| Record Information |
|---|
| Version | 2.0 |
|---|
| Creation Date | 2014-08-29 06:16:39 UTC |
|---|
| Update Date | 2014-12-24 20:26:45 UTC |
|---|
| Accession Number | T3D4295 |
|---|
| Identification |
|---|
| Common Name | Citrulline |
|---|
| Class | Small Molecule |
|---|
| Description | Citrulline is an amino acid. It is made from ornithine and carbamoyl phosphate in one of the central reactions in the urea cycle. It is also produced from arginine as a by-product of the reaction catalyzed by NOS family. Its name is derived from citrullus, the Latin word for watermelon, from which it was first isolated. |
|---|
| Compound Type | - Amide
- Amine
- Animal Toxin
- Dietary Supplement
- Drug
- Food Toxin
- Metabolite
- Micronutrient
- Natural Compound
- Non-Essential Amino Acid
- Nutraceutical
- Organic Compound
- Supplement
|
|---|
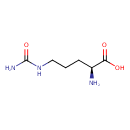
| Chemical Structure | |
|---|
| Synonyms | | Synonym | | (2S)-2-amino-5-(carbamoylamino)pentanoate | | (2S)-2-amino-5-(carbamoylamino)pentanoic acid | | (S)-2-amino-5-(aminocarbonyl)aminopentanoate | | (S)-2-amino-5-(aminocarbonyl)aminopentanoic acid | | (S)-2-Amino-5-ureidopentanoate | | (S)-2-Amino-5-ureidopentanoic acid | | 2-Amino-5-uredovalerate | | 2-Amino-5-uredovaleric acid | | 2-Amino-5-ureidovalerate | | 2-Amino-5-ureidovaleric acid | | A-Amino-D-ureidovalerate | | A-Amino-D-ureidovaleric acid | | alpha-Amino-delta-ureidovalerate | | alpha-Amino-delta-ureidovaleric acid | | alpha-Amino-gamma-ureidovalerate | | alpha-Amino-gamma-ureidovaleric acid | | Amino-ureidovalerate | | Amino-ureidovaleric acid | | CIR | | CIT | | Cytrulline | | D-Ureidonorvaline | | delta-Ureidonorvaline | | DL-citrulline | | Gammaureidonorvaline | | H-Cit-oh | | L(+)-2-Amino-5-ureidovalerate | | L(+)-2-Amino-5-ureidovaleric acid | | L(+)-Citrulline | | L-2-Amino-5-ureido-valerate | | L-2-Amino-5-ureido-valeric acid | | L-2-Amino-5-ureidovalerate | | L-2-Amino-5-ureidovaleric acid | | L-Citrulline | | L-Cytrulline | | L-N5-carbamoyl-Ornithine | | N()-Carbamylornithine | | N(5)-(Aminocarbonyl)-DL-Ornithine | | N(5)-(Aminocarbonyl)-L-ornithine | | N(delta)-Carbamylornithine | | N-Carbamylornithine | | N5-(Aminocarbonyl)-L-ornithine | | N5-(aminocarbonyl)-Ornithine | | N5-(Aminocarbonyl)ornithine | | N5-Carbamoyl-L-ornithine | | N5-carbamoylornithine | | N5-carbamylornithine | | N<SUP>5</SUP>-(aminocarbonyl)ornithine | | ND-carbamylornithine | | Ndelta-carbamy-ornithine | | Ndelta-carbamylornithine | | Ngamma-carbamylornithine | | Sitrulline | | Ureidonorvaline | | Ureidovalerate | | Ureidovaleric acid | | α-amino-δ-ureidovaleric acid | | δ-ureidonorvaline |
|
|---|
| Chemical Formula | C6H13N3O3 |
|---|
| Average Molecular Mass | 175.186 g/mol |
|---|
| Monoisotopic Mass | 175.096 g/mol |
|---|
| CAS Registry Number | 372-75-8 |
|---|
| IUPAC Name | (2S)-2-amino-5-(carbamoylamino)pentanoic acid |
|---|
| Traditional Name | L-citrulline |
|---|
| SMILES | [H][C@](N)(CCCNC(O)=N)C(O)=O |
|---|
| InChI Identifier | InChI=1S/C6H13N3O3/c7-4(5(10)11)2-1-3-9-6(8)12/h4H,1-3,7H2,(H,10,11)(H3,8,9,12)/t4-/m0/s1 |
|---|
| InChI Key | InChIKey=RHGKLRLOHDJJDR-BYPYZUCNSA-N |
|---|
| Chemical Taxonomy |
|---|
| Description | belongs to the class of organic compounds known as l-alpha-amino acids. These are alpha amino acids which have the L-configuration of the alpha-carbon atom. |
|---|
| Kingdom | Organic compounds |
|---|
| Super Class | Organic acids and derivatives |
|---|
| Class | Carboxylic acids and derivatives |
|---|
| Sub Class | Amino acids, peptides, and analogues |
|---|
| Direct Parent | L-alpha-amino acids |
|---|
| Alternative Parents | |
|---|
| Substituents | - L-alpha-amino acid
- Fatty acid
- Isourea
- Amino acid
- Carboximidic acid derivative
- Carboxylic acid
