| Record Information |
|---|
| Version | 2.0 |
|---|
| Creation Date | 2014-08-29 06:03:11 UTC |
|---|
| Update Date | 2018-03-21 17:46:11 UTC |
|---|
| Accession Number | T3D4247 |
|---|
| Identification |
|---|
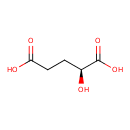
| Common Name | L-2-Hydroxyglutaric acid |
|---|
| Class | Small Molecule |
|---|
| Description | L-2-Hydroxyglutaric acid is a metabolite that accumulates in L-2-hydroxyglutaric aciduria, which is a neurometabolic disorder (OMIM: 236792), and has been reported in multiple patients who have a clinical phenotype of progressive neurodegeneration with extrapyramidal and cerebellar signs, seizures, and spongiform changes in the white matter (OMIM: 600721). In humans, 2-hydroxyglutarate is formed by a hydroxyacid-oxoacid transhydrogenase whereas in bacteria it is formed by a 2-hydroxyglutarate synthase. L-2-Hydroxyglutaric acid can be converted to alpha-ketoglutaric acid through the action of 2-hydroxyglutarate dehydrogenase (EC 1.1.99.2). In humans, there are two such enzymes (D2HGDH and L2HGDH). Both the D and L stereoisomers of hydroxyglutaric acid (EC 1.1.99.2) are found in body fluids. L-2-Hydroxyglutaric acid can also be produced via gain-of-function mutations in the cytosolic and mitochondrial isoforms of isocitrate dehydrogenase (IDH). IDH is part of the TCA cycle and this compound is generated in high abundance when IDH is mutated. Since L-2-hydroxyglutaric acid is sufficiently similar in structure to 2-oxoglutarate (2OG), it is able to inhibit a range of 2OG-dependent dioxygenases, including histone lysine demethylases (KDMs) and members of the ten-eleven translocation (TET) family of 5-methylcytosine (5mC) hydroxylases. This inhibitory effect leads to alterations in the hypoxia-inducible factor (HIF)-mediated hypoxic response and alterations in gene expression through global epigenetic remodeling. The net effect is that L-2-hydroxyglutaric acid causes a cascading effect that leads genetic perturbations and malignant transformation. Depending on the circumstances, L-2-hydroxyglutaric acid can function as an oncometabolite, a neurotoxin, an acidogen, and a metabotoxin. An oncometabolite is a compound that promotes tumour growth and survival. A neurotoxin is a compound that is toxic to neurons or neural tissue. An acidogen is an acidic compound that induces acidosis, which has multiple adverse effects on many organ systems. A metabotoxin is an endogenously produced metabolite that causes adverse health effects at chronically high levels. As an oncometabolite, L-2-hydroxyglutaric acid is a competitive inhibitor of multiple alpha-ketoglutarate-dependent dioxygenases, including histone demethylases and the TET family of 5mC hydroxylases. As a result, high levels of 2-hydroxyglutarate lead to genome-wide histone and DNA methylation alterations, which in turn lead to mutations that ultimately cause cancer (PMID: 29038145). As a neurotoxin, L-2-hydroxyglutaric acid mediates its neurotoxicity through activation of N-methyl-D-aspartate receptors. L-2-Hydroxyglutaric acid is structurally similar to the excitatory amino acid glutamate and stimulates neurodegeneration by mechanisms similar to glutamate, NMDA, or mitochondrial toxins (PMID: 12153528). As an acidogen, L-2-hydroxyglutaric acid is classified as an alpha hydroxy acid belonging to the general class of compounds known as organic acids. Chronically high levels of L-2-hydroxyglutaric acid are characteristic of the inborn error of metabolism called L-2-hydroxyglutaric aciduria. Abnormally high levels of organic acids in the blood (organic acidemia), urine (organic aciduria), the brain, and other tissues lead to general metabolic acidosis. Acidosis typically occurs when arterial pH falls below 7.35. In infants with acidosis, the initial symptoms include poor feeding, vomiting, loss of appetite, weak muscle tone (hypotonia), and lack of energy (lethargy). These can progress to heart abnormalities, kidney abnormalities, liver damage, seizures, coma, and possibly death. These are the symptoms typical of untreated L-2-hydroxyglutaric aciduria. Many affected children with organic acidemias experience intellectual disability or delayed development. In adults, acidosis or acidemia is characterized by headaches, confusion, feeling tired, tremors, sleepiness, and seizures. |
