| Record Information |
|---|
| Version | 2.0 |
|---|
| Creation Date | 2009-07-05 02:56:35 UTC |
|---|
| Update Date | 2014-12-24 20:25:42 UTC |
|---|
| Accession Number | T3D2557 |
|---|
| Identification |
|---|
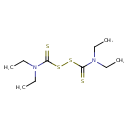
| Common Name | Disulfiram |
|---|
| Class | Small Molecule |
|---|
| Description | A carbamate derivative used as an alcohol deterrent. It is a relatively nontoxic substance when administered alone, but markedly alters the intermediary metabolism of alcohol. When alcohol is ingested after administration of disulfiram, blood acetaldehyde concentrations are increased, followed by flushing, systemic vasodilation, respiratory difficulties, nausea, hypotension, and other symptoms (acetaldehyde syndrome). It acts by inhibiting aldehyde dehydrogenase. [PubChem] |
|---|
| Compound Type | - Alcohol Deterrent
- Amine
- Drug
- Enzyme Inhibitor
- Metabolite
- Organic Compound
- Synthetic Compound
|
|---|
| Chemical Structure | |
|---|
| Synonyms | | Synonym | | 1,1'-dithiobis(N,N-diethylthioformamide) | | Antabus | | Antabuse | | Antaethyl | | Antalcol | | Anticol | | Antietanol | | Bis(diethylthiocarbamoyl) disulfide | | Chronol | | Disulfuram | | Disulphuram | | Dupon 4472 | | Dupont Fungicide 4472 | | Esperal | | N,N,N',N'-tetraethylthiuram disulfide | | TATD | | TETD | | Tetraethylthioperoxydicarbonic Diamide | | Tetraethylthiram Disulfide | | Tetraethylthiram Disulphide | | Tetraethylthiuram | | Tetraethylthiuram Disulfide | | Tetraethylthiuram Disulphide | | Tetraethylthiuram Sulfide | | Tetraethylthiuran Disulfide | | TTD | | Usaf B-33 |
|
|---|
| Chemical Formula | C10H20N2S4 |
|---|
| Average Molecular Mass | 296.539 g/mol |
|---|
| Monoisotopic Mass | 296.051 g/mol |
|---|
| CAS Registry Number | 97-77-8 |
|---|
| IUPAC Name | N,N-diethyl[(diethylcarbamothioyl)disulfanyl]carbothioamide |
|---|
| Traditional Name | disulfiram |
|---|
| SMILES | CCN(CC)C(=S)SSC(=S)N(CC)CC |
|---|
| InChI Identifier | InChI=1S/C10H20N2S4/c1-5-11(6-2)9(13)15-16-10(14)12(7-3)8-4/h5-8H2,1-4H3 |
|---|
| InChI Key | InChIKey=AUZONCFQVSMFAP-UHFFFAOYSA-N |
|---|
| Chemical Taxonomy |
|---|
| Description | belongs to the class of organic compounds known as thiuram disulfides. These are organic disulfides that have the general structural formula RN(R')C(=S)SSC(=S)N(R\")R\"', where R-R\"'=alkyl groups. |
|---|
| Kingdom | Organic compounds |
|---|
| Super Class | Organic nitrogen compounds |
|---|
| Class | Organonitrogen compounds |
|---|
| Sub Class | Amines |
|---|
| Direct Parent | Thiuram disulfides |
|---|
| Alternative Parents | |
|---|
| Substituents | - Thiuram disulfide
- Organic disulfide
- Sulfenyl compound
- Organopnictogen compound
- Hydrocarbon derivative
- Organosulfur compound
- Aliphatic acyclic compound
|
|---|
| Molecular Framework | Aliphatic acyclic compounds |
|---|
| External Descriptors | |
|---|
| Biological Properties |
|---|
| Status | Detected and Not Quantified |
|---|
| Origin | Exogenous |
|---|
| Cellular Locations | |
|---|
| Biofluid Locations | Not Available |
|---|
| Tissue Locations | Not Available |
|---|
| Pathways | | Name | SMPDB Link | KEGG Link |
|---|
| Disulfiram Pathway | Not Available | Not Available |
|
|---|
| Applications | Not Available |
|---|
| Biological Roles | |
|---|
| Chemical Roles | |
|---|
| Physical Properties |
|---|
| State | Solid |
|---|
| Appearance | White powder. |
|---|
| Experimental Properties | | Property | Value |
|---|
| Melting Point | 71.5°C | | Boiling Point | 117°C at 1.70E+01 mm Hg | | Solubility | 4.09 mg/L (at 25°C) | | LogP | 3.88 |
|
|---|
| Predicted Properties | |
|---|
| Spectra |
|---|
| Spectra | | Spectrum Type | Description | Splash Key | Deposition Date | View |
|---|
| Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | splash10-016r-8930000000-ff4b800eacdb52d66d80 | 2017-09-01 | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - GC-MS (Non-derivatized) - 70eV, Positive | Not Available | 2021-10-12 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-qTof , Positive | splash10-014i-0900000000-274003127d4c34f9abcc | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-qTof , Positive | splash10-002b-1910000000-0371815b89a4378bec81 | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-qTof , Positive | splash10-004i-0950000000-ca28d8dfdf780c201716 | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-qTof , Positive | splash10-004i-0950000000-ca28d8dfdf780c201716 | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - LC-ESI-qTof , Positive | splash10-0a4i-5910000000-7dd7b7fbb29b6ae08f4c | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - , positive | splash10-00xr-0904000000-2d29367629d096da4708 | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - , positive | splash10-00xr-1914000000-0022574151f222c8050f | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - , positive | splash10-014j-0900000000-07fd862cac7fb47fa007 | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - , positive | splash10-00di-0903000000-bb7b2b338b8981b249ac | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - , positive | splash10-00kb-3910100000-263484085094fbdd45da | 2017-09-14 | View Spectrum | | LC-MS/MS | LC-MS/MS Spectrum - 35V, Positive | splash10-014s-5900000000-fe9f3fb3c9d4c94f6496 | 2021-09-20 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-0002-1970000000-87dad5bd832c248e4c73 | 2016-08-01 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-014i-3900000000-f7637bebace8002bdd78 | 2016-08-01 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-00di-9500000000-491cc5b9b0d4455ff674 | 2016-08-01 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-01vk-4690000000-64528d1f9696aa7a5d5d | 2016-08-03 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-02ft-9810000000-825b92854771a0e58c9d | 2016-08-03 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-0fdo-8910000000-afd3a638d74e8761c518 | 2016-08-03 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-014i-0910000000-cf16eb2676d288479eac | 2021-10-11 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-014i-2900000000-2765ff0428d56cf81049 | 2021-10-11 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-000i-9200000000-7f8c0eb726cf3929fa7f | 2021-10-11 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-0002-3690000000-f14241348cdf9f8196d2 | 2021-10-11 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-01ot-7900000000-1843982004d8ffd33a5f | 2021-10-11 | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-03di-9200000000-28f4b4954161ce60f733 | 2021-10-11 | View Spectrum | | MS | Mass Spectrum (Electron Ionization) | splash10-014r-9510000000-2e18f478f9c0f9a2958a | 2014-09-20 | View Spectrum | | 1D NMR | 1H NMR Spectrum (1D, 90 MHz, CDCl3, experimental) | Not Available | 2014-09-20 | View Spectrum | | 1D NMR | 13C NMR Spectrum (1D, 25.16 MHz, CDCl3, experimental) | Not Available | 2014-09-23 | View Spectrum |
|
|---|
| Toxicity Profile |
|---|
| Route of Exposure | Inhalation (MSDS, A308); ingestion (MSDS, A308); dermal (MSDS, A308) ; eye contact (MSDS, A308)
Disulfiram is absorbed slowly from the gastrointestinal tract (80 to 90% of oral dose). |
|---|
| Mechanism of Toxicity | Disulfiram blocks the oxidation of alcohol at the acetaldehyde stage during alcohol metabolism following disulfiram intake causing an accumulation of acetaldehyde in the blood producing highly unpleasant symptoms. Disulfiram blocks the oxidation of alcohol through its irreversible inactivation of aldehyde dehydrogenase, which acts in the second step of ethanol utilization. In addition, disulfiram competitively binds and inhibits the peripheral benzodiazepine receptor, which may indicate some value in the treatment of the symptoms of alcohol withdrawal, however this activity has not been extensively studied. |
|---|
| Metabolism | Disulfiram is completely absorbed from the human GI tract. However, a period of 12 hr is required for its full action, perhaps because, being highly sol in lipid, it is initially localized in fat. It is slowly metabolized in the liver to diethyldithiocarbamate, diethylamine, and carbon disulfide. Six hr after oral administration of the drug, one third of plasma disulfiram is in the form of diethyldithiocarbamate. Elimination is relatively slow, and about 1/5 still remains in body at end of a week. The greater part of the absorbed drug is excreted in the urine as the sulfate, partly free and partly esterified (4, 5). |
|---|
| Toxicity Values | LD50: 8.6g/kg (oral, rat). |
|---|
| Lethal Dose | Not Available |
|---|
| Carcinogenicity (IARC Classification) | 3, not classifiable as to its carcinogenicity to humans. (10) |
|---|
| Uses/Sources | For the treatment and management of chronic alcoholism (1). |
|---|
| Minimum Risk Level | Not Available |
|---|
| Health Effects | Occasionally implicated in producing psychosis, optic neuritis, and encephalopathy. Hematologic, neuromuscular, and gastrointestinal toxicity and hepatotoxicity may occur 10 days to 12 months after therapy is begun (7).