- Monocarboxylic acid or derivatives
- Carboximidamide
- Amine
- Hydrocarbon derivative
- Primary amine
- Organooxygen compound
- Organonitrogen compound
- Organic oxide
- Primary aliphatic amine
- Organopnictogen compound
- Imine
- Organic oxygen compound
- Organic nitrogen compound
- Carbonyl group
- Aliphatic acyclic compound
|
|---|
| Molecular Framework | Aliphatic acyclic compounds |
|---|
| External Descriptors | |
|---|
| Biological Properties |
|---|
| Status | Detected and Not Quantified |
|---|
| Origin | Endogenous |
|---|
| Cellular Locations | |
|---|
| Biofluid Locations | Not Available |
|---|
| Tissue Locations | - Bladder
- Epidermis
- Fibroblasts
- Intestine
- Kidney
- Liver
- Myelin
- Nerve Cells
- Neuron
- Placenta
- Platelet
- Prostate
|
|---|
| Pathways | |
|---|
| Applications | |
|---|
| Biological Roles | |
|---|
| Chemical Roles | |
|---|
| Physical Properties |
|---|
| State | Solid |
|---|
| Appearance | White powder. |
|---|
| Experimental Properties | | Property | Value |
|---|
| Melting Point | 235.5°C | | Boiling Point | Not Available | | Solubility | 200 g/L (at 20°C) | | LogP | -3.19 |
|
|---|
| Predicted Properties | |
|---|
| Spectra |
|---|
| Spectra | | Spectrum Type | Description | Splash Key | Deposition Date | View |
|---|
| GC-MS | GC-MS Spectrum - GC-EI-TOF (Pegasus III TOF-MS system, Leco; GC 6890, Agilent Technologies) (Non-derivatized) | splash10-0a4i-0920000000-2d92b63cd5d9648023b8 | 2014-06-16 | View Spectrum | | GC-MS | GC-MS Spectrum - GC-MS (3 TMS) | splash10-00di-9610000000-2e7cd23afc2adcef35a3 | 2014-06-16 | View Spectrum | | GC-MS | GC-MS Spectrum - GC-EI-TOF (Non-derivatized) | splash10-0a4i-0920000000-2d92b63cd5d9648023b8 | 2017-09-12 | View Spectrum | | GC-MS | GC-MS Spectrum - GC-MS (Non-derivatized) | splash10-00di-9610000000-2e7cd23afc2adcef35a3 | 2017-09-12 | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | splash10-007o-9100000000-1f8dd2c6648b104639c7 | 2016-09-22 | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (1 TMS) - 70eV, Positive | splash10-00dl-9410000000-37909012a777213f8566 | 2017-10-06 | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | Not Available | 2021-10-12 | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | Not Available | 2021-10-12 | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_1_2) - 70eV, Positive | Not Available | 2021-11-06 | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_1_3) - 70eV, Positive | Not Available | 2021-11-06 | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_1_4) - 70eV, Positive | Not Available | 2021-11-06 | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TBDMS_1_1) - 70eV, Positive | Not Available | 2021-11-06 | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TBDMS_1_2) - 70eV, Positive | Not Available | 2021-11-06 | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TBDMS_1_3) - 70eV, Positive | Not Available | 2021-11-06 | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TBDMS_1_4) - 70eV, Positive | Not Available | 2021-11-06 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - Quattro_QQQ 10V, Positive (Annotated) | splash10-0a4i-0900000000-4c1d7af748a47e489949 | 2012-07-24 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - Quattro_QQQ 25V, Positive (Annotated) | splash10-00di-9000000000-988fced362fc0da157c9 | 2012-07-24 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - Quattro_QQQ 40V, Positive (Annotated) | splash10-00di-9000000000-0818e0e8bcee12692498 | 2012-07-24 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-ITFT (LTQ Orbitrap XL, Thermo Scientfic) , Positive | splash10-004j-0900000000-5fa8a338dcd2f2a6bdd2 | 2012-08-31 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-ITFT (LTQ Orbitrap XL, Thermo Scientfic) , Positive | splash10-004i-0900000000-16763200aa07f7629ad4 | 2012-08-31 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-ITFT (LTQ Orbitrap XL, Thermo Scientfic) , Positive | splash10-03di-3900000000-d9cfc5187aa799f6f978 | 2012-08-31 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-ITFT (LTQ Orbitrap XL, Thermo Scientfic) , Positive | splash10-0a4i-0900000000-10ee9a593e13550bec1c | 2012-08-31 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-ITFT (LTQ Orbitrap XL, Thermo Scientfic) , Positive | splash10-004i-0900000000-45d272576af34c9512a3 | 2012-08-31 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-ITFT (LTQ Orbitrap XL, Thermo Scientfic) , Positive | splash10-014i-3900000000-6177a284fdea5a3f1306 | 2012-08-31 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-ITFT (LTQ Orbitrap XL, Thermo Scientfic) , Positive | splash10-0a4i-0900000000-d9456d45e2dbd7a3df10 | 2012-08-31 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-ITFT (LTQ Orbitrap XL, Thermo Scientfic) , Positive | splash10-004i-0900000000-ada57cdc73bda93be483 | 2012-08-31 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-ITFT (LTQ Orbitrap XL, Thermo Scientfic) , Negative | splash10-008a-0904000000-23fbe48f82e515087d68 | 2012-08-31 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-ITFT (LTQ Orbitrap XL, Thermo Scientfic) , Negative | splash10-03di-5900000000-78afcbaf8b8b3eabf174 | 2012-08-31 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-ITFT (LTQ Orbitrap XL, Thermo Scientfic) , Negative | splash10-001i-0900000000-8fb191d4c20fd54b9282 | 2012-08-31 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-ITFT (LTQ Orbitrap XL, Thermo Scientfic) , Negative | splash10-00di-0900000000-da484f0362a8dca5127e | 2012-08-31 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-QQ (API3000, Applied Biosystems) 10V, Negative | splash10-00e9-0900000000-46229b4f77feabb3f857 | 2012-08-31 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-QQ (API3000, Applied Biosystems) 20V, Negative | splash10-001i-0900000000-4aca1022c393602a297d | 2012-08-31 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-QQ (API3000, Applied Biosystems) 30V, Negative | splash10-001i-0900000000-3bc2eff2e907b7734cc8 | 2012-08-31 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-QQ (API3000, Applied Biosystems) 40V, Negative | splash10-001i-3900000000-2613bf40e3be814da86f | 2012-08-31 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-QQ (API3000, Applied Biosystems) 50V, Negative | splash10-0006-9300000000-e83287bbc060eb9cf6f3 | 2012-08-31 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-QQ (API3000, Applied Biosystems) 10V, Positive | splash10-056r-0900000000-694a8872bdfd7eec1f2b | 2012-08-31 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-QQ (API3000, Applied Biosystems) 20V, Positive | splash10-08fr-2900000000-15b4711991ea9985fb2b | 2012-08-31 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-QQ (API3000, Applied Biosystems) 30V, Positive | splash10-00di-9300000000-915fbb73e0b728420e4a | 2012-08-31 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-QQ (API3000, Applied Biosystems) 40V, Positive | splash10-00di-9000000000-67e60567f5c062728350 | 2012-08-31 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-QQ (API3000, Applied Biosystems) 50V, Positive | splash10-00di-9000000000-25140713431edd7c5eea | 2012-08-31 | View Spectrum | | 1D NMR | 13C NMR Spectrum (1D, 125 MHz, H2O, experimental) | Not Available | 2012-12-04 | View Spectrum | | 1D NMR | 1H NMR Spectrum (1D, 500 MHz, H2O, experimental) | Not Available | 2012-12-04 | View Spectrum | | 1D NMR | 1H NMR Spectrum (1D, D2O, experimental) | Not Available | 2016-10-22 | View Spectrum | | 1D NMR | 1H NMR Spectrum (1D, 100 MHz, D2O, predicted) | Not Available | 2021-09-24 | View Spectrum | | 1D NMR | 13C NMR Spectrum (1D, 100 MHz, D2O, predicted) | Not