification) | No indication of carcinogenicity to humans (not listed by IARC). |
|---|
| Uses/Sources | For use as adjunctive therapy to diet to reduce elevated LDL-C, Total-C,Triglycerides and Apo B, and to increase HDL-C in adult patients with primary hypercholesterolemia or mixed dyslipidemia (Fredrickson Types IIa and IIb) |
|---|
| Minimum Risk Level | Not Available |
|---|
| Health Effects | Investigated as a teratogen and reproductive hazard. |
|---|
| Symptoms | Not Available |
|---|
| Treatment | Not Available |
|---|
| Normal Concentrations |
|---|
| Not Available |
|---|
| Abnormal Concentrations |
|---|
| Not Available |
|---|
| External Links |
|---|
| DrugBank ID | DB01039 |
|---|
| HMDB ID | HMDB15173 |
|---|
| PubChem Compound ID | 3339 |
|---|
| ChEMBL ID | CHEMBL672 |
|---|
| ChemSpider ID | 3222 |
|---|
| KEGG ID | C07586 |
|---|
| UniProt ID | Not Available |
|---|
| OMIM ID | |
|---|
| ChEBI ID | 5001 |
|---|
| BioCyc ID | Not Available |
|---|
| CTD ID | Not Available |
|---|
| Stitch ID | Not Available |
|---|
| PDB ID | Not Available |
|---|
| ACToR ID | Not Available |
|---|
| Wikipedia Link | Fenofibrate |
|---|
| References |
|---|
| Synthesis Reference | Jean-Francois Boyer, “Medicine based on fenofibrate, and a method of preparing it.” U.S. Patent US4800079, issued January, 1988. |
|---|
| MSDS | Link |
|---|
| General References | - Wysocki J, Belowski D, Kalina M, Kochanski L, Okopien B, Kalina Z: Effects of micronized fenofibrate on insulin resistance in patients with metabolic syndrome. Int J Clin Pharmacol Ther. 2004 Apr;42(4):212-7. [15124979 ]
- Keech A, Simes RJ, Barter P, Best J, Scott R, Taskinen MR, Forder P, Pillai A, Davis T, Glasziou P, Drury P, Kesaniemi YA, Sullivan D, Hunt D, Colman P, d'Emden M, Whiting M, Ehnholm C, Laakso M: Effects of long-term fenofibrate therapy on cardiovascular events in 9795 people with type 2 diabetes mellitus (the FIELD study): randomised controlled trial. Lancet. 2005 Nov 26;366(9500):1849-61. [16310551 ]
|
|---|
| Gene Regulation |
|---|
| Up-Regulated Genes | Not Available |
|---|
| Down-Regulated Genes | Not Available |
|---|