|
|---|
| Symptoms | Symptoms of overdose include irritation, slight drowsiness, unpleasant taste, mild GI disturbances, and orthostatic hypotension. |
|---|
| Treatment | Not Available |
|---|
| Normal Concentrations |
|---|
| Not Available |
|---|
| Abnormal Concentrations |
|---|
| Not Available |
|---|
| External Links |
|---|
| DrugBank ID | DB00822 |
|---|
| HMDB ID | HMDB14960 |
|---|
| PubChem Compound ID | 3117 |
|---|
| ChEMBL ID | CHEMBL964 |
|---|
| ChemSpider ID | 3005 |
|---|
| KEGG ID | C01692 |
|---|
| UniProt ID | Not Available |
|---|
| OMIM ID | |
|---|
| ChEBI ID | 4659 |
|---|
| BioCyc ID | DISULFIRAM |
|---|
| CTD ID | D004221 |
|---|
| Stitch ID | Disulfiram |
|---|
| PDB ID | Not Available |
|---|
| ACToR ID | 1353 |
|---|
| Wikipedia Link | Disulfiram |
|---|
| References |
|---|
| Synthesis Reference | Adams, H.S. and Meuser, L.; US.Patent 1,782,111; November 18,1930; assigned to The
Naugatuck Chemical Company.
Bailey, G.C.; U.S.Patent 1,796,977; March 17,1931; assigned to The Roessler & Hasslacher
Chemical Company. |
|---|
| MSDS | Link |
|---|
| General References | - Wishart DS, Knox C, Guo AC, Cheng D, Shrivastava S, Tzur D, Gautam B, Hassanali M: DrugBank: a knowledgebase for drugs, drug actions and drug targets. Nucleic Acids Res. 2008 Jan;36(Database issue):D901-6. Epub 2007 Nov 29. [18048412 ]
- Nash T, Rice WG: Efficacies of zinc-finger-active drugs against Giardia lamblia. Antimicrob Agents Chemother. 1998 Jun;42(6):1488-92. [9624499 ]
- Bouma MJ, Snowdon D, Fairlamb AH, Ackers JP: Activity of disulfiram (bis(diethylthiocarbamoyl)disulphide) and ditiocarb (diethyldithiocarbamate) against metronidazole-sensitive and -resistant Trichomonas vaginalis and Tritrichomonas foetus. J Antimicrob Chemother. 1998 Dec;42(6):817-20. [10052908 ]
- Segura-Aguilar J: Peroxidase activity of liver microsomal vitamin D 25-hydroxylase and cytochrome P450 1A2 catalyzes 25-hydroxylation of vitamin D3 and oxidation of dopamine to aminochrome. Biochem Mol Med. 1996 Jun;58(1):122-9. [8809353 ]
- David DJ, Bourin M, Hascoet M, Colombel MC, Baker GB, Jolliet P: Comparison of antidepressant activity in 4- and 40-week-old male mice in the forced swimming test: involvement of 5-HT1A and 5-HT1B receptors in old mice. Psychopharmacology (Berl). 2001 Feb;153(4):443-9. [11243491 ]
- Gaval-Cruz M, Weinshenker D: mechanisms of disulfiram-induced cocaine abstinence: antabuse and cocaine relapse. Mol Interv. 2009 Aug;9(4):175-87. doi: 10.1124/mi.9.4.6. [19720750 ]
- Rumack BH (2009). POISINDEX(R) Information System. Englewood, CO: Micromedex, Inc. CCIS Volume 141, edition expires Aug, 2009.
- McEvoy GK (ed) (2005). American Hospital Formulary Service - Drug Information 2005. Bethesda, MD: American Society of Health-System Pharmacists, Inc.
- Clayton GD and Clayton FE (eds) (1981-1982). Patty's Industrial Hygiene and Toxicology: Volume 2A, 2B, 2C: Toxicology. 3rd ed. New York: John Wiley Sons.
- International Agency for Research on Cancer (2014). IARC Monographs on the Evaluation of Carcinogenic Risks to Humans. [Link]
- Drugs.com [Link]
- Gaval-Cruz M, Weinshenker D: mechanisms of disulfiram-induced cocaine abstinence: antabuse and cocaine relapse. Mol Interv. 2009 Aug;9(4):175-87. [Link]
|
|---|
| Gene Regulation |
|---|
| Up-Regulated Genes | | Gene | Gene Symbol | Gene ID | Interaction | Chromosome | Details |
|---|
|
|---|
| Down-Regulated Genes | | Gene | Gene Symbol | Gene ID | Interaction | Chromosome | Details |
|---|
|
|---|