|---|
| Compound Type | - Animal Toxin
- Food Toxin
- Metabolite
- Natural Compound
- Organic Compound
|
|---|
| Chemical Structure | |
|---|
| Synonyms | | Synonym | | (S)-2-Hydroxyglutarate | | (S)-2-Hydroxyglutaric acid | | (S)-a-Hydroxyglutarate | | (S)-a-Hydroxyglutaric acid | | (S)-alpha-Hydroxyglutarate | | (S)-alpha-Hydroxyglutaric acid | | 2-Hydroxy-(S)-Pentanedioate | | 2-Hydroxy-(S)-Pentanedioic acid | | 2-Hydroxy-L-Glutarate | | 2-Hydroxy-L-Glutaric acid | | L-2-Hydroxyglutarate | | L-a-Hydroxyglutarate | | L-a-Hydroxyglutaric acid | | L-alpha-Hydroxyglutarate | | L-alpha-Hydroxyglutaric acid |
|
|---|
| Chemical Formula | C5H8O5 |
|---|
| Average Molecular Mass | 148.114 g/mol |
|---|
| Monoisotopic Mass | 148.037 g/mol |
|---|
| CAS Registry Number | 13095-48-2 |
|---|
| IUPAC Name | (2S)-2-hydroxypentanedioic acid |
|---|
| Traditional Name | L-2-hydroxyglutaric acid |
|---|
| SMILES | [H][C@](O)(CCC(O)=O)C(O)=O |
|---|
| InChI Identifier | InChI=1S/C5H8O5/c6-3(5(9)10)1-2-4(7)8/h3,6H,1-2H2,(H,7,8)(H,9,10)/t3-/m0/s1 |
|---|
| InChI Key | InChIKey=HWXBTNAVRSUOJR-VKHMYHEASA-N |
|---|
| Chemical Taxonomy |
|---|
| Description | belongs to the class of organic compounds known as short-chain hydroxy acids and derivatives. These are hydroxy acids with an alkyl chain the contains less than 6 carbon atoms. |
|---|
| Kingdom | Organic compounds |
|---|
| Super Class | Organic acids and derivatives |
|---|
| Class | Hydroxy acids and derivatives |
|---|
| Sub Class | Short-chain hydroxy acids and derivatives |
|---|
| Direct Parent | Short-chain hydroxy acids and derivatives |
|---|
| Alternative Parents | |
|---|
| Substituents | - Short-chain hydroxy acid
- Fatty acid
- Monosaccharide
- Dicarboxylic acid or derivatives
- Alpha-hydroxy acid
- Secondary alcohol
- Carboxylic acid
- Carboxylic acid derivative
- Organic oxygen compound
- Organic oxide
- Hydrocarbon derivative
- Organooxygen compound
- Carbonyl group
- Alcohol
- Aliphatic acyclic compound
|
|---|
| Molecular Framework | Aliphatic acyclic compounds |
|---|
| External Descriptors | |
|---|
| Biological Properties |
|---|
| Status | Detected and Not Quantified |
|---|
| Origin | Endogenous |
|---|
| Cellular Locations | |
|---|
| Biofluid Locations | Not Available |
|---|
| Tissue Locations | Not Available |
|---|
| Pathways | | Name | SMPDB Link | KEGG Link |
|---|
| 2-Hydroxyglutric Aciduria (D And L Form) | SMP00136 | Not Available |
|
|---|
| Applications | Not Available |
|---|
| Biological Roles | Not Available |
|---|
| Chemical Roles | Not Available |
|---|
| Physical Properties |
|---|
| State | Solid |
|---|
| Appearance | White powder. |
|---|
| Experimental Properties | | Property | Value |
|---|
| Melting Point | Not Available | | Boiling Point | Not Available | | Solubility | Not Available | | LogP | Not Available |
|
|---|
| Predicted Properties | |
|---|
| Spectra |
|---|
| Spectra | | Spectrum Type | Description | Splash Key | Deposition Date | View |
|---|
| Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | Not Available | 2021-10-12 | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | Not Available | 2021-10-12 | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_1_1) - 70eV, Positive | Not Available | 2021-11-05 | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_1_2) - 70eV, Positive | Not Available | 2021-11-05 | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_1_3) - 70eV, Positive | Not Available | 2021-11-05 | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_2_1) - 70eV, Positive | Not Available | 2021-11-05 | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_2_2) - 70eV, Positive | Not Available | 2021-11-05 | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_2_3) - 70eV, Positive | Not Available | 2021-11-05 | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TMS_3_1) - 70eV, Positive | Not Available | 2021-11-05 | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TBDMS_1_1) - 70eV, Positive | Not Available | 2021-11-05 | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TBDMS_1_2) - 70eV, Positive | Not Available | 2021-11-05 | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TBDMS_1_3) - 70eV, Positive | Not Available | 2021-11-05 | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TBDMS_2_1) - 70eV, Positive | Not Available | 2021-11-05 | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TBDMS_2_2) - 70eV, Positive | Not Available | 2021-11-05 | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TBDMS_2_3) - 70eV, Positive | Not Available | 2021-11-05 | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (TBDMS_3_1) - 70eV, Positive | Not Available | 2021-11-05 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - Quattro_QQQ 10V, Positive (Annotated) | splash10-05mk-1900000000-c8b7818c2a1224f0b694 | 2012-07-24 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - Quattro_QQQ 25V, Positive (Annotated) | splash10-000i-9000000000-fc923f875e23e766078c | 2012-07-24 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - 40V, Negative | splash10-0a4i-9000000000-9fb296154571d7ea7d13 | 2021-09-20 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - 20V, Negative | splash10-0a4i-9400000000-821ae25c1b9ac04003b7 | 2021-09-20 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - 10V, Negative | splash10-004i-3900000000-dc8454bb187902d1a08b | 2021-09-20 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-001i-1900000000-68ac8b23cf0ed0cf09f5 | 2019-02-23 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-0m5r-9600000000-fb8fb6611cf2b1bca9f7 | 2019-02-23 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-0a4i-9000000000-4912ecf3489ec68e9641 | 2019-02-23 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-002b-0900000000-116ef9fca434a654a8f9 | 2019-02-23 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-0zi9-5900000000-76cefbf1a7e32ce74bef | 2019-02-23 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-0a4i-9000000000-c4562468d1b81effdb63 | 2019-02-23 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-0w39-4900000000-1385b471f7cdf14ca14d | 2021-09-22 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-052r-9500000000-17ece6956644d4cf2069 | 2021-09-22 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-0a4r-9000000000-dbb4195080bddb5f6210 | 2021-09-22 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-0fbi-3900000000-f88fcf3ac0dd06636b16 | 2021-09-23 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-0k9i-9400000000-31bf8d3bb7203e7bf200 | 2021-09-23 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-0a4u-9000000000-c0ac45c4d66178ac20cf | 2021-09-23 | View Spectrum | | 1D NMR | 1H NMR Spectrum (1D, 500 MHz, H2O, experimental) | Not Available | 2012-12-04 | View Spectrum | | 1D NMR | 13C NMR Spectrum (1D, 100 MHz, D2O, predicted) | Not Available | 2021-09-29 | View Spectrum | | 1D NMR | 1H NMR Spectrum (1D, 100 MHz, D2O, predicted) | Not Available | 2021-09-29 | View Spectrum | | 1D NMR | 13C NMR Spectrum (1D, 1000 