Available | 2021-09-24 | View Spectrum | | 1D NMR | 1H NMR Spectrum (1D, 1000 MHz, D2O, predicted) | Not Available | 2021-09-24 | View Spectrum | | 1D NMR | 13C NMR Spectrum (1D, 1000 MHz, D2O, predicted) | Not Available | 2021-09-24 | View Spectrum | | 1D NMR | 1H NMR Spectrum (1D, 200 MHz, D2O, predicted) | Not Available | 2021-09-24 | View Spectrum | | 1D NMR | 13C NMR Spectrum (1D, 200 MHz, D2O, predicted) | Not Available | 2021-09-24 | View Spectrum | | 1D NMR | 1H NMR Spectrum (1D, 300 MHz, D2O, predicted) | Not Available | 2021-09-24 | View Spectrum | | 1D NMR | 13C NMR Spectrum (1D, 300 MHz, D2O, predicted) | Not Available | 2021-09-24 | View Spectrum | | 1D NMR | 1H NMR Spectrum (1D, 400 MHz, D2O, predicted) | Not Available | 2021-09-24 | View Spectrum | | 1D NMR | 13C NMR Spectrum (1D, 400 MHz, D2O, predicted) | Not Available | 2021-09-24 | View Spectrum | | 1D NMR | 1H NMR Spectrum (1D, 500 MHz, D2O, predicted) | Not Available | 2021-09-24 | View Spectrum | | 1D NMR | 13C NMR Spectrum (1D, 500 MHz, D2O, predicted) | Not Available | 2021-09-24 | View Spectrum | | 1D NMR | 1H NMR Spectrum (1D, 600 MHz, D2O, predicted) | Not Available | 2021-09-24 | View Spectrum | | 1D NMR | 13C NMR Spectrum (1D, 600 MHz, D2O, predicted) | Not Available | 2021-09-24 | View Spectrum | | 1D NMR | 1H NMR Spectrum (1D, 700 MHz, D2O, predicted) | Not Available | 2021-09-24 | View Spectrum | | 1D NMR | 13C NMR Spectrum (1D, 700 MHz, D2O, predicted) | Not Available | 2021-09-24 | View Spectrum | | 1D NMR | 1H NMR Spectrum (1D, 800 MHz, D2O, predicted) | Not Available | 2021-09-24 | View Spectrum | | 1D NMR | 13C NMR Spectrum (1D, 800 MHz, D2O, predicted) | Not Available | 2021-09-24 | View Spectrum | | 1D NMR | 1H NMR Spectrum (1D, 900 MHz, D2O, predicted) | Not Available | 2021-09-24 | View Spectrum | | 1D NMR | 13C NMR Spectrum (1D, 900 MHz, D2O, predicted) | Not Available | 2021-09-24 | View Spectrum | | 1D NMR | 13C NMR Spectrum (1D, 400 MHz, H2O, experimental) | Not Available | 2021-10-10 | View Spectrum | | 2D NMR | [1H, 1H]-TOCSY. Unexported temporarily by An Chi on Oct 15, 2021 until json or nmrML file is generated. 2D NMR Spectrum (experimental) | Not Available | 2012-12-04 | View Spectrum | | 2D NMR | [1H, 13C]-HSQC NMR Spectrum (2D, 600 MHz, H2O, experimental) | Not Available | 2012-12-05 | View Spectrum |
|
|---|
| Toxicity Profile |
|---|
| Route of Exposure | Not Available |
|---|
| Mechanism of Toxicity | L-citrulline is converted to L-arginine by argininosuccinate synthase. L-arginine is in turn responsible for citrulline's therapeutic affects. Many of L-arginine's activities, including its possible anti-atherogenic actions, may be accounted for by its role as the precursor to nitric oxide or NO. NO is produced by all tissues of the body and plays very important roles in the cardiovascular system, immune system and nervous system. NO is formed from L-arginine via the enzyme nitric oxide synthase or synthetase (NOS), and the effects of NO are mainly mediated by 3',5' -cyclic guanylate or cyclic GMP. NO activates the enzyme guanylate cyclase, which catalyzes the synthesis of cyclic GMP from guanosine triphosphate or GTP. Cyclic GMP is converted to guanylic acid via the enzyme cyclic GMP phosphodiesterase.
NOS is a heme-containing enzyme with some sequences similar to cytochrome P-450 reductase. Several isoforms of NOS exist, two of which are constitutive and one of which is inducible by immunological stimuli. The constitutive NOS found in the vascular endothelium is designated eNOS and that present in the brain, spinal cord and peripheral nervous system is designated nNOS. The form of NOS induced by immunological or inflammatory stimuli is known as iNOS. iNOS may be expressed constitutively in select tissues such as lung epithelium.
