| Record Information |
|---|
| Version | 2.0 |
|---|
| Creation Date | 2014-10-02 18:59:38 UTC |
|---|
| Update Date | 2018-03-21 17:46:18 UTC |
|---|
| Accession Number | T3D4963 |
|---|
| Identification |
|---|
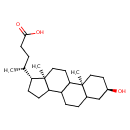
| Common Name | Lithocholic acid |
|---|
| Class | Small Molecule |
|---|
| Description | Lithocholic acid, also known as 3α-hydroxy-5β-cholan-24-oic acid or LCA, is a secondary bile acid. It is formed from chenodeoxycholate by bacterial action, and is usually conjugated with glycine or taurine. It acts as a detergent to solubilize fats for absorption and is itself absorbed. It is used as cholagogue and choleretic. Bile acids are steroid acids found predominantly in the bile of mammals. The distinction between different bile acids is minute, and depends only on the presence or absence of hydroxyl groups on positions 3, 7, and 12. Bile acids are physiological detergents that facilitate excretion, absorption, and transport of fats and sterols in the intestine and liver. Bile acids are also steroidal amphipathic molecules derived from the catabolism of cholesterol. They modulate bile flow and lipid secretion, are essential for the absorption of dietary fats and vitamins, and have been implicated in the regulation of all the key enzymes involved in cholesterol homeostasis. Bile acids recirculate through the liver, bile ducts, small intestine, and portal vein to form an enterohepatic circuit. They exist as anions at physiological pH, and consequently require a carrier for transport across the membranes of the enterohepatic tissues. The unique detergent properties of bile acids are essential for the digestion and intestinal absorption of hydrophobic nutrients. Bile acids have potent toxic properties (e.g. membrane disruption) and there are a plethora of mechanisms to limit their accumulation in blood and tissues (PMID: 11316487, 16037564, 12576301, 11907135). When present in sufficiently high levels, lithocholic acid can act as an oncometabolite. An oncometabolite is a compound that when present at chronically high levels promotes tumour growth and survival. Chronically high levels of lithocholic acid are associated with several forms of cancer including colon cancer, pancreatic cancer, esophageal cancer, and many other GI cancers. High bile acid levels lead to the generation of reactive oxygen species and reactive nitrogen species, disruption of the cell membrane and mitochondria, induction of DNA damage, mutation and apoptosis, and the development of reduced apoptosis capability upon chronic exposure (PMID: 24884764). Dietary fibre can bind to lithocholic acid and aid in its excretion in stool. As such, fibre can protect against colon cancer. |
|---|
| Compound Type | - Animal Toxin
- Metabolite
- Natural Compound
|
|---|
| Chemical Structure | |
|---|
| Synonyms | | Synonym | | (3a,5b)-3-hydroxy-cholan-24-oate | | (3a,5b)-3-hydroxy-cholan-24-oic acid | | 3a-Hydroxy-5b-cholan-24-oate | | 3a-Hydroxy-5b-cholan-24-oic acid | | Lithocholate |
|
|---|
| Chemical Formula | C24H40O3 |
|---|
| Average Molecular Mass | 376.573 g/mol |
|---|
| Monoisotopic Mass | 376.298 g/mol |
|---|
| CAS Registry Number | 434-13-9 |
|---|
| IUPAC Name | (4R)-4-[(2S,5R,14R,15R)-5-hydroxy-2,15-dimethyltetracyclo[8.7.0.0²,⁷.0¹¹,¹⁵]heptadecan-14-yl]pentanoic acid |
|---|
| Traditional Name | (4R)-4-[(2S,5R,14R,15R)-5-hydroxy-2,15-dimethyltetracyclo[8.7.0.0²,⁷.0¹¹,¹⁵]heptadecan-14-yl]pentanoic acid |
|---|
| SMILES | C[C@H](CCC(O)=O)[C@H]1CCC2C3CCC4C[C@H](O)CC[C@]4(C)C3CC[C@]12C |
|---|
| InChI Identifier | InChI=1S/C24H40O3/c1-15(4-9-22(26)27)19-7-8-20-18-6-5-16-14-17(25)10-12-23(16,2)21(18)11-13-24(19,20)3/h15-21,25H,4-14H2,1-3H3,(H,26,27)/t15-,16?,17-,18?,19-,20?,21?,23+,24-/m1/s1 |
|---|
| InChI Key | InChIKey=SMEROWZSTRWXGI-HRFHTWGISA-N |
|---|
| Chemical Taxonomy |
|---|
