| Record Information |
|---|
| Version | 2.0 |
|---|
| Creation Date | 2014-09-11 05:16:36 UTC |
|---|
| Update Date | 2014-12-24 20:26:57 UTC |
|---|
| Accession Number | T3D4788 |
|---|
| Identification |
|---|
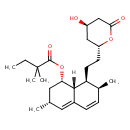
| Common Name | Simvastatin |
|---|
| Class | Small Molecule |
|---|
| Description | Simvastatin is a lipid-lowering agent that is derived synthetically from the fermentation of Aspergillus terreus. It is a potent competitive inhibitor of 3-hydroxy-3-methylglutaryl coenzyme A reductase (hydroxymethylglutaryl COA reductases), which is the rate-limiting enzyme in cholesterol biosynthesis. It may also interfere with steroid hormone production. Due to the induction of hepatic LDL receptors, it increases breakdown of LDL cholesterol. |
|---|
| Compound Type | - Anticholesteremic Agent
- Drug
- Ester
- Ether
- Food Toxin
- Fungal Toxin
- Hydroxymethylglutaryl-CoA Reductase Inhibitor
- Hypolipidemic Agent
- Metabolite
- Organic Compound
- Synthetic Compound
|
|---|
| Chemical Structure | |
|---|
| Synonyms | | Synonym | | (+)-Simvastatin | | 2,2-Dimethylbutyric acid, 8-ester with (4R,6R)-6-(2-((1S,2S,6R,8S,8ar)-1,2,6,7,8,8a-hexahydro-8-hydroxy-2,6-dimethyl-1-naphthyl)ethyl)tetrahydro-4-hydroxy-2H-pyran-2-one | | Cholestat | | Colemin | | Denan | | Eucor | | Kolestevan | | Labistatin | | Lipex | | Lipinorm | | Liponorm | | Lipovas | | Lodales | | Medipo | | MK-733 | | Modutrol | | Nivelipol | | Nor-Vastina | | Pepstatin | | Rechol | | Simcor | | Simovil | | Simvastatin lactone | | Simvastatina | | Simvastatine | | Simvastatinum | | Simvotin | | Sinvacor | | Sinvascor | | Sivastin | | Sivatin | | Sivinar | | Sorfox | | Sotovastin | | Starezin | | Starstat | | Starzoko | | Stasiva | | Statex | | Staticor | | Statin | | Statinal | | Stativer | | Synvinolin | | Valemia | | Velostatin | | Zocor | | Zocord |
|
|---|
| Chemical Formula | C25H38O5 |
|---|
| Average Molecular Mass | 418.566 g/mol |
|---|
| Monoisotopic Mass | 418.272 g/mol |
|---|
| CAS Registry Number | 79902-63-9 |
|---|
| IUPAC Name | (1S,3R,7S,8S,8aR)-8-{2-[(2R,4R)-4-hydroxy-6-oxooxan-2-yl]ethyl}-3,7-dimethyl-1,2,3,7,8,8a-hexahydronaphthalen-1-yl 2,2-dimethylbutanoate |
|---|
| Traditional Name | simvastatin |
|---|
| SMILES | [H][C@]1(O)CC(=O)O[C@]([H])(CC[C@@]2([H])[C@@]([H])(C)C=CC3=C[C@]([H])(C)C[C@]([H])(OC(=O)C(C)(C)CC)[C@]23[H])C1 |
|---|
| InChI Identifier | InChI=1S/C25H38O5/c1-6-25(4,5)24(28)30-21-12-15(2)11-17-8-7-16(3)20(23(17)21)10-9-19-13-18(26)14-22(27)29-19/h7-8,11,15-16,18-21,23,26H,6,9-10,12-14H2,1-5H3/t15-,16-,18+,19+,20-,21-,23-/m0/s1 |
|---|
| InChI Key | InChIKey=RYMZZMVNJRMUDD-HGQWONQESA-N |
|---|
| Chemical Taxonomy |
|---|
| Description | belongs to the class of organic compounds known as delta valerolactones. These are cyclic organic compounds containing an oxan-2- one moiety. |
|---|
| Kingdom | Organic compounds |
|---|
| Super Class | Organoheterocyclic compounds |
|---|
| Class | Lactones |
|---|
| Sub Class | Delta valerolactones |
|---|
| Direct Parent | Delta valerolactones |
|---|
| Alternative Parents | |
|---|
| Substituents | - Delta_valerolactone
- Fatty acid ester
- Delta valerolactone
- Fatty acyl
- Oxane
- Dicarboxylic acid or derivatives
- Secondary alcohol
- Carboxylic acid ester
- Oxacycle
- Carboxylic acid derivative
- Organic oxygen compound
- Organic oxide
- Hydrocarbon derivative
- Organooxygen compound
- Carbonyl group
- Alcohol
- Aliphatic heteropolycyclic compound
|
|---|
| Molecular Framework | Aliphatic heteropolycyclic compounds |
|---|
| External Descriptors | |
|---|
| Biological Properties |
|---|
| Status | Detected and Not Quantified |
|---|
| Origin | Exogenous |
|---|
| Cellular Locations | |
|---|
| Biofluid Locations | Not Available |
|---|
| Tissue Locations | |
|---|
| Pathways | Not Available |
|---|
| Applications | Not Available |
|---|
| Biological Roles | Not Available |
|---|
| Chemical Roles | Not Available |
|---|
| Physical Properties |
|---|
| State | Solid |
|---|
| Appearance | White powder. |
|---|
| Experimental Properties | | Property | Value |
|---|
| Melting Point | 135-138°C | | Boiling Point | Not Available | | Solubility | Insoluble | | LogP | 4.68 |
|
|---|
| Predicted Properties | |
|---|
| Spectra |
|---|
| Spectra | | Spectrum Type | Description | Splash Key | Deposition Date | View |
|---|
| Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | splash10-0fk9-9154200000-9a8f0d47976b67e5c9d5 | 2017-09-01 | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (1 TMS) - 70eV, Positive | splash10-00b9-9137500000-2fdfacbca82ff764afb2 | 2017-10-06 | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | Not Available | 2021-10-12 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-qTof , Positive | splash10-0229-0967000000-6c51c479e90818ec703b | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-qTof , Positive | splash10-0229-0967000000-6c51c479e90818ec703b | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-QTOF , positive | splash10-0fya-0988400000-d9778b2f11696b81fb07 | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-QTOF , positive | splash10-006w-0790000000-4f23e0376d85845e0964 | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-QTOF , positive | splash10-05i1-0940000000-058f6298af54c05532ed | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-QTOF , positive | splash10-0092-0910000000-6ecd838b2f6fddf67dad | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-QTOF , positive | splash10-0597-0900000000-012d0d6fcabbd21d696d | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-QFT , positive | splash10-000e-0590000000-7f99e0b9531d4f42831a | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-QFT , positive | splash10-0092-1960000000-e8429bbec985c5ecdd2d | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-QFT , positive | splash10-0592-1910000000-dc09a51ca37fe4d24a5a | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-QFT , positive | splash10-0abd-1900000000-9b55821f49270e5dd598 | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-QFT , positive | splash10-0536-1900000000-d8a41a97ccbdb30e5a5e | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-QFT , positive | splash10-002f-2900000000-abe04d7f99c3eb0f2fcb | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - 60V, Positive | splash10-0abd-1900000000-78fbdab0700c9c2d0ea4 | 2021-09-20 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - 45V, Positive | splash10-0592-1910000000-88e71bd3ee3ad70a99d1 | 2021-09-20 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - 50V, Positive | splash10-0597-0900000000-093875a17b3bb042dc1b | 2021-09-20 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - 40V, Positive | splash10-0092-0910000000-6ecd838b2f6fddf67dad | 2021-09-20 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - 30V, Positive | splash10-05i1-0940000000-058f6298af54c05532ed | 2021-09-20 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - 10V, Positive | splash10-0fya-0988400000-d9778b2f11696b81fb07 | 2021-09-20 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-0uxr-3028900000-ac9ec6a75c0ccc802808 | 2016-06-03 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-0fr2-9156100000-30ee09c0a70d4a8c7b20 | 2016-06-03 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-00kb-9120000000-4cb8e77d058b47cb3d82 | 2016-06-03 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-014i-0009500000-4449db04da972dd8d05f | 2016-08-03 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-01dj-4109100000-a14d60d17f23d1fb9872 | 2016-08-03 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-0006-9131000000-24b3e278d457c517b414 | 2016-08-03 | View Spectrum | | MS | Mass Spectrum (Electron Ionization) | splash10-0a4i-3900000000-d04758924bf88a90c017 | 2014-09-20 | View Spectrum |
|
|---|
| Toxicity Profile |
|---|
| Route of Exposure | Absorption of simvastatin, estimated relative to an intravenous reference dose, in each of two animal species tested, averaged about 85% of an oral dose. In animal studies, after oral dosing, simvastatin achieved substantially higher concentrations in the liver than in non-target tissues. However, because simvastatin undergoes extensive first-pass metabolism, the availability of the drug in the systemic is low. Peak plasma concentration occurs 1.3 - 2.4 hours after administration. |
|---|
| Mechanism of Toxicity | Simvastatin is a prodrug in which the 6-membered lactone ring of simvastatin is hydrolyzed in vivo to generate the beta,delta-dihydroxy acid, an active metabolite structurally similar to HMG-CoA (hydroxymethylglutaryl CoA). Once hydrolyzed, simvastatin competes with HMG-CoA for HMG-CoA reductase, a hepatic microsomal enzyme. Interference with the activity of this enzyme reduces the quantity of mevalonic acid, a precursor of cholesterol. |
|---|
| Metabolism | Hepatic, simvastatin is a substrate for CYP3A4. The major active metabolites of simvastatin are ‘_-hydroxyacid metabolite and its 6'-hydroxy, 6'-hydroxymethyl, and 6'-exomethylene derivatives
Route of Elimination: Following an oral dose of 14C-labeled simvastatin in man, 13% of the dose was excreted in urine and 60% in feces.