MHz, D2O, predicted) | Not Available | 2021-09-29 | View Spectrum | | 1D NMR | 1H NMR Spectrum (1D, 1000 MHz, D2O, predicted) | Not Available | 2021-09-29 | View Spectrum | | 1D NMR | 13C NMR Spectrum (1D, 200 MHz, D2O, predicted) | Not Available | 2021-09-29 | View Spectrum | | 1D NMR | 1H NMR Spectrum (1D, 200 MHz, D2O, predicted) | Not Available | 2021-09-29 | View Spectrum | | 1D NMR | 13C NMR Spectrum (1D, 300 MHz, D2O, predicted) | Not Available | 2021-09-29 | View Spectrum | | 1D NMR | 1H NMR Spectrum (1D, 300 MHz, D2O, predicted) | Not Available | 2021-09-29 | View Spectrum | | 1D NMR | 13C NMR Spectrum (1D, 400 MHz, D2O, predicted) | Not Available | 2021-09-29 | View Spectrum | | 1D NMR | 1H NMR Spectrum (1D, 400 MHz, D2O, predicted) | Not Available | 2021-09-29 | View Spectrum | | 1D NMR | 13C NMR Spectrum (1D, 500 MHz, D2O, predicted) | Not Available | 2021-09-29 | View Spectrum | | 1D NMR | 1H NMR Spectrum (1D, 500 MHz, D2O, predicted) | Not Available | 2021-09-29 | View Spectrum | | 1D NMR | 13C NMR Spectrum (1D, 600 MHz, D2O, predicted) | Not Available | 2021-09-29 | View Spectrum | | 1D NMR | 1H NMR Spectrum (1D, 600 MHz, D2O, predicted) | Not Available | 2021-09-29 | View Spectrum | | 1D NMR | 13C NMR Spectrum (1D, 700 MHz, D2O, predicted) | Not Available | 2021-09-29 | View Spectrum | | 1D NMR | 1H NMR Spectrum (1D, 700 MHz, D2O, predicted) | Not Available | 2021-09-29 | View Spectrum | | 1D NMR | 13C NMR Spectrum (1D, 800 MHz, D2O, predicted) | Not Available | 2021-09-29 | View Spectrum | | 1D NMR | 1H NMR Spectrum (1D, 800 MHz, D2O, predicted) | Not Available | 2021-09-29 | View Spectrum | | 1D NMR | 13C NMR Spectrum (1D, 900 MHz, D2O, predicted) | Not Available | 2021-09-29 | View Spectrum | | 1D NMR | 1H NMR Spectrum (1D, 900 MHz, D2O, predicted) | Not Available | 2021-09-29 | View Spectrum | | 1D NMR | 1H NMR Spectrum (1D, 500 MHz, H2O, experimental) | Not Available | 2021-10-10 | View Spectrum | | 1D NMR | 1H NMR Spectrum (1D, 500 MHz, H2O, experimental) | Not Available | 2021-10-10 | View Spectrum | | 2D NMR | [1H, 13C]-HSQC NMR Spectrum (2D, 600 MHz, H2O, experimental) | Not Available | 2012-12-05 | View Spectrum |
|
|---|
| Toxicity Profile |
|---|
| Route of Exposure | Endogenous, Injection |
|---|
| Mechanism of Toxicity | 2-hydroxyglutarate is an oncometabolite. It is a competitive inhibitor of multiple α-ketoglutarate-dependent dioxygenases, including histone demethylases and the TET family of 5-methlycytosine (5mC) hydroxylases. As a result, high levels of 2-hydroxyglutarate lead to genome-wide histone and DNA methylation alterations, which in turn lead to mutations that ultimately cause cancer. L-2-hydroxyglutarate mediates its neurotoxicity through activation of N-methyl-D-aspartate receptors. L-2-hydroxyglutarate is structurally similar to the excitatory amino acid glutamate and stimulates neurodegeneration by mechanisms well-known for glutamate, NMDA or mitochondrial toxins (7). |
|---|
| Metabolism | Not Available |
|---|
| Toxicity Values | Not Available |
|---|
| Lethal Dose | Not Available |
|---|
| Carcinogenicity (IARC Classification) | Not listed by IARC. Has been implicated in oncogenesis (8, 9). |
|---|
| Uses/Sources | Not Available |
|---|
| Minimum Risk Level | Not Available |
|---|
| Health Effects | Chronically high levels of L-2-hydroxyglutaric acid are associated with the inborn error of metabolism called: 2-Hydroxyglutaric Aciduria. L-2-hydroxyglutarate is neurotoxic. L-2-hydroxyglutarate mediates its neurotoxicity through activation of N-methyl-D-aspartate receptors. Symptoms of L-2-Hydroxyglutaric Aciduria include hypotonia, tremors, and epilepsy declining into spongiform leukoencephalopathy, muscular choreodystonia, mental retardation, and psychomotor regression. |