All the nitric oxide synthases use NADPH (reduced nicotinamide adenine dinucleotide phosphate) and oxygen (O2) as cosubstrates, as well as the cofactors FAD (flavin adenine dinucleotide), FMN (flavin mononucleotide), tetrahydrobiopterin and heme. Interestingly, ascorbic acid appears to enhance NOS activity by increasing intracellular tetrahydrobiopterin. eNOS and nNOS synthesize NO in response to an increased concentration of calcium ions or in some cases in response to calcium-independent stimuli, such as shear stress. In vitro studies of NOS indicate that the Km of the enzyme for L-arginine is in the micromolar range. The concentration of L-arginine in endothelial cells, as well as in other cells, and in plasma is in the millimolar range. What this means is that, under physiological conditions, NOS is saturated with its L-arginine substrate. In other words, L-arginine would not be expected to be rate-limiting for the enzyme, and it would not appear that supraphysiological levels of L-arginine which could occur with oral supplementation of the amino acid would make any difference with regard to NO production. The reaction would appear to have reached its maximum level. However, in vivo studies have demonstrated that, under certain conditions, e.g. hypercholesterolemia, L-arginine could enhance endothelial-dependent vasodilation and NO production. |
|---|
| Metabolism | Not Available |
|---|
| Toxicity Values | Not Available |
|---|
| Lethal Dose | Not Available |
|---|
| Carcinogenicity (IARC Classification) | No indication of carcinogenicity to humans (not listed by IARC). |
|---|
| Uses/Sources | Used for nutritional supplementation, also for treating dietary shortage or imbalance. |
|---|
| Minimum Risk Level | Not Available |
|---|
| Health Effects | Not Available |
|---|
| Symptoms | Not Available |
|---|
| Treatment | Not Available |
|---|
| Normal Concentrations |
|---|
| Not Available |
|---|
| Abnormal Concentrations |
|---|
| Not Available |
|---|
| External Links |
|---|
| DrugBank ID | DB00155 |
|---|
| HMDB ID | HMDB00904 |
|---|
| PubChem Compound ID | 9750 |
|---|
| ChEMBL ID | CHEMBL444814 |
|---|
| ChemSpider ID | 9367 |
|---|
| KEGG ID | C00327 |
|---|
| UniProt ID | Not Available |
|---|
| OMIM ID | |
|---|
| ChEBI ID | 16349 |
|---|
| BioCyc ID | L-CITRULLINE |
|---|
| CTD ID | Not Available |
|---|
| Stitch ID | Not Available |
|---|
| PDB ID | CIR |
|---|
| ACToR ID | Not Available |
|---|
| Wikipedia Link | Citrulline |
|---|
| References |
|---|
| Synthesis Reference | Hua Bai, Peijie Yang, Zhengjie Chen, Chongyan Xu, Zhaorul Li, Zigang Zhao, Luyan Jiang, Zongyi Yang, Jiang Li, “PROCESSES FOR THE PRODUCTION OF L-CITRULLINE.” U.S. Patent US20090142813, issued June 04, 2009. |
|---|
| MSDS | Link |
|---|
| General References | - DeLong GR, Glick TH: Ammonia metabolism in Reye syndrome and the effect of citrulline. Ann Neurol. 1982 Jan;11(1):53-8. [7059128 ]
- Jianfeng G, Weiming Z, Ning L, Fangnan L, Li T, Nan L, Jieshou L: Serum citrulline is a simple quantitative marker for small intestinal enterocytes mass and absorption function in short bowel patients. J Surg Res. 2005 Aug;127(2):177-82. [15921697 ]
- Cynober LA: Plasma amino acid levels with a note on membrane transport: characteristics, regulation, and metabolic significance. Nutrition. 2002 Sep;18(9):761-6. [12297216 ]
- Liappis N, Pohl B, Weber HP, el-Karkani H: [Free amino acids in the saliva of children with phenylketonuria]. Klin Padiatr. 1986 Jan-Feb;198(1):25-8. [3959486 ]
- Rainesalo S, Keranen T, Palmio J, Peltola J, Oja SS, Saransaari P: Plasma and cerebrospinal fluid amino acids in epileptic patients. Neurochem Res. 2004 Jan;29(1):319-24. [14992292 ]