Half Life: 3 hours |
|---|
| Toxicity Values | Not Available |
|---|
| Lethal Dose | Not Available |
|---|
| Carcinogenicity (IARC Classification) | No indication of carcinogenicity to humans (not listed by IARC). |
|---|
| Uses/Sources | For the treatment of hypercholesterolemia and for the reduction in the risk of cardiac heart disease mortality and cardiovascular events. It can also be used in adolescent patients for the treatment of heterozygous familial hypercholesterolemia. |
|---|
| Minimum Risk Level | Not Available |
|---|
| Health Effects | The most common adverse reactions that lead to discontinuation of therapy include gastrointestinal disorders (0.5%), myalgia (0.1%), and arthralgia (0.1%). |
|---|
| Symptoms | Not Available |
|---|
| Treatment | Not Available |
|---|
| Normal Concentrations |
|---|
| Not Available |
|---|
| Abnormal Concentrations |
|---|
| Not Available |
|---|
| External Links |
|---|
| DrugBank ID | DB00641 |
|---|
| HMDB ID | HMDB05007 |
|---|
| PubChem Compound ID | 54454 |
|---|
| ChEMBL ID | CHEMBL1064 |
|---|
| ChemSpider ID | 49179 |
|---|
| KEGG ID | Not Available |
|---|
| UniProt ID | Not Available |
|---|
| OMIM ID | |
|---|
| ChEBI ID | 9150 |
|---|
| BioCyc ID | Not Available |
|---|
| CTD ID | Not Available |
|---|
| Stitch ID | Not Available |
|---|
| PDB ID | Not Available |
|---|
| ACToR ID | Not Available |
|---|
| Wikipedia Link | Simvastatin |
|---|
| References |
|---|
| Synthesis Reference | Shieh-Shung J. Chen, Byron H. Arison, “Process for the preparation of 3-keto, 5-hydroxy simvastatin analogs.” U.S. Patent US4965200, issued April, 1981. |
|---|
| MSDS | Link |
|---|
| General References | - Wolozin B, Wang SW, Li NC, Lee A, Lee TA, Kazis LE: Simvastatin is associated with a reduced incidence of dementia and Parkinson's disease. BMC Med. 2007 Jul 19;5:20. [17640385 ]
- Mahboobi SK, Shohat EZ, Jellinek SP, Rose M: Systemic infections can decrease the threshold of statin-induced muscle injury. South Med J. 2006 Apr;99(4):403-4. [16634254 ]
- Antons KA, Williams CD, Baker SK, Phillips PS: Clinical perspectives of statin-induced rhabdomyolysis. Am J Med. 2006 May;119(5):400-9. [16651050 ]
- Tsivgoulis G, Spengos K, Karandreas N, Panas M, Kladi A, Manta P: Presymptomatic neuromuscular disorders disclosed following statin treatment. Arch Intern Med. 2006 Jul 24;166(14):1519-24. [16864763 ]
- Finsterer J, Zuntner G: Rhabdomyolysis from Simvastatin triggered by infection and muscle exertion. South Med J. 2005 Aug;98(8):827-9. [16144183 ]
- Soininen K, Niemi M, Kilkki E, Strandberg T, Kivisto KT: Muscle symptoms associated with statins: a series of twenty patients. Basic Clin Pharmacol Toxicol. 2006 Jan;98(1):51-4. [16433891 ]
- Davidson MH, Maccubbin D, Stepanavage M, Strony J, Musliner T: Striated muscle safety of ezetimibe/simvastatin (Vytorin). Am J Cardiol. 2006 Jan 15;97(2):223-8. Epub 2005 Nov 21. [16442367 ]
- Hellinger FJ, Encinosa WE: Inappropriate drug combinations among privately insured patients with HIV disease. Med Care. 2005 Sep;43(9 Suppl):III53-62. [16116309 ]
|
|---|
| Gene Regulation |
|---|
| Up-Regulated Genes | Not Available |
|---|
| Down-Regulated Genes | Not Available |
|---|