|---|
| Symptoms | Chronic exposure of high levels of L-2-hydroxyglutarate leads to hypotonia, tremors, and epilepsy declining into spongiform leukoencephalopathy, muscular choreodystonia, mental retardation, and psychomotor regression. |
|---|
| Treatment | Not Available |
|---|
| Normal Concentrations |
|---|
| Not Available |
|---|
| Abnormal Concentrations |
|---|
| Not Available |
|---|
| External Links |
|---|
| DrugBank ID | Not Available |
|---|
| HMDB ID | HMDB00694 |
|---|
| PubChem Compound ID | 439939 |
|---|
| ChEMBL ID | CHEMBL1615211 |
|---|
| ChemSpider ID | 388969 |
|---|
| KEGG ID | C03196 |
|---|
| UniProt ID | Not Available |
|---|
| OMIM ID | |
|---|
| ChEBI ID | 32797 |
|---|
| BioCyc ID | Not Available |
|---|
| CTD ID | Not Available |
|---|
| Stitch ID | Not Available |
|---|
| PDB ID | S2G |
|---|
| ACToR ID | Not Available |
|---|
| Wikipedia Link | Not Available |
|---|
| References |
|---|
| Synthesis Reference | Kobayashi, Hidehiko; Yamaguchi, Koretaka; Yamashita, Takeshi. a-Hydroxyglutaric acid from glutamic acid. Jpn. Tokkyo Koho (1968), 3 pp. |
|---|
| MSDS | Link |
|---|
| General References | - Guneral F, Bachmann C: Age-related reference values for urinary organic acids in a healthy Turkish pediatric population. Clin Chem. 1994 Jun;40(6):862-6. [8087979 ]
- Gibson KM, ten Brink HJ, Schor DS, Kok RM, Bootsma AH, Hoffmann GF, Jakobs C: Stable-isotope dilution analysis of D- and L-2-hydroxyglutaric acid: application to the detection and prenatal diagnosis of D- and L-2-hydroxyglutaric acidemias. Pediatr Res. 1993 Sep;34(3):277-80. [8134166 ]
- Fujitake J, Ishikawa Y, Fujii H, Nishimura K, Hayakawa K, Inoue F, Terada N, Okochi M, Tatsuoka Y: L-2-hydroxyglutaric aciduria: two Japanese adult cases in one family. J Neurol. 1999 May;246(5):378-82. [10399870 ]
- Barth PG, Hoffmann GF, Jaeken J, Lehnert W, Hanefeld F, van Gennip AH, Duran M, Valk J, Schutgens RB, Trefz FK, et al.: L-2-hydroxyglutaric acidemia: a novel inherited neurometabolic disease. Ann Neurol. 1992 Jul;32(1):66-71. [1642474 ]
- Rashed MS, AlAmoudi M, Aboul-Enein HY: Chiral liquid chromatography tandem mass spectrometry in the determination of the configuration of 2-hydroxyglutaric acid in urine. Biomed Chromatogr. 2000 Aug;14(5):317-20. [10960831 ]
- de Klerk JB, Huijmans JG, Stroink H, Robben SG, Jakobs C, Duran M: L-2-hydroxyglutaric aciduria: clinical heterogeneity versus biochemical homogeneity in a sibship. Neuropediatrics. 1997 Dec;28(6):314-7. [9453028 ]
- Kölker S, Pawlak V, Ahlemeyer B, Okun JG, Hörster F, Mayatepek E, Krieglstein J, Hoffmann GF, Köhr G. NMDA receptor activation and respiratory chain complex V inhibition contribute to neurodegeneration in d-2-hydroxyglutaric aciduria. Eur J Neurosci. 2002 Jul;16(1):21-8. [12153528 ]
- Chowdhury R, Yeoh KK, Tian YM, Hillringhaus L, Bagg EA, Rose NR, Leung IK, Li XS, Woon EC, Yang M, McDonough MA, King ON, Clifton IJ, Klose RJ, Claridge TD, Ratcliffe PJ, Schofield CJ, Kawamura A. The oncometabolite 2-hydroxyglutarate inhibits histone lysine demethylases. EMBO Rep. 2011 May;12(5):463-9. doi: 10.1038/embor.2011.43. Epub 2011 Apr 1. [21460794 ]
- Xu W, Yang H, Liu Y, Yang Y, Wang P, Kim SH, Ito S, Yang C, Wang P, Xiao MT, Liu LX, Jiang WQ, Liu J, Zhang JY, Wang B, Frye S, Zhang Y, Xu YH, Lei QY, Guan KL, Zhao SM, Xiong Y. Oncometabolite 2-hydroxyglutarate is a competitive inhibitor of α-ketoglutarate-dependent dioxygenases. Cancer Cell. 2011 Jan 18;19(1):17-30. doi: 10.1016/j.ccr.2010.12.014. [21251613 ]
- International Agency for Research on Cancer (2014). IARC Monographs on the Evaluation of Carcinogenic Risks to Humans. [Link]
|
|---|
| Gene Regulation |
|---|
| Up-Regulated Genes | Not Available |
|---|
| Down-Regulated Genes | Not Available |
|---|