- Fleisher LD, Harris CJ, Mitchell DA, Nadler HL: Citrullinemia: prenatal diagnosis of an affected fetus. Am J Hum Genet. 1983 Jan;35(1):85-90. [6823975 ]
- Potter MA, Zeesman S, Brennan B, Kobayashi K, Gao HZ, Tabata A, Saheki T, Whelan DT: Pregnancy in a healthy woman with untreated citrullinemia. Am J Med Genet A. 2004 Aug 15;129A(1):77-82. [15266621 ]
- Wheatley DN, Kilfeather R, Stitt A, Campbell E: Integrity and stability of the citrulline-arginine pathway in normal and tumour cell lines. Cancer Lett. 2005 Sep 28;227(2):141-52. [16112417 ]
- Melis GC, Boelens PG, van der Sijp JR, Popovici T, De Bandt JP, Cynober L, van Leeuwen PA: The feeding route (enteral or parenteral) affects the plasma response of the dipetide Ala-Gln and the amino acids glutamine, citrulline and arginine, with the administration of Ala-Gln in preoperative patients. Br J Nutr. 2005 Jul;94(1):19-26. [16115328 ]
- Reparon-Schuijt CC, van Esch WJ, van Kooten C, Schellekens GA, de Jong BA, van Venrooij WJ, Breedveld FC, Verweij CL: Secretion of anti-citrulline-containing peptide antibody by B lymphocytes in rheumatoid arthritis. Arthritis Rheum. 2001 Jan;44(1):41-7. [11212174 ]
- Maruyama H, Ogawa M, Nishio T, Kobayashi K, Saheki T, Sunohara N: Citrullinemia type II in a 64-year-old man with fluctuating serum citrulline levels. J Neurol Sci. 2001 Jan 1;182(2):167-70. [11137523 ]
- McLaurin J, Moscarello MA: The preparation of antibodies reactive against citrulline-containing charge isomers of myelin basic protein but not against the arginine-containing charge isomers. Anal Biochem. 1990 Dec;191(2):272-7. [1707596 ]
- Facchinetti F, Longo M, Piccinini F, Neri I, Volpe A: L-arginine infusion reduces blood pressure in preeclamptic women through nitric oxide release. J Soc Gynecol Investig. 1999 Jul-Aug;6(4):202-7. [10486782 ]
- Origuchi Y, Ushijima T, Sakaguchi M, Akaboshi I, Matsuda I: Citrullinemia presenting as uncontrollable epilepsy. Brain Dev. 1984;6(3):328-31. [6486381 ]
- Booth FA, Haworth JC, Dilling LA, Perry TL, Greenberg CR, Seargeant LE, Penn AM, Rhead WJ: Mitochondrial encephalomyopathy with associated aminoacidopathy in a male sibship. J Pediatr. 1989 Jul;115(1):81-8. [2738799 ]
- Engelborghs S, Marescau B, De Deyn PP: Amino acids and biogenic amines in cerebrospinal fluid of patients with Parkinson's disease. Neurochem Res. 2003 Aug;28(8):1145-50. [12834252 ]
- Le Boucher J, Charret C, Coudray-Lucas C, Giboudeau J, Cynober L: Amino acid determination in biological fluids by automated ion-exchange chromatography: performance of Hitachi L-8500A. Clin Chem. 1997 Aug;43(8 Pt 1):1421-8. [9267323 ]
- Crenn P, Coudray-Lucas C, Thuillier F, Cynober L, Messing B: Postabsorptive plasma citrulline concentration is a marker of absorptive enterocyte mass and intestinal failure in humans. Gastroenterology. 2000 Dec;119(6):1496-505. [11113071 ]
- Hagenfeldt L, Bjerkenstedt L, Edman G, Sedvall G, Wiesel FA: Amino acids in plasma and CSF and monoamine metabolites in CSF: interrelationship in healthy subjects. J Neurochem. 1984 Mar;42(3):833-7. [6198473 ]
- Sreekumar A, Poisson LM, Rajendiran TM, Khan AP, Cao Q, Yu J, Laxman B, Mehra R, Lonigro RJ, Li Y, Nyati MK, Ahsan A, Kalyana-Sundaram S, Han B, Cao X, Byun J, Omenn GS, Ghosh D, Pennathur S, Alexander DC, Berger A, Shuster JR, Wei JT, Varambally S, Beecher C, Chinnaiyan AM: Metabolomic profiles delineate potential role for sarcosine in prostate cancer progression. Nature. 2009 Feb 12;457(7231):910-4. doi: 10.1038/nature07762. [19212411 ]
|
|---|
| Gene Regulation |
|---|
| Up-Regulated Genes | Not Available |
|---|
| Down-Regulated Genes | Not Available |
|---|