| Description | belongs to the class of organic compounds known as monohydroxy bile acids, alcohols and derivatives. These are bile acids, alcohols or any of their derivatives bearing a hydroxyl group. |
|---|
| Kingdom | Organic compounds |
|---|
| Super Class | Lipids and lipid-like molecules |
|---|
| Class | Steroids and steroid derivatives |
|---|
| Sub Class | Bile acids, alcohols and derivatives |
|---|
| Direct Parent | Monohydroxy bile acids, alcohols and derivatives |
|---|
| Alternative Parents | |
|---|
| Substituents | - Monohydroxy bile acid, alcohol, or derivatives
- 3-alpha-hydroxysteroid
- Hydroxysteroid
- 3-hydroxysteroid
- Cyclic alcohol
- Secondary alcohol
- Monocarboxylic acid or derivatives
- Carboxylic acid
- Carboxylic acid derivative
- Organic oxygen compound
- Organic oxide
- Hydrocarbon derivative
- Organooxygen compound
- Carbonyl group
- Alcohol
- Aliphatic homopolycyclic compound
|
|---|
| Molecular Framework | Aliphatic homopolycyclic compounds |
|---|
| External Descriptors | Not Available |
|---|
| Biological Properties |
|---|
| Status | Detected and Not Quantified |
|---|
| Origin | Not Available |
|---|
| Cellular Locations | |
|---|
| Biofluid Locations | Not Available |
|---|
| Tissue Locations | |
|---|
| Pathways | Not Available |
|---|
| Applications | Not Available |
|---|
| Biological Roles | Not Available |
|---|
| Chemical Roles | Not Available |
|---|
| Physical Properties |
|---|
| State | Solid |
|---|
| Appearance | White powder. |
|---|
| Experimental Properties | | Property | Value |
|---|
| Melting Point | 186 °C | | Boiling Point | Not Available | | Solubility | 0.000377 mg/mL | | LogP | Not Available |
|
|---|
| Predicted Properties | |
|---|
| Spectra |
|---|
| Spectra | | Spectrum Type | Description | Splash Key | Deposition Date | View |
|---|
| Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-0a6r-0009000000-f631439c42560b6112dc | 2016-08-02 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-0bu3-0029000000-77d9dd354e5acfcbb059 | 2016-08-02 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-0002-1195000000-4e93d1691883a980ad09 | 2016-08-02 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-004i-0009000000-bc8a31ce5625ab01cbf9 | 2016-08-03 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-056r-1009000000-703fb777c311a25b0d15 | 2016-08-03 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-0a4l-9006000000-6c71a6df88d4f84121d0 | 2016-08-03 | View Spectrum |
|
|---|
| Toxicity Profile |
|---|
| Route of Exposure | Not Available |
|---|
| Mechanism of Toxicity | Not Available |
|---|
| Metabolism | Not Available |
|---|
| Toxicity Values | Not Available |
|---|
| Lethal Dose | Not Available |
|---|
| Carcinogenicity (IARC Classification) | Not listed by IARC. |
|---|
| Uses/Sources | Not Available |
|---|
| Minimum Risk Level | Not Available |
|---|
| Health Effects | Chronically high levels of lithocholic acid are associated with several forms of cancer including colon cancer. |
|---|
| Symptoms | Not Available |
|---|
| Treatment | Not Available |
|---|
| Normal Concentrations |
|---|
| Not Available |
|---|
| Abnormal Concentrations |
|---|
| Not Available |
|---|
| External Links |
|---|
| DrugBank ID | Not Available |
|---|
| HMDB ID | HMDB00761 |
|---|
| PubChem Compound ID | 11740284 |
|---|
| ChEMBL ID | Not Available |
|---|
| ChemSpider ID | 9914991 |
|---|
| KEGG ID | C03990 |
|---|
| UniProt ID | Not Available |
|---|
| OMIM ID | |
|---|
| ChEBI ID | 16325 |
|---|
| BioCyc ID | TAUROLITHOCHOLATE-SULFATE |
|---|
| CTD ID | D008095 |
|---|
| Stitch ID | Not Available |
|---|
| PDB ID | Not Available |
|---|
| ACToR ID | Not Available |
|---|
| Wikipedia Link | Not Available |
|---|
| References |
|---|
| Synthesis Reference | Not Available |
|---|
| MSDS | T3D4963.pdf |
|---|
| General References | - Greco AV, Mingrone G: Serum bile acid concentrations in mild liver cirrhosis. Clin Chim Acta. 1993 Nov 30;221(1-2):183-9. [8149635 ]
- Kitahara M, Sakata S, Sakamoto M, Benno Y: Comparison among fecal secondary bile acid levels, fecal microbiota and Clostridium scindens cell numbers in Japanese. Microbiol Immunol. 2004;48(5):367-75. [15215624 ]
- Dew MJ, van Berge Henegouwen GP, Huybregts AW, Allan RN: Hepatotoxic effect of bile acids in inflammatory bowel disease. Gastroenterology. 1980 Jun;78(6):1393-401. [7372059 ]
- Ceryak S, Bouscarel B, Fromm H: Comparative binding of bile acids to serum lipoproteins and albumin. J Lipid Res. 1993 Oct;34(10):1661-74. [8245717 ]
- Eklund A, Norlander A, Norman A: Bile acid synthesis and excretion following release of total extrahepatic cholestasis by percutaneous transhepatic drainage. Eur J Clin Invest. 1980 Oct;10(5):349-55. [6777167 ]
- Balistreri WF, Suchy FJ, Farrell MK, Heubi JE: Pathologic versus physiologic cholestasis: elevated serum concentration of a secondary bile acid in the presence of hepatobiliary disease. J Pediatr. 1981 Mar;98(3):399-402. [7205448 ]
- Deleze G, Paumgartner G, Karlaganis G, Giger W, Reinhard M, Sidiropoulos D: Bile acid pattern in human amniotic fluid. Eur J Clin Invest. 1978 Feb;8(1):41-5. [417931 ]
- Beher WT, Gabbard A, Norum RA, Stradnieks S: Effect of blood high density lipoprotein cholesterol concentration on fecal steroid excretion in humans. Life Sci. 1983 Jun 27;32(26):2933-7. [6865641 ]
- Fouin-Fortunet H, Le Quernec L, Erlinger S, Lerebours E, Colin R: Hepatic alterations during total parenteral nutrition in patients with inflammatory bowel disease: a possible consequence of lithocholate toxicity. Gastroenterology. 1982 May;82(5 Pt 1):932-7. [6800873 ]
- Rudi J, Schonig T, Stremmel W: -Therapy with ursodeoxycholic acid in primary biliary cirrhosis in pregnancy-. Z Gastroenterol. 1996 Mar;34(3):188-91. [8650973 ]
- Hofmann AF: [Enterohepatic circulation of bile acids and biliary lipid secretion]. Minerva Med. 1977 Sep 19;68(43):3011-7. [409965 ]
- Stadler J, Yeung KS, Furrer R, Marcon N, Himal HS, Bruce WR: Proliferative activity of rectal mucosa and soluble fecal bile acids in patients with normal colons and in patients with colonic polyps or cancer. Cancer Lett. 1988 Jan;38(3):315-20. [3349450 ]
- Tadano T, Kanoh M, Matsumoto M, Sakamoto K, Kamano T: Studies of serum and feces bile acids determination by gas chromatography-mass spectrometry. Rinsho Byori. 2006 Feb;54(2):103-10. [16548228 ]
- Loof L, Wengle B: Enzymatic sulphation of bile salts in human liver. Biochim Biophys Acta. 1978 Sep 28;530(3):451-60. [698243 ]
- Salen G, Tint GS, Eliav B, Deering N, Mosbach EH: Increased formation of ursodeoxycholic acid in patients treated with chenodeoxycholic acid. J Clin Invest. 1974 Feb;53(2):612-21. [11344576 ]
- Tinker LF, Schneeman BO, Davis PA, Gallaher DD, Waggoner CR: Consumption of prunes as a source of dietary fiber in men with mild hypercholesterolemia. Am J Clin Nutr. 1991 May;53(5):1259-65. [1850578 ]
- Farrell GC, Duddy SK, Kass GE, Llopis J, Gahm A, Orrenius S: Release of Ca2+ from the endoplasmic reticulum is not the mechanism for bile acid-induced cholestasis and hepatotoxicity in the intact rat liver. J Clin Invest. 1990 Apr;85(4):1255-9. [2318979 ]
- St-Pierre MV, Kullak-Ublick GA, Hagenbuch B, Meier PJ: Transport of bile acids in hepatic and non-hepatic tissues. J Exp Biol. 2001 May;204(Pt 10):1673-86. [11316487 ]
- Claudel T, Staels B, Kuipers F: The Farnesoid X receptor: a molecular link between bile acid and lipid and glucose metabolism. Arterioscler Thromb Vasc Biol. 2005 Oct;25(10):2020-30. Epub 2005 Jul 21. [16037564 ]
- Chiang JY: Bile acid regulation of hepatic physiology: III. Bile acids and nuclear receptors. Am J Physiol Gastrointest Liver Physiol. 2003 Mar;284(3):G349-56. [12576301 ]
- Davis RA, Miyake JH, Hui TY, Spann NJ: Regulation of cholesterol-7alpha-hydroxylase: BAREly missing a SHP. J Lipid Res. 2002 Apr;43(4):533-43. [11907135 ]
|
|---|
| Gene Regulation |
|---|
| Up-Regulated Genes | Not Available |
|---|
| Down-Regulated Genes | Not Available |
|---|